Abstract

Microfluidic devices are made up of interconnected networks of microchannels and tiny volume reservoirs in which all the processes required for single analyses may be miniaturized, integrated, and automated within a single substrate the size of a human hand or smaller. However, LabCard devices are more than tiny replicates of existing equipment. They are characterized by high speed, parallel analysis using designs devised to eliminate sample cross contamination while automating processes that are otherwise undesirably cumbersome at the macro laboratory scale.
Electrophoresis is one of the most widely used analytical separation methodologies for life science research. Electrophoresis refers to the movement of a charged molecule under the influence of an electric field. Electrophoresis can be used to separate molecules that have different charge-to-mass ratios such as proteins; or, molecules that have similar charge-to-mass ratios but different masses such as DNA fragments. In recent years, the development of capillary electrophoresis (CE), which is performed in a fused silica capillary filled with a buffer or polymer solution, has increased the speed of analytical separations.
Planar, microchannel devices offer further improvements over capillary electrophoresis. In general, both electrophoresis and elec-troosmosis occur when a high electric field is applied along a microchannel (Figure 1). In practice, these effects can be comparable in magnitude, in which case ions of one charge move rapidly-at velocities of millimeters per second-while those of opposite charge move backward or slowly forward, depending upon which of the two effects dominates. The amount of fixed charge on the inner surface of a channel can be controlled by pH, specific adsorption of charged species onto the surface, or surface chemical modification. Therefore, the contribution of electroosmosis (the pumping mechanism) can be tuned relative to electrophoresis (the separating mechanism).
The capability to transport and separate exceptionally small liquid volumes with precise electrical control is a powerful tool, which is complemented by an additional benefit of using microchannels with micrometer cross-sections. While electrophoresis channels are of similar cross section in both capillaries and microchannel devices, the planar format of the microchannel devices enables structures more complex than a single, non-intersecting channel to be used. Two or more channels can intersect to form a tee, a cross, or other such structure. The ability to fabricate intersecting channels has led to sample injectors that introduce shorter, better-defined sample plugs into separation channels than those typically encountered in CE. Because shorter sample plugs can be injected, shorter separation lengths are required, and separations become faster without loss of resolution. Additionally, the ability to independently control the electrical potential on intersecting channels permits fluid streams to be metered quantitatively to create mixtures of desired composition. In essence, this electronically controlled metering mimics valve functionality.
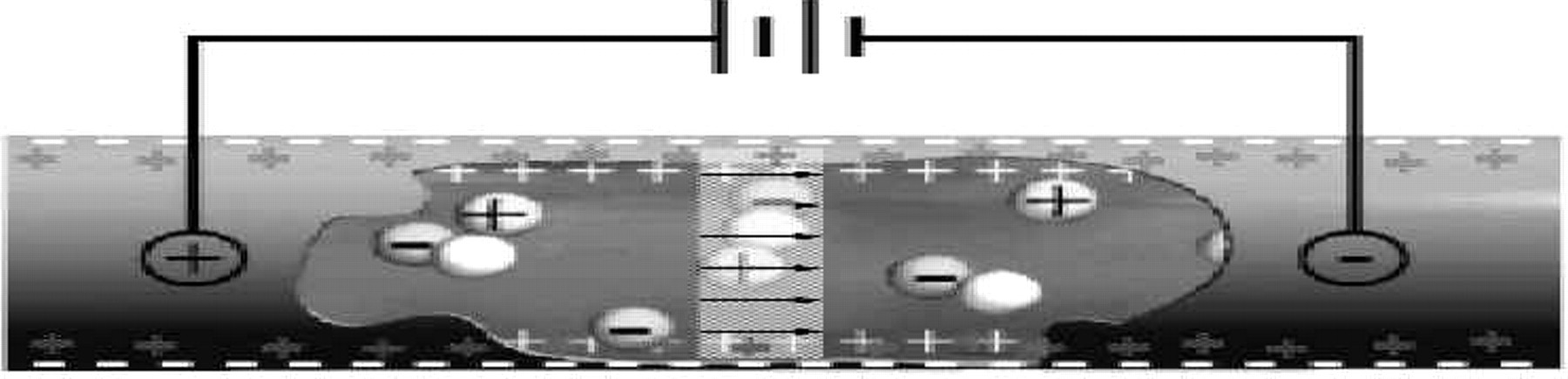
Microchannel cut-away showing electroosmotic flow in which a moving layer of counter ions (positive) attracted to the partial charge (negative) on the channel wall results in plug flow of a bolus of material in the capillary.
