Abstract
Objective:
To develop a mouse cell biosensor system for the high-throughput genotoxicity detection of chemicals, such as environmental pollutants.
Method:
We developed a novel reporter vector pGL4-GFP, wherein the firefly luciferase reporter gene in the pGL4.82 vector was replaced by the green fluorescent protein (GFP) gene from the pAcGFP1-N1 vector. To construct the reporter pGL4-p53-GFP (p53 promoter linked to GFP), a fragment containing the p53 gene promoter was generated by amplifying a region from −481 to +180 of mouse genomic DNA isolated from mouse tail tissue. We developed a mouse cell biosensor system for the high-throughput genotoxicity detection of new drugs by stably integrating the reporter plasmid of pGL4-p53-GFP into the mouse embryonic fibroblast cells. Various genotoxic agents were used to treat this biosensor system. The resulting fluorescence was directly observed under a fluorescence microscope, and the GFP protein level was measured through Western blot analysis.
Result:
The biosensor system was treated with genotoxic agents, such as doxorubicin, cyclophosphamide, and benzo(a)pyrene. The GFP protein expression was significantly increased in cells exposed to genotoxic agents but negatively responded to the non-genotoxic agent dimethyl sulfoxide, thereby proving the specificity and sensitivity of the biosensor system.
Conclusion:
This novel in vitro biosensor system can be especially useful in genotoxicity detection.
Introduction
Fundamental to the initiation of most tumors is DNA damage, which, if inaccurately or inappropriately repaired, may lead to the activation, deregulation, or overexpression of oncogenes that drive cell proliferation or survival in the absence of physiological stimuli. 1 Accumulating evidence suggests that the pathways of DNA damage response may mediate tumor suppression by activating p53 in response to the persistent DNA damage and genomic instability. 2 Therefore, biosensor systems based on genotoxicity detection are useful tools for assessing the genotoxic effects of environmental chemicals on human health. 3
To date, cell-based bioassays that include bacterial and mammalian systems have been developed and are currently in use to detect genotoxic agents. 4 However, all these assays have their shortcomings. For example, the Ames test is the most widely used assay for bacteria; it is based on the reverse mutation of genetically modified bacterial strains. 5 This method is used to detect gene mutations caused by chemicals or irradiation, but it has relatively low sensitivity, which results in its frequent failures to identify the genotoxic properties of some compounds. 6 The comet assay is a mammalian cell-based genotoxicity test widely used to detect DNA damage, such as DNA strand breaks in mammalian cells; it is rapid and sensitive but has a high rate of false positives. 7 Several cell-based gene mutation assays are available, such as the hamster v97 and CHO cells, the mouse lymphoma L5178Y cells, and the human lymphoblastoid TK6 cells, all of which are well validated and widely used. However, these cell-based gene mutation assays have relatively low sensitivity. 8
Previous studies indicated that the p53 gene expression is highly inducible by several stimuli, such as serum stimulation and phorbol esters. 9 More recent studies have shown that the p53 gene expression is regulated in response to various genotoxic agents. 10 Various genotoxic agents, including some anticancer drugs, can also induce p53 promoter activation. 11 Therefore, we hypothesize that the p53 promoter sequence can be used to detect DNA damage and the high-throughput detection of genotoxic chemicals.
We describe the development and initial validation of a p53-mediated green fluorescent protein (GFP) reporter gene bioassay system for genotoxicity detection in vitro. The obtained results show that this biosensor system can be used for the rapid high-throughput screening of genotoxic agents.
Materials and methods
Chemicals
Doxorubicin, dimethyl sulfoxide, cyclophosphamide, and cisplatin were purchased from Sigma-Aldrich (St Louis, Missouri, USA). Benzo(a)pyrene was a gift from Dr Yao (Tobacco Research Institute of Yunnan Province).
