Abstract
Our objective is to eliminate the digital divide that persists between the physical and information spaces of wet-lab based enterprises by embedding computational resources into the shared laboratory environment. Our first challenge is to enable individual lab workers to contribute to a fine-grained formal representation of ongoing lab activities — to build the database by doing the work, without having to stop and write things down in a notebook or to enter information into a computer. By eliminating the redundancy of doing the work and then recording it, accuracy and completeness will be improved. And, by capturing information at a finer detail than is practical for manual entry systems, unanticipated applications can be supported.
INTRODUCTION
For computing to play an even greater role in the advancement of molecular biology than it already does today, easier and better access to computing must be made available within the laboratory environment. Some of the application areas that can benefit from integration of the physical and information spaces of the biology lab include:
scientific applications such as data mining and analysis, experiment design, and modeling and simulation;
collaboration between individuals and groups;
quality control and inventory management in production and research environments;
automation and integration of low and high throughput experimentation; and
record keeping for legal and attribution purposes.
While computing is applied to some degree in all of these areas today, our research indicates that loss of information is a major limiting factor. Much of the data produced by measurement devices in the laboratory are abstract and disconnected from the history of the corresponding physical samples, so the meaning of the data (image or graph, e.g.) cannot be directly understood by people or by machines. Instead, potential consumers of the experimental results must turn to a specific individual to supply essential background information. We refer to this background information as experimental meta-data. Experimental meta-data is essential for all of the applications listed above, yet the perceived value of metadata tends to be lower than the cost associated with documentation and dissemination.
The barrier to wider utilization of information technology in the biology laboratory can be reduced to two specific problems related to meta-data: the lack of a formal, portable representation for experimental meta-data; and the lack of easy ways for people to update and query the meta-data along with their measured results. We contend that these problems cannot be solved by improved desktop applications or interfaces alone, rather, data gathering and first order interpretation must be a direct result of the physical activity taking place in the laboratory environment combined with input from the laboratory worker. Our objective is to enable new applications through meta-data capture in the laboratory environment, while decreasing the overall time and attention that lab workers must devote to direct interaction with computers in the conventional sense.
Though the formal meta-data representation itself is not our primary research focus, we recognize the importance of this aspect of the problem, and the difficulties associated with ontologies for disciplines as broad as molecular biology. To this end, we are cooperating in the development of a representation called MetaGraph 8 that, among other things, can express the primitive physical operations that occur in the wet-lab environment.
BACK GROUND
Desktop and client-server applications called Electronic Lab Notebooks (ELNs) are available today that allow biologists to enter experimental information into a database before or after protocols are performed as described in research by Myers 1 and Lysakowski 2 in respectively. While ELNs have the potential to overcome the deadend nature of paper based laboratory notebooks, 3 they are less portable, and still suffer from the other fundamental weakness of paper systems — the overhead in time and attention required to use them effectively stemming in part from lack of integration into the physical environment. Our approach complements existing work in ELNs through development and deployment of ubiquitous computing technologies in a unique experiment capture system. Rather than striving for a paperless laboratory in which notebooks are replaced by displays and keyboards, we are striving for an apparently computer-less laboratory that enables people to build the database by doing the work while placing a wealth of sophisticated information services at their disposal.
In the emerging field of post-PC era computing that is sometimes referred to as “ubiquitous”, “pervasive”, or “invisible” computing 4,5 activities of large numbers of heterogeneous devices, dispersed throughout an environment, are easily coordinated to deliver powerful information services to people. The objective of ubiquitous computing research is to provide these capabilities in a general way so that they can be readily applied to a variety of domains. We are in the process of building and evaluating a smart biology laboratory environment, called Labscape, for the Cell Systems Initiative (CSI) 6 based on ubiquitous computing technologies that are emerging from the Portolano Expedition into Invisible Computing. 7 There are two main research thrusts within the Labscape project:
User Experience
To discover a paradigm for computer-human interaction that is appropriate for experiment capture and information access in a smart laboratory environment.
Infrastructure
To apply and invent general networking and computing technologies that are capable of delivering the user experience defined above in real laboratory environments, which require high levels of reliability, scalability, security, and flexibility.
CASE STUDIES
The development process that we have applied to this project began with simple observation of laboratory work. In our observations to date, we witnessed several specific instances of lost meta-data that could have been applied directly to one or another of the target applications that we have identified. Two of those examples are presented here to motivate the proposed model of interaction that we have implemented in Labscape.