The combination of electrokinetic pumping, valving and separation within interconnected networks of microchannels on planar substrates was described in publications in the early to mid 90's by Soane 1 and groups at Ciba-Geigy, 2 – 5 the University of Alberta, 6,7 Oak Ridge National Laboratory, 8,9 the University of California at Berkeley, 10,11 and PerSeptive Biosystems. 12 Each of these groups implemented approaches to electrophoretically separate species in solution using microfabricated chip devices. In addition to separations, several groups explored “integrated” chip-CE, meaning that two or more analytical protocols (e.g., mixing, incubation, and separation) were included on a single, monolithic structure. 13 – 15
Though much of the work on microfluidic devices has utilized glass or silica substrates, Soane 1 and Ekström 16 proposed using polymeric materials in chip-CE as early as 1990. Soane focused on thermoplastics such as PMMA, while Ekström et al., investigated elastomeric polymers such as poly(dimethylsiloxane), PDMS. Recently, researchers at Novartis 15 have also demonstrated electrophoretic separations in PDMS microchannel devices.
ACLARA's LabCard devices are manufactured by injection and compression molding methods or by embossing, which contributes to making card-based, high-throughput analysis cost effective. Perhaps more importantly, these devices are designed to be compatible with existing laboratory automation. Many of the most developed of the initial microfluidic systems are focused on biotechnology applications in genetic analysis, pharmaceutical compound screening, and medical diagnosis. Here, we describe the promising technology of plastic microfluidic systems, which are sufficiently low in manufacturing cost to be both disposable and adaptable-in the form of highly customized designs for each application-to a wide range of clinically and academically interesting analytical applications.
MINIATURIZED PLASTIC ELECTROPHORESIS DEVICES
ACLARA BioSciences has applied polymer fabrication technology to manufacture miniaturized channel networks for electrophoresis in thin, plastic substrates. These plastic LabCard devices are replicated from a solid master (typically glass, silicon, or nickel) by processes, which include injection molding, compression molding, casting, and hot embossing. ACLARA BioSciences uses all of these processes to generate substrates with high precision patterns of micron-size channel features. The master is produced either by direct chemical etching of a material, or, in the case of nickel masters, is created by electroplating nickel from an aqueous solution directly onto an etched substrate. The master is an exact mirror image of the topography of the features (such as channels) in the finished device. The embossed or molded, open channel card is then sealed with a second, flat cover of plastic to enclose the embossed or molded features. The cover and embossed or molded base can be sealed by thermally fusing the two pieces together or by lamination using a layer of pressure sensitive adhesive (Figure 2). The cover may be of the same or different material and thickness as the base. A range of plastic materials may be molded and sealed using these general techniques. Here we report analytical results obtained primarily with devices molded in acrylic-based polymers.
