Abstract
To provide biological specimens for scientific studies, the Medical Automation Research Center (MARC) designed and constructed a large-scale device that emphasizes the use of robotics and automation to integrate many associated laboratory operations. These included analysis, dilution, archival storage, and retrieval of purified human-derived specimens. Designers of automated biological repositories are challenged by complex engineering problems. In this paper, we present an overview of the biological repository (biorepository) and give details of the software architecture.
INTRODUCTION
One consequence of completing the human genome project will be a corresponding need for purified human genomic DNA from a characterized population of subjects as starting material for scientific studies [1,2,3]. While investigators can acquire this material by several means, an attractive source would be one providing representative samples of genomic DNA, RNA, and blood plasma and the corresponding donor's personal information and medical history. While historically the process of storing such samples and tracking background information has been time-consuming and reliant on manual operations, the ability to automate many of these procedures now affords an opportunity to provide these samples in less time and at a lower cost. An increasing number of automated biorepositories are being developed and used for scientific research.
Many of the laboratory operations associated with cataloging samples, such as analysis, dilution, archival storage, and retrieval of purified human genomic DNA specimens, human RNA extracts, and plasma derived from peripheral blood, have a potential for automation. The Medical Automation Research Center (MARC) has constructed a large scale biological repository (biorepository) that maximizes