Cell lines and plasmids
The pAcGFP1-N1 vector was purchased from Clontech (Mountain View, California, USA), while the pGL4.82 vector was purchased from Promega (Madison, Wisconsin, USA). The firefly luciferase reporter gene in the pGL4.82 vector was replaced by the GFP gene, which was obtained from the pAcGFP1-N1 vector by digestion with the EcoRI and AflII restriction enzymes. The resulting promoterless plasmid was designated as pGL4-GFP. To construct the reporter pGL4-p53-GFP, a fragment containing the p53 gene promoter was generated by amplifying a region from −481 to +180 of the p53 gene in mouse genomic DNA, which was isolated from mouse tail tissue. Amplification was performed with 5′-tggctcgagGTCTTTACAGAGAGTG-3′ and 5′-cgagatctCGGAGAAGCGTGACA-3′ as forward and reverse primers, respectively. These forward and reverse primers contain recognition sites specifically designed for the XhoI and BglII restriction enzymes respectively. After digestion with XhoI and BglII and purification, the fragment was cloned into the XhoI and BglII sites upstream of the GFP reporter gene in the pGL4-GFP vector. Plasmids were purified with the TIAN prep Plasmid Kit (TIANGEN, Beijing, China) according to the manufacturer’s instructions.
The pGL4-p53-GFP construct and empty vector pGL4-GFP construct were introduced into mouse embryonic fibroblast (MEF) cells through the LipofectAMINE 2000 method (Invitrogen, Carlsbad, California, USA) according to the manufacturer’s instructions. Antibiotic selection for 1−2 months was used to select the stably transformed cells. Puromycin was used to maintain the stably transformed cells. The cells were cultured in Dulbecco’s modified eagle’s medium, supplemented with 10% fetal bovine serum (Hyclone, Novato, CaliforniaA, USA) in a 3% oxygen and 5% carbondioxide incubator at 37°C.
Challenge of the pGL4-p53-GFP cells with genotoxic agents
Doxorubicin, cyclophosphamide, cisplatin, and benzo(a)pyrene are direct-acting genotoxic agents with known mechanisms. Dimethyl sulfoxide has been reported as a non-genotoxic agent. The above-mentioned agents were used to challenge the pGL4-p53-GFP cells and examine the cellular responses to genotoxic and non-genotoxic agents.
The pGL4-p53-GFP cells were collected and diluted to a cell density of 1 × 105 cells/mL. The cell suspension was seeded into 24-well culture plates with 1 mL/well at 20°C overnight. The old medium was replaced by fresh medium the following day. The cells were treated with doxorubicin (0.5 μg/mL), cyclophosphamide (50 μg/mL), cisplatin (10 μM), or benzo(a)pyrene (10 μM) for 24 h. The treated pGL4-p53-GFP cells were directly observed under a fluorescence microscope. Fluorescence microscopy was performed with a Nikon Eclipse 90i fluorescence microscope and a Nikon Digital Sight CCD (controlled by NIS-Elements 3.0 software; Japan). The cells were lysed, and the cell lysates were collected for Western blot analysis.
Western blot analysis
Cell lysates were harvested, and the protein concentration was detected using the Bio-Rad Bradford protein assay kit (Hercules, California, USA) before 20 μg of protein was subjected to sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE). The proteins were separated by electrophoresis in 12% SDS-PAGE and transferred onto a polyvinylidene fluoride membrane (Millipore, Bedford, Massachusetts, USA). The transblotted membrane was incubated in 10% skim milk in Tris-buffered saline Tween 20 (25 mM Tris, pH 7.6; 150 mM sodium chloride; 0.1% Tween 20) for 1 h. The membrane was incubated overnight at 4°C with the respective primary antibody, that is, anti-GFP (1:2000 dilution; Clontech) or anti-tubulin (1:8000 dilution; Upstate, New York, USA). The primary antibodies were detected with an horseradish peroxidase-conjugated secondary antibody (enhanced chemiluminescence anti-mouse immunoglobulin G peroxidase, 1:5000 dilution; Sigma-Aldrich, St Louis, Missouri, USA) before the membranes were subjected to a chemiluminescence detection assay.
Results
Construction of p53 promoter-based reporter vector
Literature review and analyses with the relevant software demonstrated that the region of −481 to +180 of the mouse p53 gene contains the core promoter region of the gene. In addition, this region contained several transcription factor-binding sites, which bind to different transcription factors that influence p53 gene transcription. The constructed plasmid structure is illustrated in Figure 1, where the GFP reporter gene is under the control of the mouse p53 gene promoter. The construct was confirmed by restriction enzyme digestion and sequence analysis (Figure 2).