This first example addresses the notion of failure in the biology laboratory. The work observed was in the context of ligand-receptor assay development for high throughput screening. In this instance, the experimenter ran into difficulty obtaining a specific, proportional response using fluorescent tags as indicators. Ultimately, the fluorescent tagging approach was abandoned in favor of other means. The time spent on the unproductive approach was termed a failure by the biologist. This is a classical negative result that would normally not warrant much beyond a comment or two at a weekly research group meeting. However, it is not the outcome of the experiment that makes it a failure, rather it is the loss, to the larger community, of the knowledge that was created-someone in the future may go down this same path, as someone in the past may already have. The ability to query a database about other experimenter's experiences with the molecules in question might have saved our subject a week of work, or it may have provided clues about persistent problems with certain types of fluorescent tags.
Our second example involved direct collaboration between researchers. In a series of visits to an immunology laboratory, we observed how T-cells are grown in mice treated in different ways, and then harvested through a careful multi-step process. During one of our observations sessions, our subject offered extra cells obtained during the purification process to a colleague working in the same lab. Little historical information was exchanged at the time of hand-off. We discovered the potential value of this lost information on a subsequent visit to the same laboratory when the receiver of the cells returned to ask questions about how those particular mice had been treated. The convenience of the lab workers notwithstanding, it would be even more useful if a computer could access the joint history of these separated-at-birth samples for multi-dimensional data mining and model testing on the results generated in the divergent experimental paths. Much of this analysis could take place without the direct involvement of the experimenters as long as the measured data and corresponding meta-data are recorded and linked.
USE MODEL
The human computer interaction paradigm that we have implemented in our experiment capture prototype is based primarily on observation and interviews. Rather than viewing lab work as a set of experiments with discrete beginnings and ends, our system is based on the concept of sessions during which local names for objects can be defined and linked to global names for the same. In a session, samples of material held in containers are subject to a set of partially ordered primitive operations such as liquid transfer, environmental incubation, centrifuge, agitation, and separation by density gradient, phase or other means. The interlinked structure of the captured meta-data directly reflects the material flow between sessions. It is up to the lab worker to define the beginnings and ends of sessions primarily for the purpose of defining the scope of local names. Events involving locally named containers and various lab devices are combined with utterances from the lab worker to construct a formal representation of experimental procedures in MetaGraph. For new containers to survive the current session they must be named and scanned by the lab worker, otherwise no scanning is needed. Finally, measurements taken by various lab instruments or even by inspection would then be linked as data files to the MetaGraph representation. As a result, it would no longer be necessary to know or remember the names or locations of data files since they can be accessed in terms of the context in which they were created.
In our system, an experiment is a higher-level concept that can be defined as a view or query into the MetaGraph database produced through a series of sessions involving one or more experimenters. The history of a specific sample is yet another view, or query, into the same database. Such a database would instantly address a variety of applications that can be implemented as queries. For example, all experiments involving a poorly made solution or mis-calibrated device could be immediately identified. Personal attribution of entries in the database could have far-reaching, mostly positive effects, though some negative uses can be imaged as well.
Speech plays an important role in Labscape, but our approach is to de-emphasize speech recognition in favor of obtaining coherent input from all of the devices in the laboratory up to and including the manual pipetters that perform many of the primitive liquid handling operations. The role of speech is to fill in the gaps and to provide redundancy and feedback for error correction or disambiguation. As an example, when performing a pipette operation, we should be able to obtain accurate volumetric data from the device, while the names of the source and destinations of the liquid transfer may require direct input from the lab worker via voice or other means.
AN EXAMPLE
The following session, whose goal is to produce dilutions of two reagents in a single 96-well plate, provides a concrete example of the use model described above.