the use of robotics and automation to perform these operations (Figure 1). The biorepository will be used to support research in gene-environment interactions in cancer [4]. The biorepository has the ability to sort and retrieve personal samples based on donor attributes. In its current form, the system has the capacity to automatically access up to 250,000 individual samples.
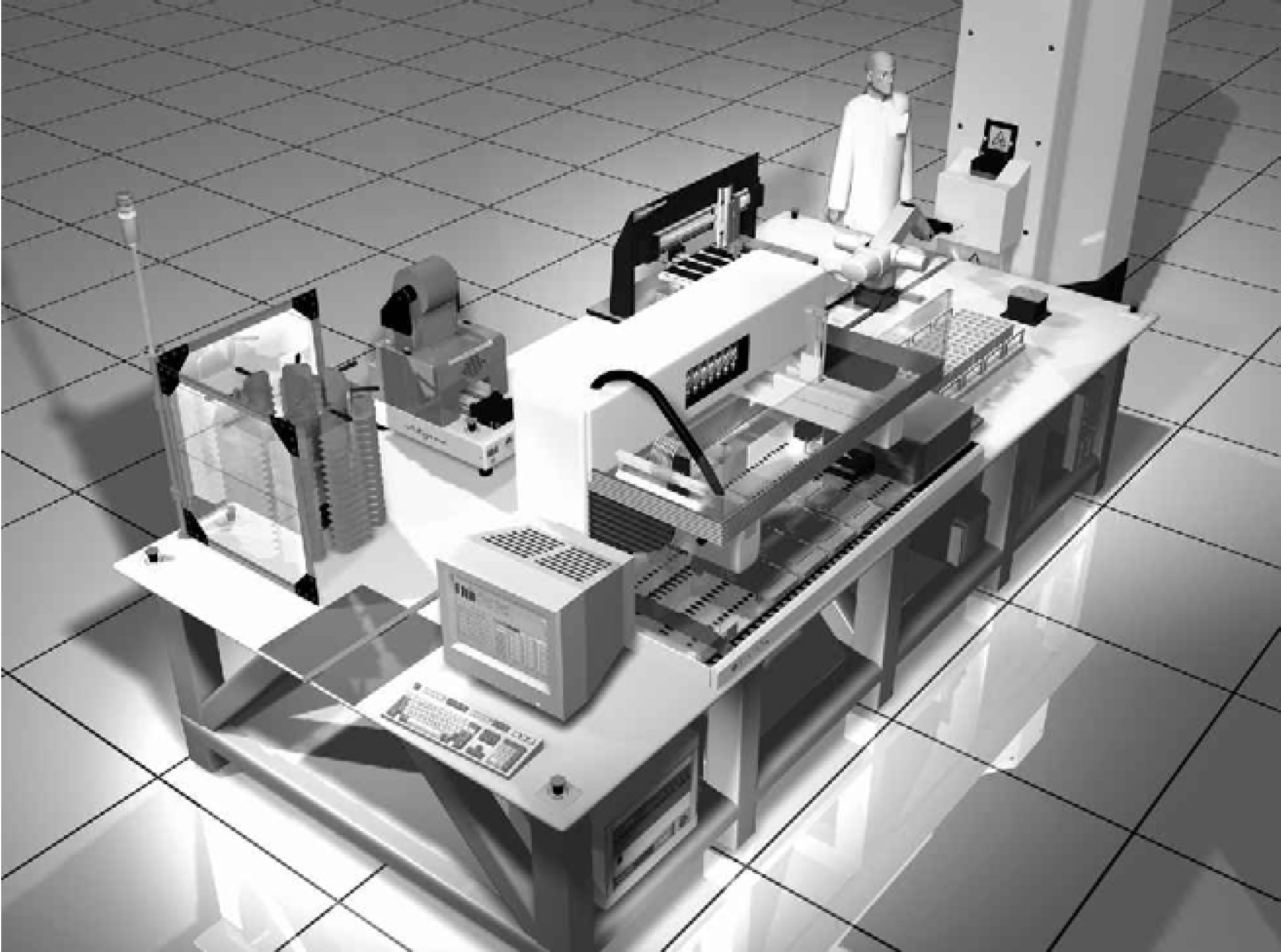

The biorepository system. This computer-generated view is from a three-dimensional animation. The animation was used to assist in system design and layout.
Our design uses a variety of off-the-shelf laboratory automation equipment, coupled with custom-designed hardware and software. The software architecture we have designed allows for simple reconfiguration of the system logic as needs require. We have created a two-level process control system using an industry standard process controller coupled with Microsoft Visual Basic. This approach allows system development to be performed by programmers with little or no robotics and automation expertise. After a brief overview of our system operation and design, we will present this software architecture in more detail.
OVERVIEW OF BIOREPOSITORY OPERATION
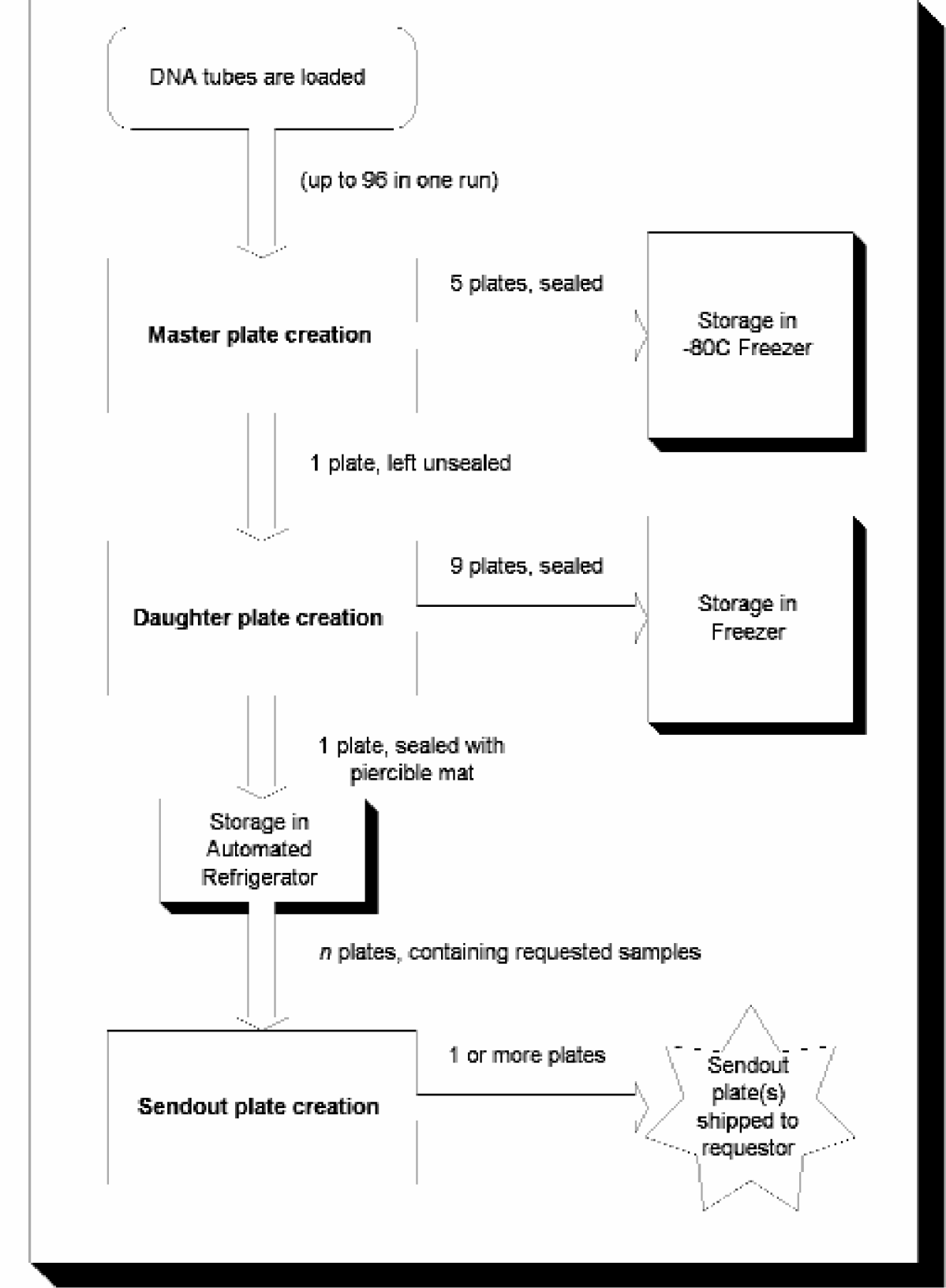
The biorepository is composed of a hardware system, a software system, and a set of standard procedures for presentation, selection, and send-out of DNA, RNA, and plasma. The primary function of the biorepository is to produce microtiter plates (MTPs) containing selected samples upon request. These samples are archived in the biorepository and are cataloged for later retrieval. When a request is made to the biorepository for DNA, RNA, or plasma samples, the samples are automatically gathered into a 350 uL round bottom MTP. This MTP is referred to as a Sendout Plate.