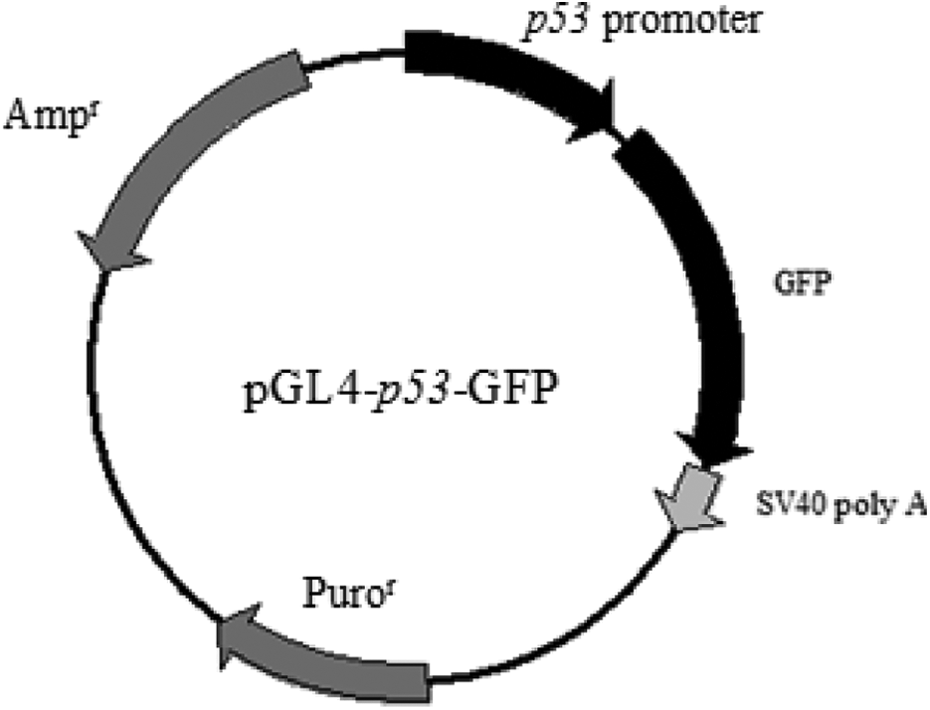
Structure of the p53 promoter-based reporter vector construct. The fragment containing the p53 core promoter region was cloned into the XhoI and BglII sites upstream of the GFP reporter gene in the pGL4-GFP vector. GFP: green fluorescent protein.

Sequence analysis of the region containing the p53 promoter in the pGL4-p53-GFP plasmid. GFP: green fluorescent protein.
Response of the pGL4-p53-GFP cells to doxorubicin and dimethyl sulfoxide
To ensure the specificity and sensitivity of the biosensor system for detecting genotoxic agents, the pGL4-p53-GFP cells were treated with 0.5 μg/mL of doxorubicin and 1% (v/v) dimethyl sulfoxide. As shown in Figure 3, the challenge of pGL4-p53-GFP cells with the genotoxic agent doxorubicin significantly upregulated the expression of the GFP in contrast to the untreated cells. As directly observed under a fluorescence microscope, the DNA damage-induced response was detected after 6 h of exposure and lasted for 48 h. However, the pGL4-p53-GFP cells treated with dimethyl sulfoxide did not induce the expression of the GFP reporter gene. The GFP protein level was confirmed by the Western blot analysis (Figure 4).

GFP induction by doxorubicin in pGL4-p53-GFP cells. (a) pGL4-p53-GFP cells with 24 h of no treatment control. (b) pGL4-p53-GFP cells treated with 0.5 μg/mL doxorubicin for 6 h (B), 24 h (c), and 48 h (d). (e) pGL4-p53-GFP cells treated with 1% (v/v) dimethyl sulfoxide for 24 h. GFP: green fluorescent protein.