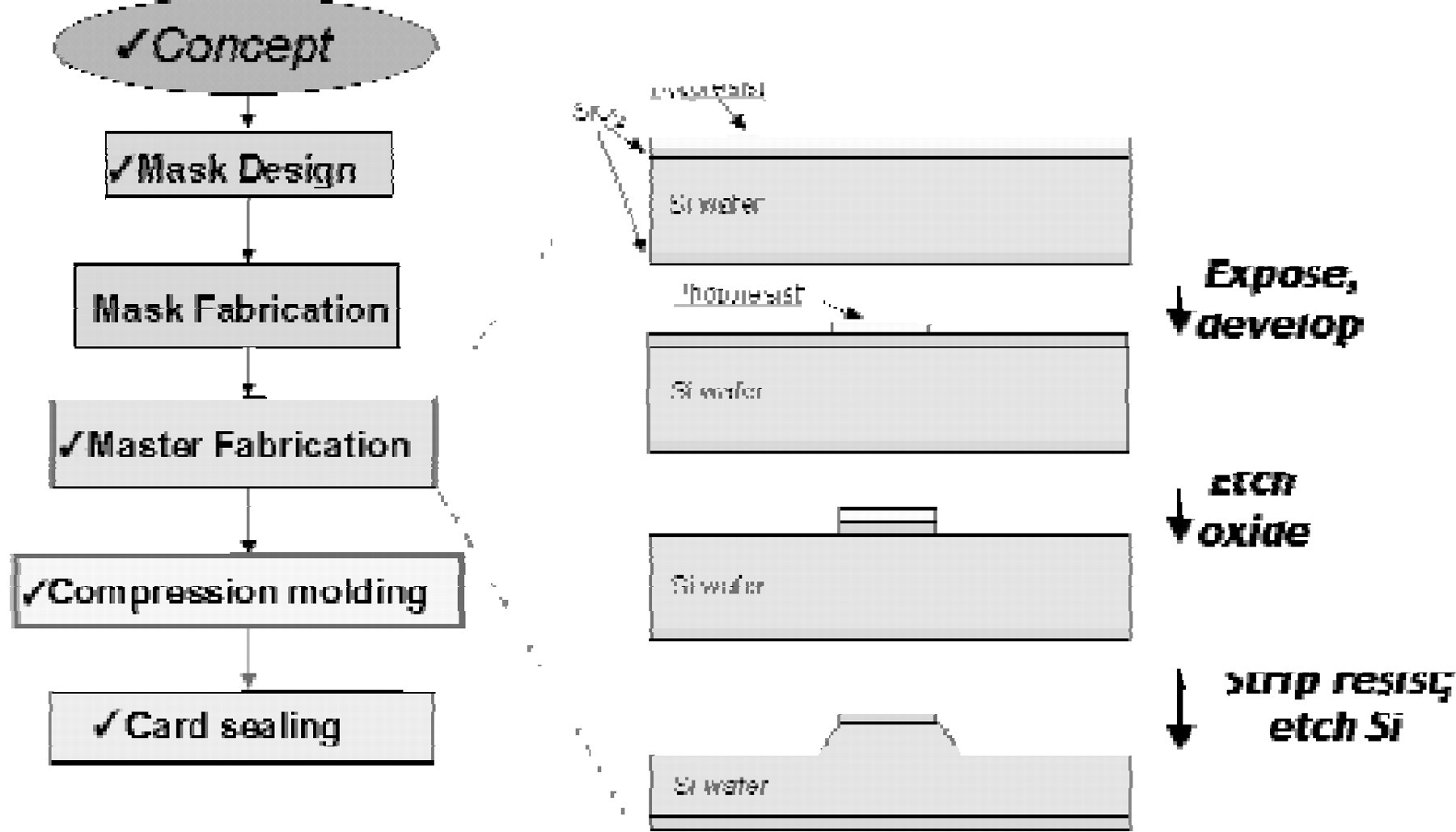
Fabrication process. A microfluidic design is patterned and etched on a glass or silicon substrate using standard micromachining techniques to form a positive master. Plastic parts with channeled structures are thermally formed by compression against the master mold tool. The molded card is then sealed with a plastic cover film enclosing the microchannels and through holes (not shown) that intersect them. Once sealed, these through holes form reagent reservoirs in fluidic contact with the microchannels.
DNA SEPARATIONS FOR GENETIC ANALYSIS
Electrophoresis has long been a fundamental technique used in nucleic acid analysis. This is exemplified in the genomic sequencing programs for the human genome and other model organism genomes, which rely primarily on electrophoretic separation of these enzymatically generated, fluorescently labeled DNA fragments. Separation and analysis of enzymatic sequencing reaction products identifies the sequence of nucleotide bases in a target DNA sample. Electrophoresis is also used for genotyping applications, including human identity testing and mutation detection, 17,18 as well as expression analysis, pathogen detection and pathogen characterization. 19,20 The market for DNA analysis is expected to continue to follow a steep growth curve, even with the completion of the Human Genome Project. This is due to increasing interest in pharmacogenetics, the science of using genetics to direct both the discovery of therapeutic agents and their rational prescription.
Rapid dsDNA fragment separations are achieved on microchannel devices by taking advantage of the narrow injection plug width. At short separation distances under high electric field strengths, diffusional broadening of analyte zones is small and the contribution of injection to the total peak width is significant. Under these conditions a reduction in injected plug size translates into improved peak separation efficiencies. This is compatible with short separation channels, including arrays of channel patterns, which are easily fabricated on microfluidic devices and allow peak detection at distances closer to the point of injection than in conventional CE systems. Rapid separation of DNA fragments from a standard ØX174 phagemid DNA endonuclease digestion has been successfully performed in a plastic LabCard device (Figure 3A). Using standard electrokinetic injection methods nearly baseline resolution was achieved for all fragments in less than one minute.
