Abstract

For every experimental compound in the drug discovery pipeline, pharmaceutical companies generate huge amounts of data: chemical and physical data derived from various analytical techniques, and biological data that flow out of large-scale screening programs as well as lead optimization work. Drug discovery efforts have been hampered not by a lack of lead compounds or a dearth of experimental data, but by the need for effective and efficient computational tools to collect, store, manipulate, and analyze large amounts of data.
Scientists in many of the major pharmaceutical and biotechnology companies, including GlaxoSmithKline, Aventis, AstraZeneca, Hoffmann-La Roche, Merck, Novartis, Millennium, Exelixis, and Immunex, Cytokinetics, Evotec and Monsanto are using ActivityBase (Figure 1), an integrated data management system, to collect and analyze data generated by high throughput screening (HTS), to store chemical structures and register novel compounds, and to integrate cheminformatic and bioinformatic datasets.
ActivityBase manages data produced by HTS (with sustained screening volumes exceeding 30,000 to 40,000 wells/working day) and ultra-HTS (sustained volumes of greater than 100,000 wells/working day) and some companies are populating ActivityBase databases at the rate of approximately 20 million data points per six months. Many operational databases exceed tens of millions of rows, and the software's search engine can respond to typical queries in a matter of seconds.

IDBS' ActivityBase 5.0 enables for the first time the complete integration of biological and chemical data in drug discovery. Within IDBS' ActivityBase, conditional formatting is one of the ways users can quickly highlight results of interest in a screening run. Fully integrated chemistry tools mean that the chemical properties, as well as biological activity, of any of the compounds can be viewed directly from the ActivityBase user interface.
An abundance of data does not necessarily add value to an experimental compound. The data do not imply therapeutic efficacy, infer bioavailability, predict toxicity, or suggest drug-like properties. Successful discovery research depends on the ability to integrate diverse datasets from multiple sources and to extract information from raw data. It is this information that will guide and expedite decision-making, improve productivity, and add value. It is this information that will allow a company to decide whether to pursue a lead compound or to “fail” it early in the discovery process.
ActivityBase is based on IDBS' generic data model designed for discovery research and can capture, manage, and store data from biological, chemical, and robotic systems. The ActivityBase 5.0 Suite seamlessly integrates cheminformatic and bioinformatic data. It provides the framework for converting data into information that can be applied to lead discovery and optimization processes. (Figure 2)
New functionalities introduced in version 5.0 enhance the flexibility of data collection and analysis and expand data integration capabilities. Joining AssayBase, which manages biological data are three new software modules:

Biological, chemical and analytical data can be seamlessly integrated within the ActivityBase software environment from IDBS.
StructureBase – for registering chemical compounds and searching molecular structures and related physicochemical data. (Figure 3)
ReactionBase – for storing, managing, and searching chemical reactions and reaction schemes. (Figure 4)
Natural Products – for managing the process of isolating active compounds from natural materials; it generates a genealogic trail that tracks the derivation of new chemical compounds from natural products.