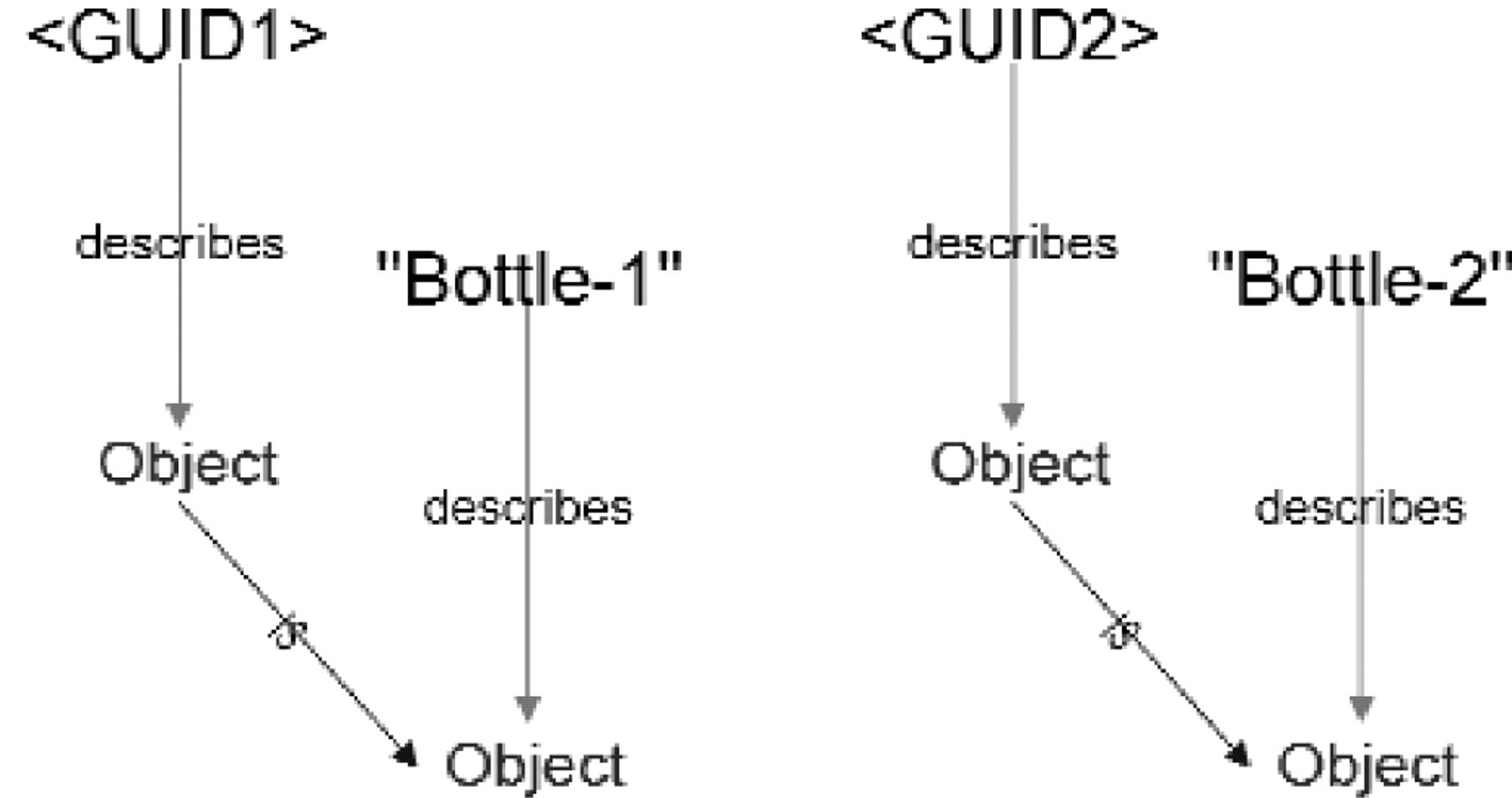
Additions to the MetaGraph model after Step 1
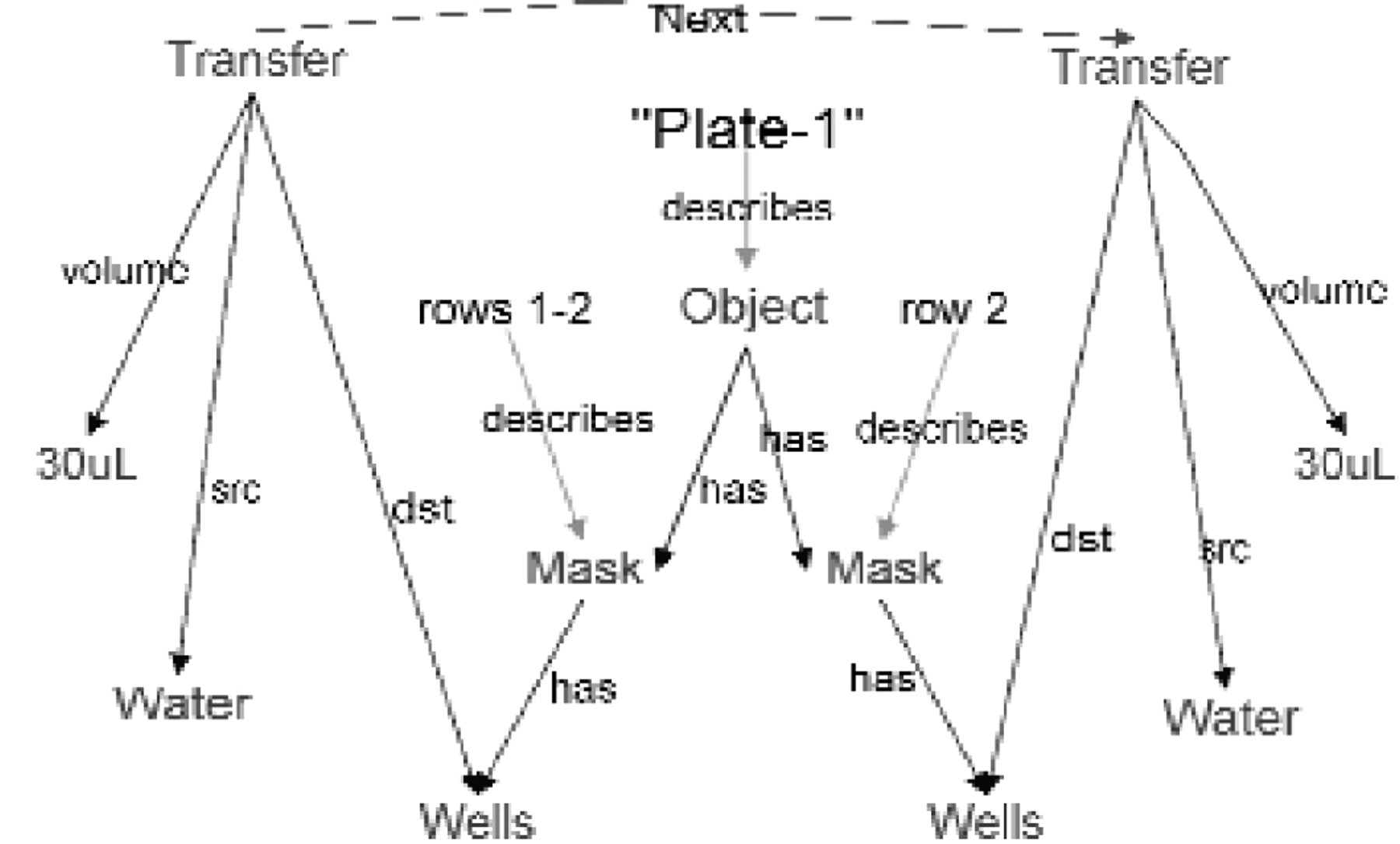
Additions to the MetaGraph model after Step 2
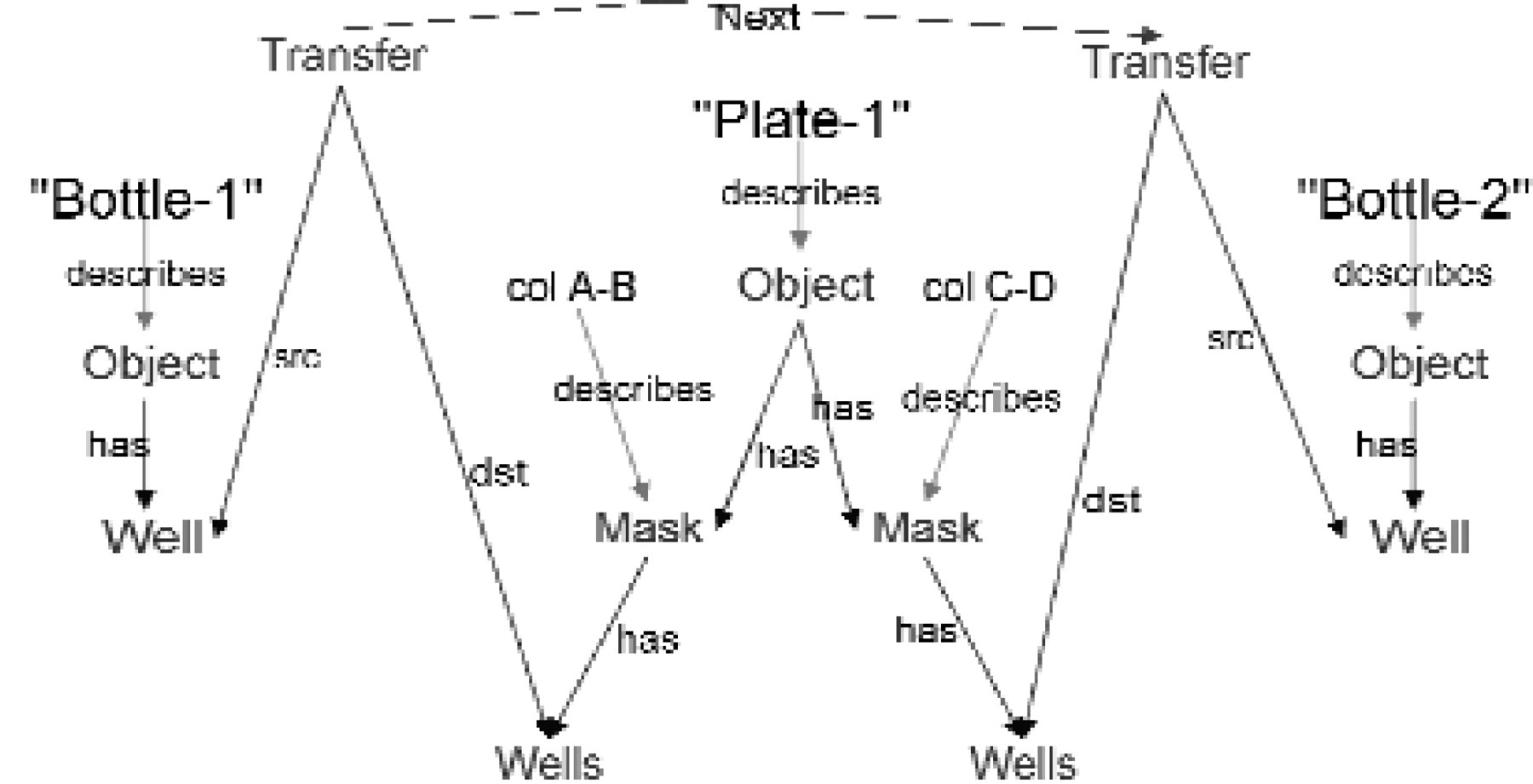
Additions to the MetaGraph model after Step 3
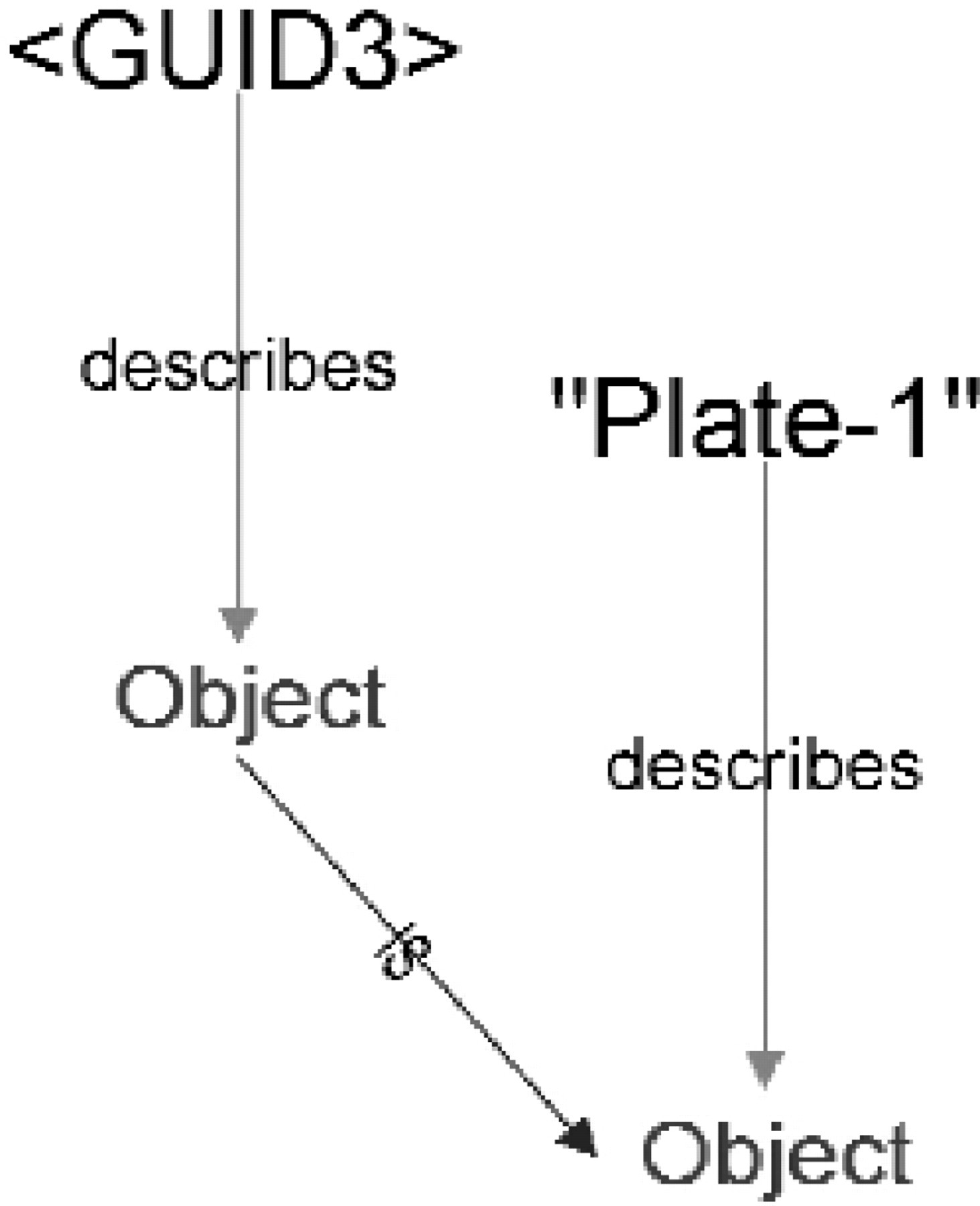
Additions to the MetaGraph model after Step 4
This scenario illustrates how speech is used to fill in the gaps that cannot be obtained from devices used in the lab. For instance, the pipetter should be able to report accurate volumes, though it is not likely to know the sources or destinations for the liquid transfer events. In the case of a device like a centrifuge that is capable of reporting the details of its use, speech may not be necessary at all, especially if it can scan the containers.
Had such a system been in use for the two case studies presented above significant benefits in time and quality could have been realized. Our challenge is to prove that