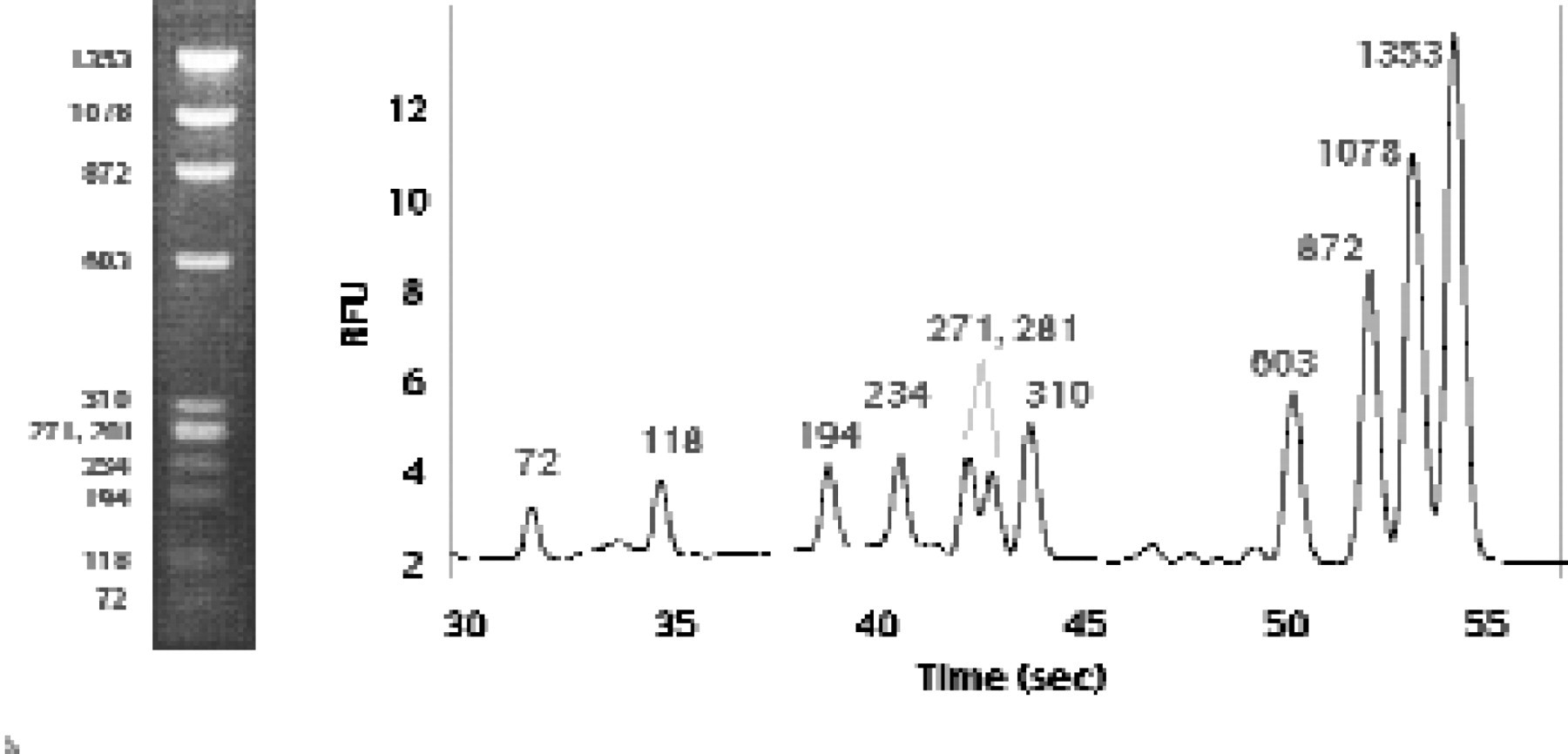
A separation of double-stranded, phagemid ØX 174 DNA fragments generated by HaeIII restriction endonuclease digestion. The polymer used for the separation is a 2% (w/v) uncrosslinked linear polymer and the fragments were detected 2.5 cm from the point of injection. At the left is an agarose gel separation of the same digest, which takes about 30 minutes to perform.
Because nucleic acid fragments have a charge-to-mass ratio essentially independent of fragment length, nucleic acid separations are typically performed in a solution of entangled polymers that act as a sieve to separate the fragments by size. Figure 3B shows an example of the rapid DNA size separation in a microchannel device. The electropherogram shows a common DNA sizing standard, consisting of a “ladder” of fragments ranging from 100 to 1000 base pairs, each differing in length by 100 base pairs. To detect the fragments, an intercalating dye was included in the polymer sieving media. When this dye intercalates into the DNA structure, it fluoresces brightly in response to laser excitation. The fragments were separated in less than half a minute over a separation distance of less than two millimeters. In a typical commercial CE or slab gel electrophoresis device, this separation would require at least half an hour. Double-stranded DNA fragments differing in length by 1.3% can be resolved almost to baseline in longer channels in less than six minutes.