StructureBase is a complete molecule registration and handling system, developed using the latest industry standard technology. Full stereochemical information can be assigned to molecules registered and retrieved using StructureBase.
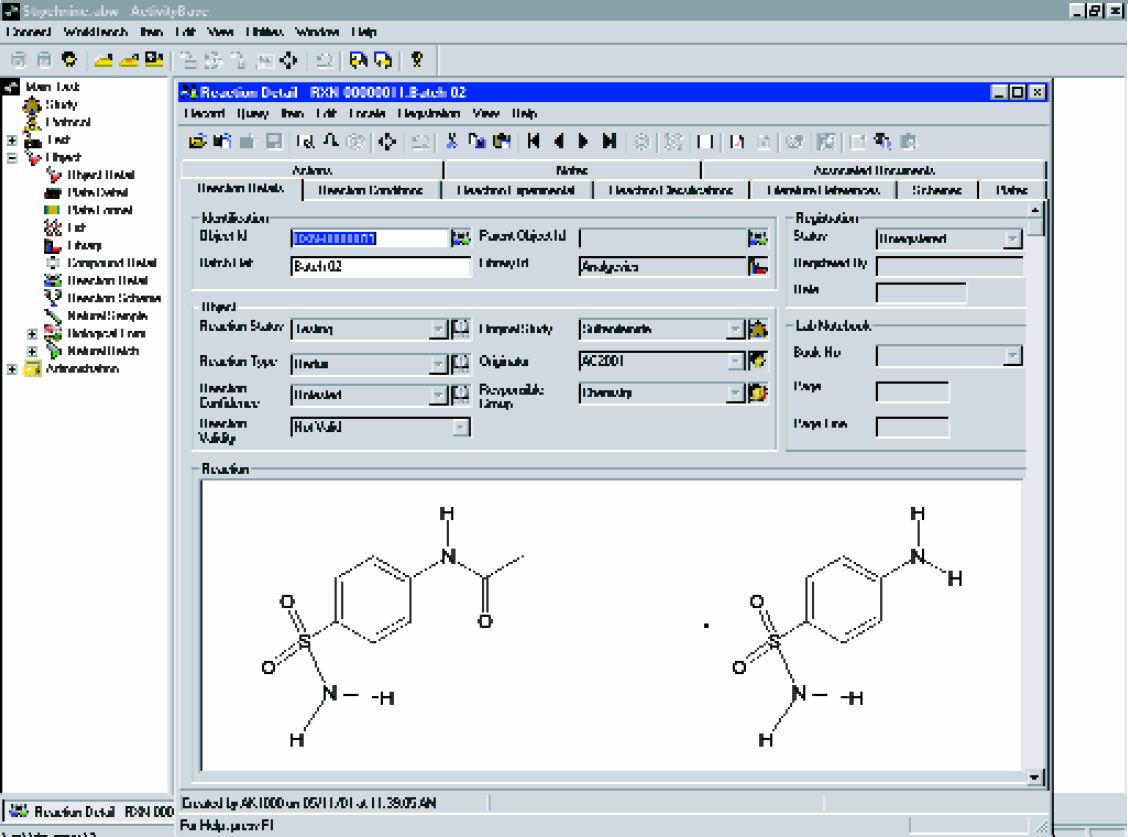
ReactionBase is a fully integrated solution to register, store, validate, search and retrieve reactions and reaction schemes.
AssayBase supports several plate pooling and corresponding result deconvolution schemes, including row collapse, column collapse and stack. Combining any two of these yields a two-dimensional pooling scheme, or users can select the built in two-dimensional orthogonal scheme.
The main advantage of an integrated data management system is that various types of information-biological and chemical, for example-are collected and stored in one system, using a common software platform. This improves the efficiency of data retrieval, manipulation, and analysis, and enhances search capabilities for compounds or chemical structures based on multiple desired characteristics. The ability to use integrated search tools to view and manipulate multiple datasets, such as chemical structure, screening results, and the output of analytical tests, facilitates data exchange within an organization and simplifies problem solving.
A seamless data management system benefits laboratory managers by improving laboratory workflow and process management. ActivityBase can generate a record of research activities, study protocols, experimental conditions, and analytical techniques and can assist in scheduling workflow. For example, using the Test Request facility, users can automatically assign compounds of interest for testing by colleagues in other groups. ActivityBase is developed using industry standard Oracle and Microsoft technologies. This allows information technology (IT) groups to configure and optimize the applications to meet the needs and match the workflow of a particular group within the company, or alternatively to integrate with other vendor solutions of their choice.
Each independent ActivityBase module brings its own functionality to the software suite. StructureBase provides for molecule registration, chemical searching, salt stripping, novelty checking, and validation rules. It automatically calculates the mass of a compound and generates possible formulas, including salts and solvates. Both the StructureBase and ReactionBase applications are based on Oracle cartridge technology.
Chemical search options include single or multiple structure queries with and/or logic, as well as substructure, superstructure, exact match, similarity, and full stereochemistry searches. Users are able to display structures as two- or three-dimensional wireframe, ball and stick, tube, or spacefilling models, to rotate molecule displays, and to “drag and drop” files around the ActivityBase desktop. StructureBase also provides links to industry standard drawing packages such as ChemDraw and ISISDraw.
StructureLoad, a novel supporting application enables batch loading of chemical structures into StructureBase and of reactions, reaction schemes, and related data (including mapping and transformation information) into ReactionBase. Users can search ReactionBase for reactions, compounds, reactants, and products, and are able to link, order, and track reactions within schemes. One of the application's functions is to manage reaction data as “testable objects”, which allows for measuring, tracking, and recording of reaction results. This information is particularly beneficial for catalyst design and optimization of reaction conditions.
In drug discovery, one of the most important factors in evaluating the therapeutic potential of an experimental compound is whether or not it binds to the target and the outcome of the binding event. Does the compound function as an agonist or antagonist for a target receptor in functional assays, does it play a catalytic or inhibitory role in a particular enzymatic reaction, or does it promote or block cell signaling in cell-based assays? Answering these types of questions and defining the characteristics of molecular structures that influence the sensitivity and specificity of a compound for a particular target requires the integration of chemical and biological data.
The merger and analysis of structural and screening data leads to the determination of a Structure-Activity Relationship (SAR). The SAR provides a critical piece of information for assessing the value of a chemical structure in drug discovery research. Structure searching is an essential component of SAR analysis. The integration of SARgen 5.0, a query engine for ActivityBase suite databases, enables structure searching and SAR determination. SARgen searches and retrieves data related to study properties, protocols, and results; it retrieves assay data and can search compound files for specific characteristics. Version 5.0 incorporates logical operators (and, or, not) for combining fields in complex queries. It also allows users to schedule automatic queries and report generation.
The SARgen suite of applications for data reporting and querying in ActivityBase includes SARgen 5.0, SARview, and ProfileView. The SARview viewer presents datasets retrieved by SARgen in an SAR report format that displays chemical structures together with their biological and chemical properties in a tabular format. (Figure 5) SARview guides the user through the report generation process, establishing settings that can be saved and applied to subsequent datasets and query results.