To archive DNA, the biorepository accepts racks of 50ml input tubes containing purified DNA. The DNA is distributed to 2.2 mL deep-well MTPs called Master Plates. These Master Plates contain up to 96 samples, and are typically sealed and frozen at −80°C. The concentration of DNA in Master Plates varies according to the concentration in the input tube. During the creation of Master Plates, the biorepository analyzes the samples using a fluorescent dye to establish the concentration of DNA of each sample. UV absorption analysis is used to measure the level of protein contamination in the samples. This information is permanently tracked in the sample database.
One Master Plate from each run is used to generate Daughter Plates. Daughter Plates are round-bottom 350 uL MTPs with the same physical arrangement of samples as the Master Plates from which they are created. However, the DNA in Daughter Plates is diluted to a standard concentration. Daughter Plates are intended to be loaded into an automated 4°C refrigerator for automated access. They are manually sealed using a pierceable silicon mat prior to their use in the system. Typically, 10 Daughter Plates are created at once from one Master Plate. Nine of these plates are placed into a freezer, and the tenth is loaded into the automated refrigerator. The process for Master and Daughter Plate creation is similar for RNA and plasma (plasma operations are performed using a chiller system to ensure sample integrity).
All input tubes and MTPs are individually bar-coded, and these barcodes are permanently tracked by the biorepository's database. Auditing and QC/QA functions are built into the biorepository. Together, these functions allow for completely automated DNA storage and retrieval. Figure 2 illustrates the operation of the biorepository.