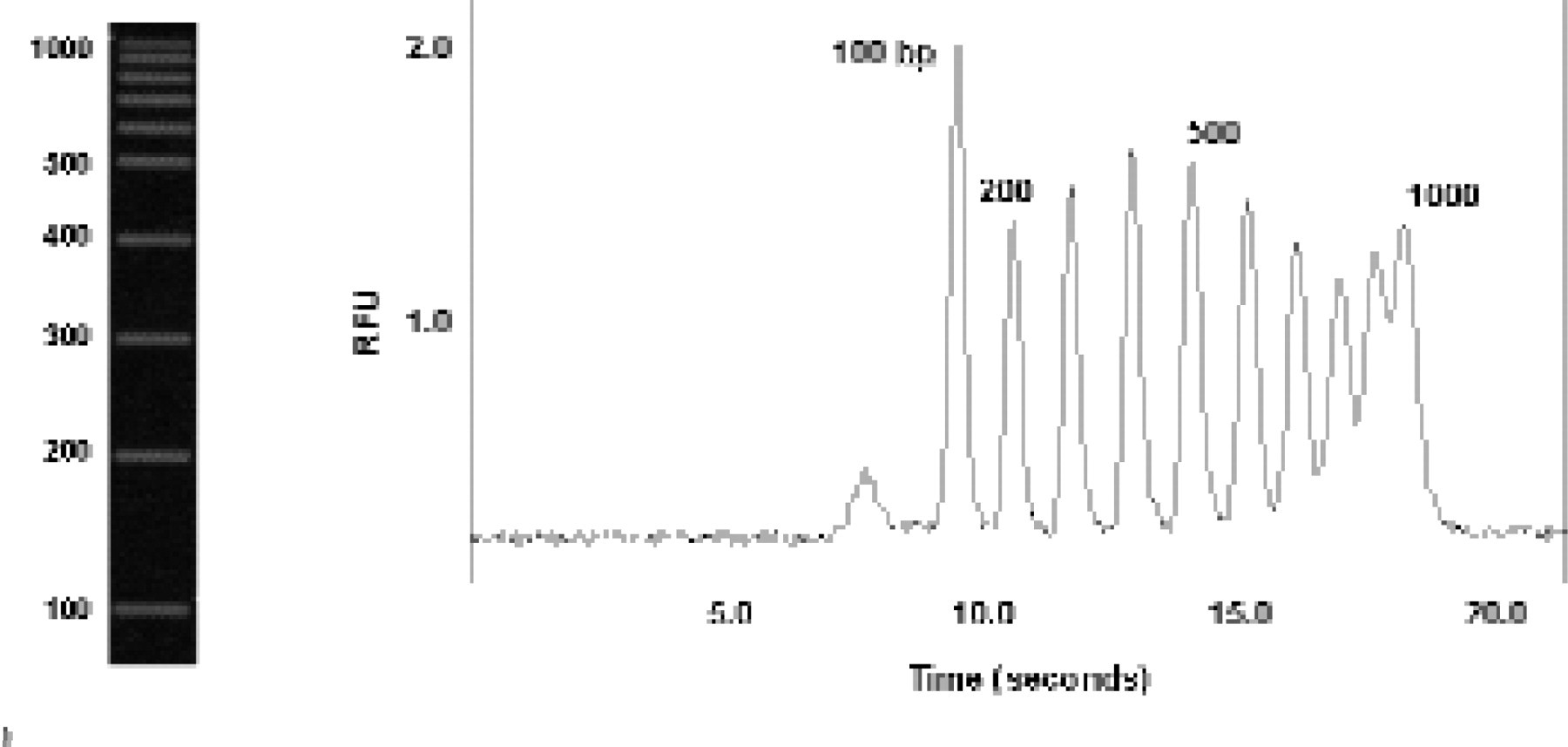
Rapid separation of fluorescein-labelled, 100 to 1000 bp, double-stranded DNA sizing ladder in a plastic microchannel. Electrophoresis was performed in a solution of 2% uncrosslinked polymer with an effective separation distance of approximately 1.7 mm. At the left is an agarose gel separation of the same DNA ladder, which takes about 30 minutes to perform.
Human identity testing is another application that requires double-stranded DNA separation that has already been adapted from traditional slab gel electrophoresis to capillary analysis 21 . Multichannel, plastic LabCard devices have been used successfully to demonstrate the feasibility of performing multiplexed, high throughput genotyping of short tandem repeat (STR) microsatellite alleles. D1S80, one such microsatellite locus, has been genotyped in a plastic multi-channel LabCard device made up of an array of 16 microchannels 22 . Alleles from the D1S80 locus range in size from 369 to more than 801 base pairs, and may differ in length by approximately 4% or less. The LabCard chips used in this study were from a batch of approximately 1000 produced by injection molding from a nickel electroform, then subsequently sealed by lamination with an acrylic film as described above.