the system can work reliably and simply enough to encourage complete and accurate capture of meta-data. To meet this challenge a variety of software and hardware infrastructure issues must be addressed, such as how events from widely distributed sensors (scanners, speech recognizers, pipetters, etc) involving a large number of workers can be converted into coherent meta-data representation, while giving the right feedback to each worker for validation and error correction.
INFRASTRUCTURE
The lab environment exhibits many characteristics that make it an ideal vehicle for evaluating and extending emerging technologies for ubiquitous computing. One of these characteristics is a high degree of device and personal mobility that stem from the task specific work areas commonly found in the lab environment. In a suitably enabled laboratory, one should expect a nearly constant reconfiguration of the environment through the addition of new devices, new people, and new software services. It is this characteristic that is particularly difficult to manage with traditional user oriented desktop computing technology. Another important characteristic that distinguishes ubiquitous computing from the desktop computing paradigm is the need to obtain coherent behavior from a collection of heterogeneous autonomous devices. The alternative practiced widely today is to enslave each device to a specific computer running an all-knowing application that is capable of coordinating the activities of those devices. This approach necessarily leads to brittle systems that cannot be easily reconfigured by the people in the lab.
Our approach is to decompose the big problem of building the MetaGraph database into a set of smaller services that can be defined and implemented independently. We then provide a means, called “worklists”, for composing those services on an event-by-event basis if necessary to produce the desired high-level functionality along with high degree of flexibility. A worklist is a lightweight, mobile list of service requests combined with event specific state that travels through the network finding services to process the requests. Conversely, a service is a relatively static program that manages a persistent state (databases) which can consume and generate worklists or process individual requests. As an example, a local network administrator can specify how local electronic pipetter events are handled by defining the following work-lists of service requests:
For any event, determine the event type and find the associated worklist specification (port listener service)
Send the raw event to the raw event log (event logging service)
Initiate (instantiate) a worklist that has been specified for this type of event (device proxy service)
Convert the raw data from the pipette into an abstract liquid handling event using the local XML schema (pipette translator service).