SARview from IDBS allows users to identify structure activity relationships in a dataset. Compounds of interest can be highlighted and structures filtered as required.
Refining the application of SAR data may require filtering through the many compounds that bind to a target and produce the desired activity in initial screening assays. The goal is to identify the structures that are most likely to make good drugs and that warrant more extensive analysis and further expenditure of resources. Analytical functions incorporated in SARview include “drug-like” filters (including Lipinski-type filters) to search datasets for compounds that have drug-like qualities, as well as filters that select for compounds having preferred structural motifs.
ProfileView (Figure 6) allows users to customize data presentation in report form. Scientists can present the same dataset in different formats, or combine various datasets and analyses, tailoring a report for a particular target audience. A sample report may include the structure of a compound presented alongside the results of tests performed on the compound, such as IC50 curves, spectra, or toxicology data. The software can also plot graphical results against condition qualifiers.

Profileview for use with the SARgen query tool from IDBS allows chemical and biological data for a compound to be viewed within a single report, enabling compounds to be “failed early.”
In summary, ActivityBase 5.0 enables the seamless integration of cheminformatic and bioinformatic data in discovery research. The software modules provide a common computational platform for capturing, managing, analyzing, and applying experimental data. This integrated framework facilitates exchange of biological and chemical information within an organization and bridges the gap between the research and IT environments, thereby maximizing the value of research data and improving overall productivity.
ABOUT THE AUTHOR
Jack Elands is Vice President of Marketing at IDBS. Drawing on his depth of experience in the biopharmaceutical industry, Jack leads IDBS' strategic marketing and business development. He is a recognized expert in the industry, with extensive experience in drug discovery including scientific research, project and product management, and leadership in sales and marketing.
Prior to joining IDBS, Jack was Head of Marketing and Sales at BioSignal Packard, Inc. Earlier in his career, Jack created a screening operation at Marion Merrell Dow and was responsible for the automation of all the company's research sites in addition to being a key player in their Bioinformatics Program. He also worked for Zymark Corporation as Business Unit Manager of Drug Discovery Products.
As a graduate of the University of Utrecht, Netherlands, Jack holds a doctorate degree in Pharmacology and Medical Biology. He has published over 50 scientific papers in journals including the American Journal of Physiology, European Journal of Pharmacology, Neuroendocrinology and Endocrinology. He is also co-author of two U.S. patents. Currently, Jack is a member of the Society of Biomolecular Screening and the Association of Laboratory Automation.