Plastic microchannel devices have also been used to perform DNA sequencing separations. For this application, a collection of fragments that differ in length by one nucleotide base (to as little as one base in 1000) must be resolved so that the sequence of bases can be interpreted from the electropherogram. The maximum number of bases that can be assigned with confidence is referred to as the sequence read length. Sequencing read lengths in excess of 600 bases have been achieved with >98% accuracy in only 30 minutes using plastic LabCard devices (Figure 4). A similar level of performance would require two or three hours in commercial, automated slab gel or capillary sequencers.
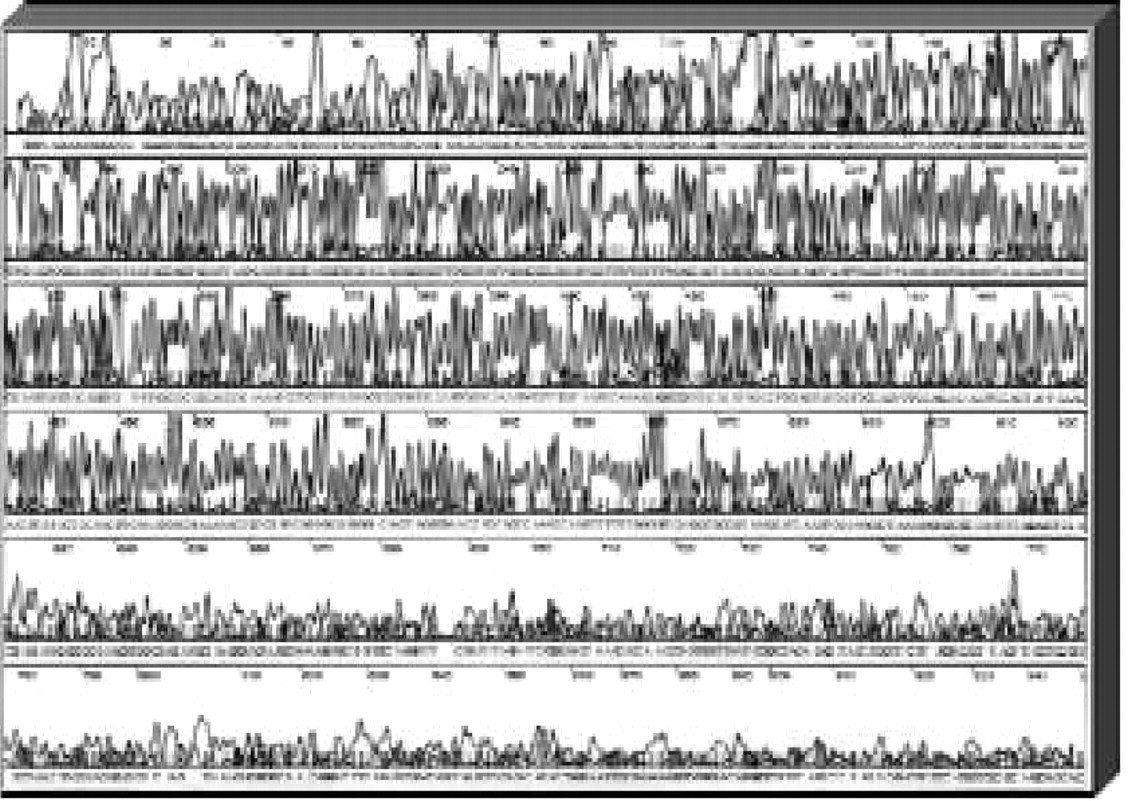
A four-color, electrophoretic separation of a DNA sequencing reaction generated from an M13mp18 template (a common cloning vector) using BigDyeb-labeled primers (Applied Biosystems, Inc., Foster City, CA); the total electrophoresis time is approximately 40 minutes and base-calling accuracy is 98% up to 640 bases.
LabCard devices offer increased analytical throughput over traditional capillary electrophoresis by providing arrays of channel patterns on each substrate, each of which is capable of rapid electrophoretic separation. However, another order of magnitude increase in throughput can be gained by multiplexing some types of analyses in each channel. This has been accomplished in such nucleic acid analysis applications as genotyping and expression profiling. Specifically, the sequence of interest is detected by hybridization with a complementary, sequence-specific, recognition probe labeled at the 5′ end using a unique class of CE reporter molecule called an eTag™ reporter. These eTag reporters are fluorescent dyes covalently modified to produce unique mobility signatures under defined electrophoretic separation conditions. These reporter molecules are enzymatically released from hybridized recognition probes during biochemical enzyme assays such as the TaqMan® or Invader™ assays. The eTag reporters released during one of these multiplexed assays can be electrophoretically separated and decoded in a single LabCard microchannel. This chemical reporter system lends itself to generating large families of generic reporters, which are ideally suited to capillary electrophoretic analysis. The additional power of multiplex assay analysis compounds the advantages of an already flexible and powerful analytical system.