Find the ID of the worker associated with this event (user association service)
Send the converted event information to user's applications (user proxy service)
Notice that the second worklist isn't really complete with respect to the processing of the pipetter event. At the time this worklist was specified by the system integrator, it may not have been known which user level services might be interested in this data. Maybe there was no StepEditor service available at that time! The role of the user's proxy service is to pass events on to user applications that have registered interest in certain event types. In this case, each new StepEditor instance (mine, yours, etc) must register interest in abstract liquid transfer events with the corresponding user's registry service (via other worklists). This Service/Worklist architecture exhibits the following properties that are important for this and other ubiquitous computing environments:
Device data and behavior is not controlled only by software running on the locally connected machine. Instead, a device's role is a function of the context in which the device is used (by whom, on what containers, etc). This means that devices can be moved and installed anywhere without further reconfiguration of the software environment. The Port Listener Service can accept events from any device and produce the appropriate bootstrap worklist shown above.
Functionality is not tied to service configuration. For example, the system will function properly whether or not the user association service is running on a wireless badge or a wired server, though the former may be quicker and more efficient. Automatic or assisted network administration tools can reconfigure services to optimize performance independent of function. We rely on service discovery 10 to locate service instances that can handler specific request types.
There is a clear separation of expertise between services that do not need to know anything about each other. As an example, the StepEditor must understand the semantics of abstract liquid transfer between containers expressed as XML, yet it need know nothing about specific devices such as pipetters from a variety of manufacturers, or how to find users for events. By providing very late binding of service instances to service requests via service discovery, we can always make sure to get the best and latest functionality, including the capabilities of new devices as they are added to the environment and new policies as they are established by local administrators or users (by installing new worklist specifications into the device registry service).
The service/worklist decomposition allows incremental addition of functionality to the network. For example a “lab solution” provider might add a pipetter worklist specification to a basic set of services and worklists already provided by a general IT company to support domain independent capabilities such as event logging and user association. Meanwhile, the local system administrator might alter existing worklists to change the policies that determine how events are associated with users.
Though it would be possible to engineer a single special work area to provide an interesting user experience on an experimental basis, our approach is to couple advances in the user interface to advances in the computing infrastructure that will be needed to deploy our system into working laboratories. We are evaluating the service/worklist decomposition as a way to achieve the flexibility of purely event driven (reactive) systems without losing the control, programmability and security associated with desktop applications.
The operations that we support in our current implementation include:
Basic liquid handling: transfer, remove, add
Separation: supernatant, pellet, density gradient layers
Containers: Single well containers (tubes, bottles, dishes), one and two dimensional plates
Other operations: Spin, Agitate, Incubate (time, temperature, other ambient conditions)
Readers and measurement devices: None integrated at this time.
Our first deployed systems are input only, not requiring error correction from the user. We are using our experience with this system to identify any additional primitive operations that must be supported and to design a fully interactive interface. Future deployments will provide appropriate user interfaces for error correction, and will expand our support for meta-data to include ambient conditions of the lab; capture and indexing of audio, image, and video data; and integration with a larger set of laboratory devices such as centrifuge machines, incubators and a variety of readers. An important aspect of the user experience that we expect to address in the future is the need for local extensibility and customizability in the primitive operations and in the formal representations.
SUMMARY
The PC-bound information technologies found in most laboratory environments today limit the complexity of the biological questions that can be investigated to those that can be scoped and comprehended by a single researcher, or by a small tightly managed team. Specifically, we are concerned about the barriers that exist between a researcher and his or her peers, students, and mentors, with regard to the tremendous loss of valuable information generated in the lab. The information is lost because the opportunity cost of capture, dissemination and application of the knowledge is less than its perceived value. We believe that we can alter both the supply side and demand side economics of meta-data utilization by deploying ubiquitous computing technologies into the laboratory environment. Biology laboratories house people engaged in a complex but well-structured goal oriented activity, that must simultaneously engage in mental and physical activity. Thus, the biology laboratory is an excellent crucible for forging post-PC computing technologies. If we can make it (work) here, we can make it (work) anywhere! Preliminary results and feedback from our colleagues and patient experimental subjects encourages us to continue down this path.
ACKNOWLEDGEMENTS
Special thanks to Tom Kowski of Immunex, and Qinghong Zhou and Eduardo Firpo of CSI. This research was funded in part by DARPA Contract Number N66001-99-2-8924 “The Portolano Expedition into Invisible Computing”.