Based on the goals of maximized automation and flexible programmability, the biorepository hardware design includes an anthropomorphic robot arm (CRS Robotics T255) in connection with a number of peripheral devices (Figure 3). These devices include:
TECAN Genesis RSP150 liquid handling station
TECAN SPECTRAFLUOR PLUS microplate scanner
TECAN MOL BANK™ automated microplate storage unit
ABGene ALPS-100 Plate Sealer
Sartorius LP 820 Balance
Torrey Pines Scientific EchoTherm Chiller
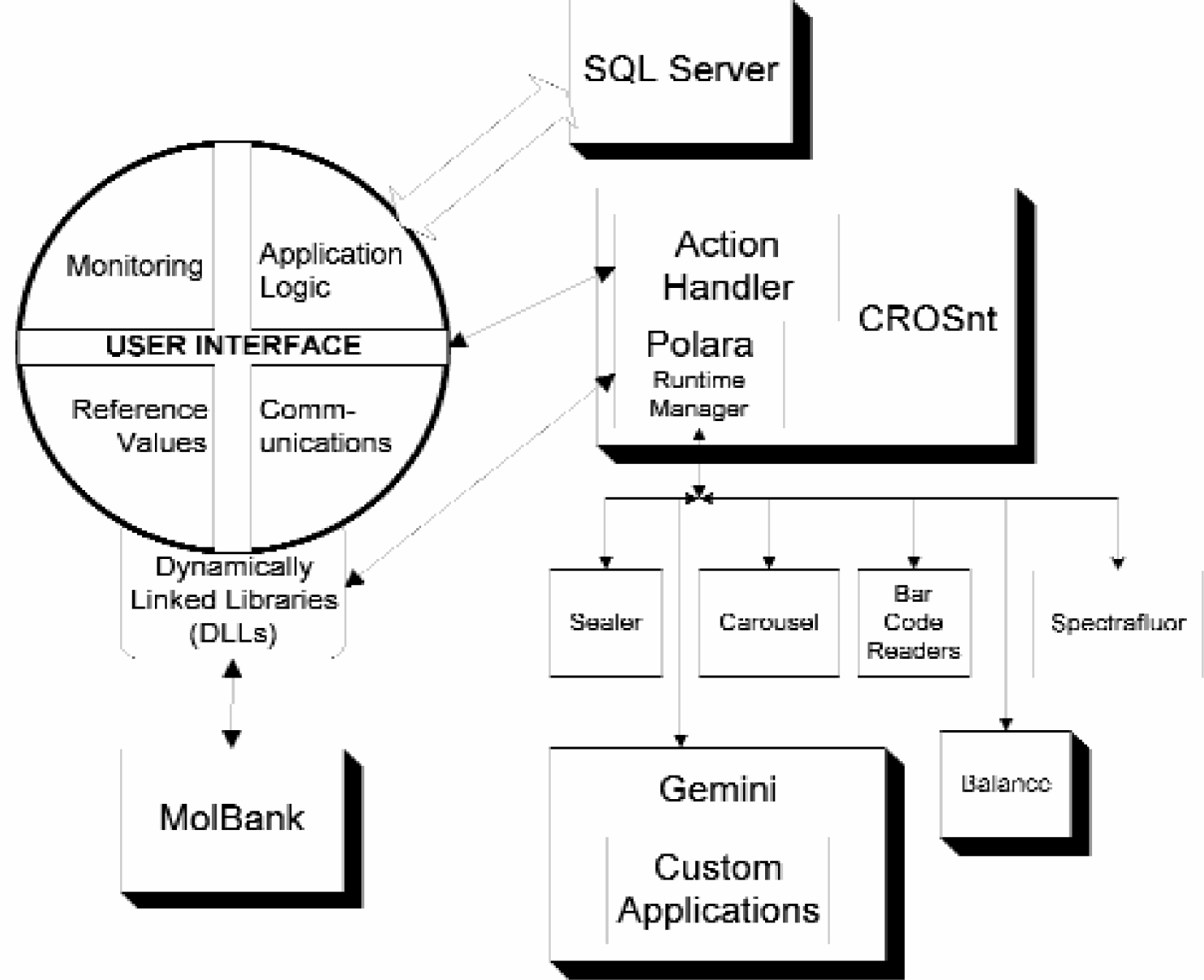
The software architecture of the biorepositiory. Arrows indicate data flow. As currently implemented, the user interface and CROSnt environment run on one Intel-based processor. The SQL Server is hosted on a separate processor.
The entire system of a robot plus its peripheral devices is interconnected via a network based upon POLARA (CRS Robotics, Inc.) software and hardware. The actual operation of the biorepository is accomplished using a custom written interface and operating system based upon Visual Basic (VB) and RAPL-3 (CRS Robotics). This software system runs on Windows NT. The software system also interacts with a Microsoft SQL (Structured Query Language) database server, which is resident on a separate Windows NT-based server. The biorepository includes expansion capabilities for PCR and DNA sequencing. Its components also support stand-alone operation for customer-developed automated tasks.
The biorepository software system coordinates the operation of separate but interrelated modules that are accessed through a screen-driven process. Accessioning of samples as well as microplates (Master Plates, Daughter Plates, and Sendout Plates) are all accomplished using a system of linear barcodes. Individual tubes in Master and Daughter Plates for the RNA and plasma samples are accessed through a 2D barcode system from Matrix corporation.
SOFTWARE ARCHITECTURE
The software architecture, like the hardware, uses a combination of off-the-shelf and custom components. The user interface (UI) is a Windows NT application. It communicates with a variety of instrument-specific applications to perform the processing tasks. For example, the robot arm is controlled by the CROSnt software package. During operation of the biorepository, the UI automatically invokes CROSnt for robot control. Similarly, the Tecan Genesis RSP150 pipetting workstation is controlled by the Gemini application. The biorepository communicates with Gemini to cause pipetting operations to occur. In both of these cases, we have developed custom applications in these control environments (CROSnt and Gemini). These custom applications are specifically designed to interact with the UI to process genetic material.
Other devices in the system (i.e., the Tecan MolBank automated refrigerator, the balance) are controlled directly from the UI using a Windows NT DLL. All database functions are handled by the Microsoft SQL Server. The SQL server runs on a second personal computer.
The biorepository's user interface allows the operator to start up the system, process samples, create plates, perform maintenance, and perform debugging operations.
During operation of the biorepository, the system uses Gemini (the control software for the TECAN Genesis 150) to initiate liquid-handling operations. The fact that the biorepository uses Gemini for liquid handling operations is actually an advantage for the Operator. Once familiar with Gemini, the Operator may adjust or modify the liquid handling operations that the biorepository uses. These operations can be run and tested independent of the biorepository operation. This enhances the flexibility of the system.
Our software architecture was originally to be based in the RAPL-3 language, that of the CRS robot arm. In this plan, application logic and scheduling would be encoded in RAPL-3, and user interface (UI) functionality would be performed by Visual Basic. As we developed the system, we realized that a more flexible approach was to isolate the atomic actions of the biorepository (i.e., moving samples, performing liquid handling, sealing plates) into primitives implemented in RAPL-3. We therefore designed a server application running continuously in the RAPL-3 environment. The server encompasses device control, low level error handling, and status monitoring. It reports status information to the VB client, which makes control decisions appropriately. Using this approach, we have enhanced the dynamic scheduling capability of the system and of application creation (changing or adding functionality of the biorepository). The UI is implemented in VB as originally intended, but with the additional control logic in place, the UI takes on additional roles of monitoring, verifying reference values (QA/QC), and implementing communications with the SQL server.
CURRENT STATUS AND FUTURE WORK
The biorepository is currently undergoing software integration testing and biochemistry verification using actual samples. Our goal is for the system to be fully operational by January 15, 2001. We have begun to assemble a list of design improvements that will be implemented in future versions of the system. These improvements will include:
smaller footprint
more intelligent error handling
more dedicated automation technology for increased throughput and increased reliability
a re-evaluation of the components integrated into the system
ACKNOWLEDGEMENTS
The authors would like to acknowledge the contributions of the MARC development team, including Chris Estey, Catherine Piche, and Kevin Bowman.