ELECTROPHORETIC-BASED ENZYME ASSAYS FOR HIGH THROUGHPUT SCREENING
The ultimate goal for pharmaceutical companies is to develop a consistent pipeline of new lead compounds for drug development. With the advent of combinatorial chemistry, an exponentially increasing number of test compounds are becoming available to screen as drug candidates, while advances in genomics have broadened the variety of candidate therapeutic targets available for screening. As a result, there is pent-up demand to perform an exponentially increasing number of screening assays. However, even a milliliter of expensive or hard-to-obtain target or test compound can dominate the cost of an assay. Therefore, it is critical to limit the reagent volumes required per assay so that libraries of pharmaceutical compounds can be screened against as many targets as possible. The goal of high throughput drug screening (HTS) is to optimize the efficiency of these screens and maximize the number of likely candidate compounds emerging from screening for potential therapeutic development.
A common method for screening test compounds involves mixing an enzyme solution with its fluorogenic substrate along with a candidate inhibitor compound. After a fixed incubation period, the mixture is read directly on a plate reader to detect substrate fluorescence. This value is used to quantitate the amount of product formed by the enzyme and when compared to the baseline product formation in the absence of test compounds shows whether the test compound alters enzyme activity. Such homogeneous assays are typically performed in approximately 200 μL volumes in standard 96-well plates. Utilizing modern robotic systems, up to one hundred thousand test compounds may be screened in a single day, which in this assay format would consume, in total, 20 liters of reagents.
ACLARA has developed microfluidic formats that address the HTS challenge to achieve more efficient assay formats while still adapting to a diverse range of assay types. One specific microfluidic solution offers reagent conservation by direct reduction in the minimum reaction volume required to perform a homogeneous, mix-and-read assay. Homogeneous assays are typically run in a minimum of 5 μL since volumes less than this are inadequate to withstand evaporative effects during assay incubation and therefore result in unreliable assay results. ACLARA has broken this evaporation barrier to homogeneous assay volume reduction by developing a unique, microfluidic LabCard design that provides intrinsic evaporation-control. The unique channel pattern that interfaces with the assay incubation reservoir in this device permits assay volumes to be reliably reduced from 200 μL to <300 nL.
In a second microfluidic design strategy, ACLARA is integrating assay incubation reservoirs directly with electrophoretic separation channels on single LabCard substrates. This provides an opportunity to miniaturize heterogeneous assay formats, which depend on assay product separation to analyze the output. Assays for kinase, phosphatase, and protease-catalyzed enzyme reactions performed over a wide range of pH and assay conditions have been demonstrated using both of these LabCard designs. Incubations typically occur in 30 minutes or less and analytical separations are generally complete in seconds. This is possible because the reaction products usually have significantly altered charge-to-mass ratios relative to the assay substrates and separate readily in high voltage fields in short separation channels. By combining fluidic metering with separation capability, which is easily accomplished in these microfluidic systems; it is also possible to integrate assay mixing, reading and component separations to monitor the progress of a heterogeneous assay reaction in real time. Furthermore, by performing electrophoretic separations in parallel on LabCard devices, enzyme inhibition data as a function of concentration can be collected in parallel on a single device, providing dose-response curves in approximately the same time it would normally take to perform a single assay (Figure 5). The electropherograms shown here were obtained using a plastic microfluidic system to monitor both a kinase-catalyzed reaction substrate and its product. The first peak in the electropherograms shown in this figure is an internal standard used to account for possible variations in channel dimensions or the performance of the injection process. Product, substrate, and internal control were separated in less than 7.5 sec. In general, microfluidics dramatically reduces assay volumes, which may be orders of magnitude less than those used in conventional microtitre plates.