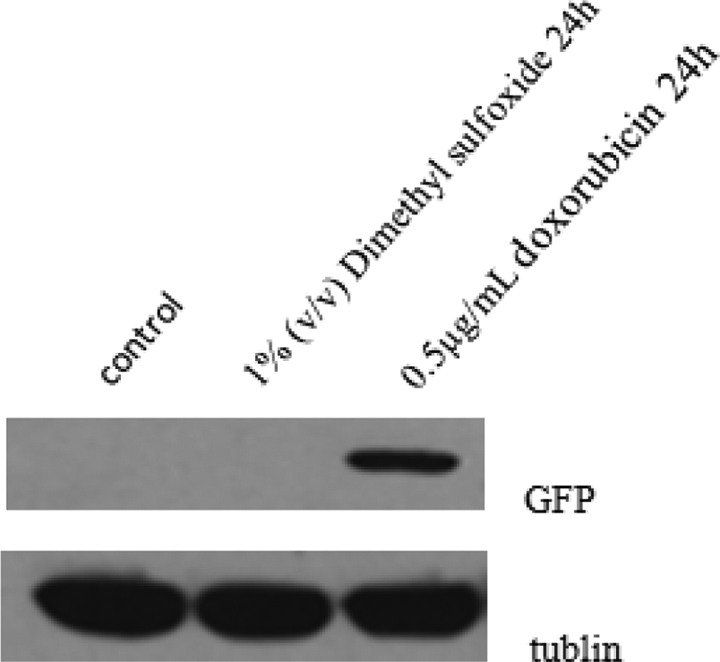
Western blot analysis of GFP protein levels. The GFP protein expression was induced in the pGL4-p53-GFP cells after treatment with 0.5 μg/mL doxorubicin for 24 h. The challenge with the non-genotoxic agent of 1% (v/v) dimethyl sulfoxide did not induce GFP expression in pGL4-p53-GFP cells. GFP: green fluorescent protein.
Response of the pGL4-p53-GFP cells to various genotoxic agents
Cyclophosphamide is a carcinogen that exerts its genotoxicity on cells. Cisplatin is a chemotherapy drug that binds to and cross-links with DNA to exert its genotoxicity on cells and ultimately trigger apoptosis. Benzo(a)pyrene is another genotoxic agent that has been shown to cause genetic damage in lung cells identical to the DNA damage observed in most malignant lung tumors. To further validate the specificity and sensitivity of the proposed biosensor system, pGL4-p53-GFP cells were exposed to 50 μg/mL of cyclophosphamide, 10 μM of cisplatin, or 10 μM of benzo(a)pyrene for 24 h. The expression of GFP was directly observed under a fluorescence microscope. As shown in Figure 5, cisplatin significantly induced the expression of the GFP gene. Similar results were observed after treatment with cyclophosphamide and benzo(a)pyrene (Figure 5). The GFP protein level was confirmed by Western blot analysis (Figure 6). Therefore, this biosensor system provides a specific and sensitive method for detecting genotoxic agents.
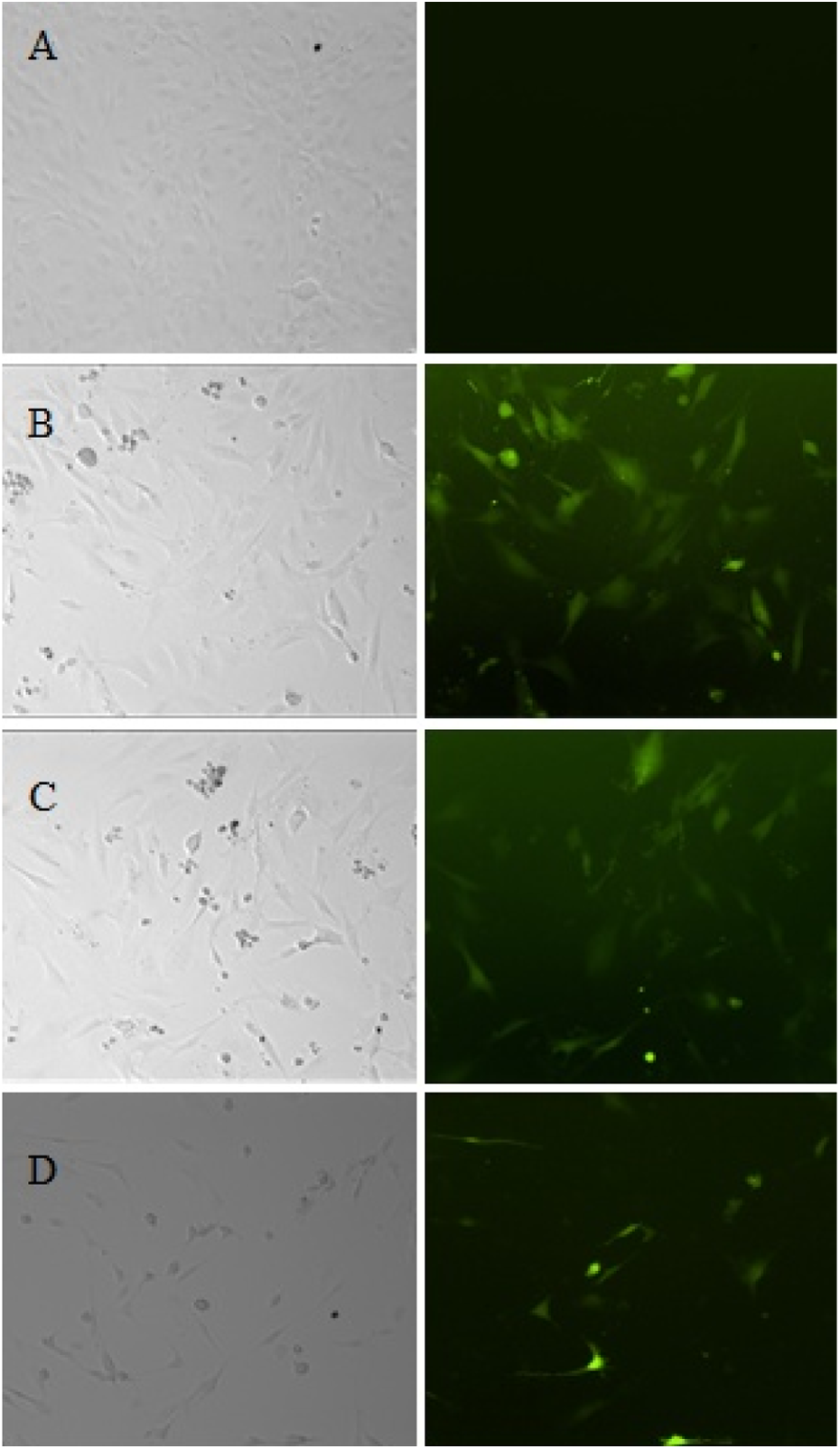
GFP induction by genotoxic agents in pGL4-p53-GFP cells. (a) pGL4-p53-GFP cells without treatment for 24 h (control). (b) pGL4-p53-GFP cells treated with 10 μM of cisplatin for 24 h. (c) pGL4-p53-GFP cells treated with 50 μg/mL of cyclophosphamide for 24 h. (d) pGL4-p53-GFP cells treated with 10 μM of benzo(a)pyrene for 24 h. GFP: green fluorescent protein.
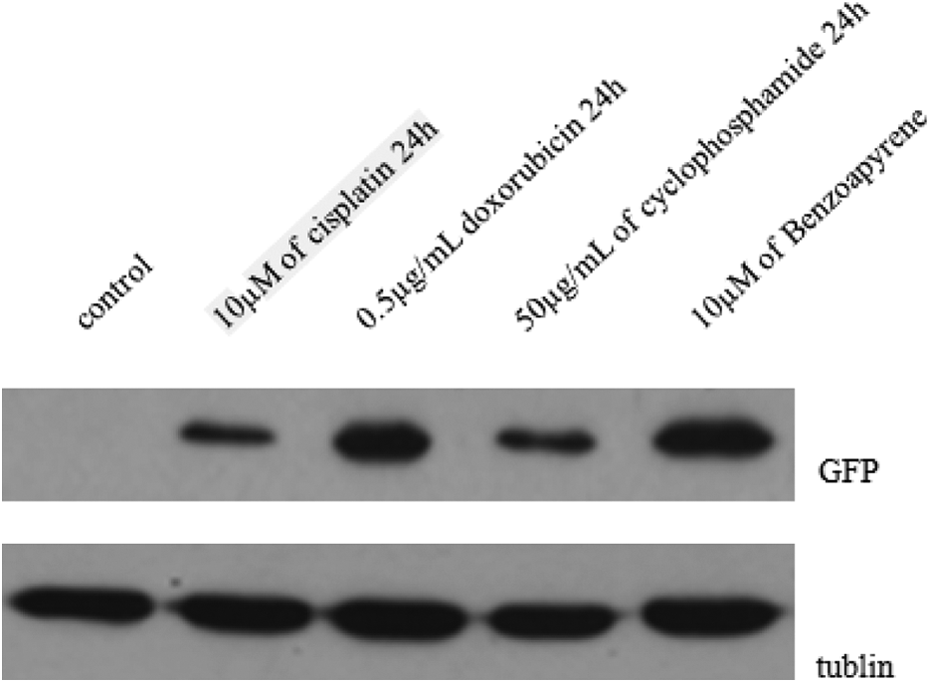
Western blot analysis of GFP protein levels. GFP proteins in pGL4-p53-GFP cells were induced after 24 h of treatment with 50 μg/mL of cyclophosphamide, 10 μM of cisplatin, or 10 μM of benzo(a)pyrene, respectively. GFP: green fluorescent protein.
Discussion
The activation of p53 in response to various genotoxic stresses is an important cellular response to maintain genome integrity. 12 Previous studies have proven that the p53 level is increased upon exposure to various genotoxic agents, a finding that suggests this increase is mainly due to the phosphorylation of p53, which enhances protein stability. However, subsequent studies showed that the p53 messenger RNA (mRNA) level is increased by two- to threefold after treatment of cells with various genotoxic agents, such as mitomycin C and 5-fluorouracil. 11 These results indicate that p53 genotoxic stress response is regulated at the protein and mRNA levels. Therefore, the p53 promoter region will be very useful for developing a reporter gene assay to detect DNA damage. The pGL4 reporter vector is a next-generation reporter gene vector that is optimized for high expression in mammalian cells and designed to reduce anomalous transcription. The vector is a promoterless basic vector that uses the luciferase reporter gene hRluc as its reporter gene. Figure 1 shows the constructed novel reporter vector from pAcGFP1-N1 vector, designated as pGL4-GFP, wherein the firefly luciferase reporter gene in pGL4.82 was replaced by the GFP gene. The GFP gene was placed under the control of the p53 promoter, such that the resulting fluorescence could be directly observed under a fluorescence microscope. This p53 promoter-based reporter vector could be useful in detecting DNA damage.
We reported the development of a p53 promoter-based biosensor system for genotoxicity detection by the stable integration of a pGL4-p53-GFP construct into MEF cells. The results for the initial validation of this biosensor system for genotoxicity detection are shown in Figures 3 and 4. The pGL4-p53-GFP cells were challenged with the model genotoxin doxorubicin and the non-genotoxin dimethyl sulfoxide. Doxorubicin is an antitumor drug commonly used to treat some leukemias and lymphomas. Previous studies showed that doxorubicin interacts with DNA through the intercalation and inhibition of macromolecular biosynthesis. 13 The treatment of mammalian cells with doxorubicin causes a significant increase in the cellular p53 level. 14 The results obtained in the challenge experiments showed that the expression of the GFP reporter gene in response to the doxorubicin-induced DNA damage is significantly upregulated but not in response to dimethyl sulfoxide. Furthermore, the response speed of this biosensor system to genotoxic agents is high, and the induced expression level of the reporter gene is relatively stable. To further validate this biosensor system, the pGL4-p53-GFP cells were treated with the genotoxic agents cisplatin, cyclophosphamide, and benzo(a)pyrene. As shown in Figure 5, cisplatin significantly induced the expression of GFP gene after 24 h of treatment. Similar results were observed after treatment with cyclophosphamide and benzo(a)pyrene. The results were further confirmed by Western blot analysis, as shown in Figure 6. Therefore, we believe this biosensor system can be used for the detection of agents with genotoxic potential.
In summary, we developed a p53 promoter-derived biosensor system, which may become a specific and sensitive tool for genotoxicity detection of novel chemicals and drugs. This biosensor system also provides a highly useful reporter system for excluding the nonspecific genotoxicity of chemicals in anticancer drug screening.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the Nature Science Foundation of Yunnan Province (No. 2013FC006)