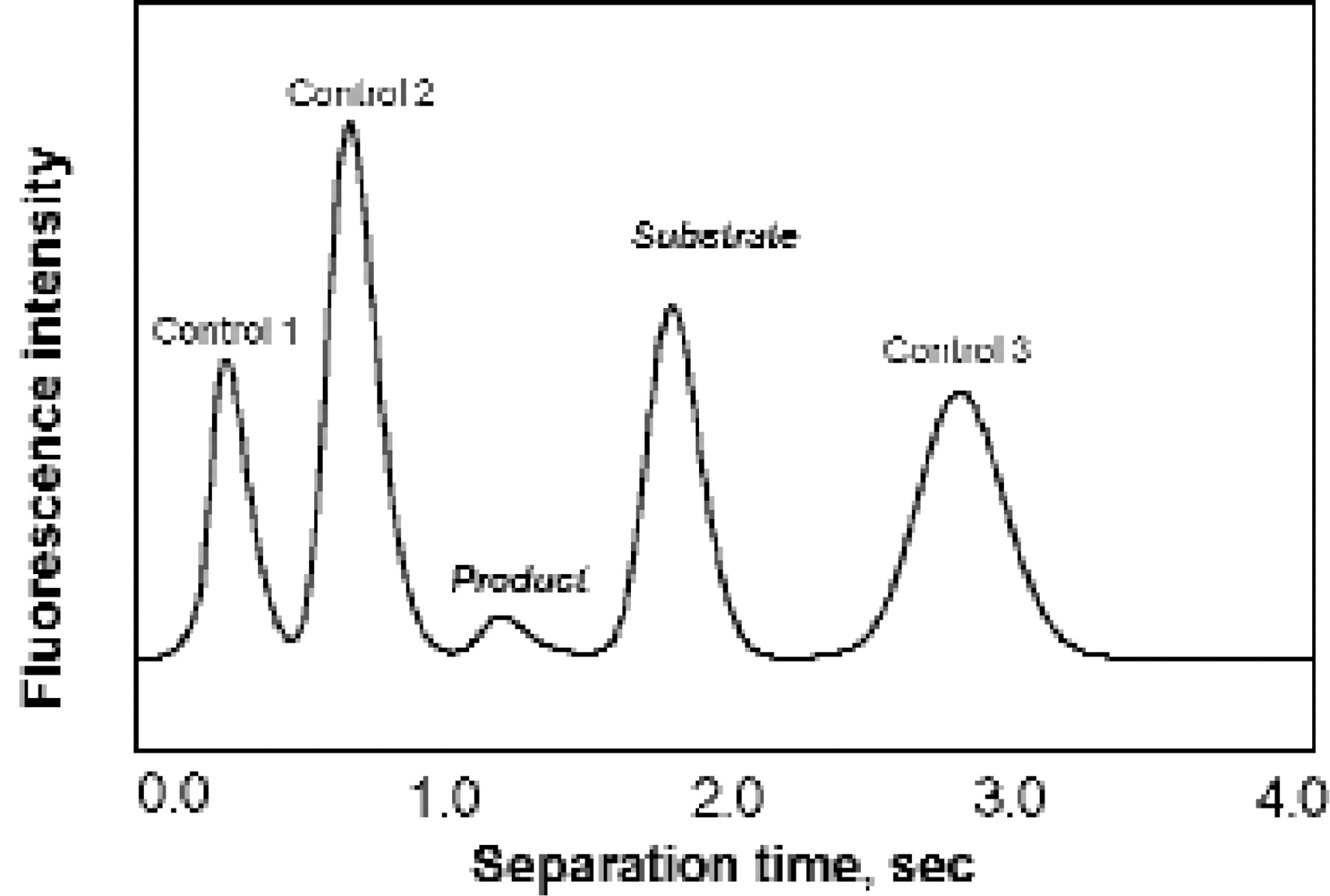
Typical enzyme inhibition assay, showing substrate and product peaks relative to internal assay and migration standards. Typical electrophoresis conditions are 300 V/cm in 1% uncrosslinked polymer, 25 mM HEPES buffer using a separation distance of approximately 2 mm.
CONCLUSION
The microfluidic devices described here, combined with analytical separations and complimentary assay formats, provide a few examples of the rapid, high throughput assays and electrophoretic separations made possible in plastic, microchannel devices. Applications in only two out of many potential application fields-genetic analysis and high throughput pharmaceutical compound screening were discussed. In the years to come, each of the analytical processes from sample collection through final analysis will become more fully integrated into interfacing structures on LabCard devices, allowing the complete realization of sample-to-answer devices, the true “labs-on-a-chip”.
ACKNOWLEDGMENTS
We would like to thank our many colleagues at ACLARA BioSciences in the MicroFabrication and Materials, New Technologies, Integrated Devices and Genomics Applications groups for their work on microfabrication, LabCard device molding, organic chemistry and analytical separations.
The authors are members of the core research and development teams at ACLARA BioSciences, Inc., 1288 Pear Avenue, Mountain View, CA 94043, tel: (650) 210–1200, fax: (650) 210–1244, e-mail:
