Abstract

INTRODUCTION
The dramatic progress in the analysis of human, animal and plant genomes as well as parallel developments such as the human cancer gene anatomy project 1 have created an enormous demand for low-cost high throughput technologies for DNA and RNA analysis. Chip-based molecular techniques — if available in satisfactory quality for diagnostic applications — will enable major analytical issues in health care such as predisposition, cancer, infectious diseases and others to be addressed.
DNA micro-arrays have been predicted to become a key technology in molecular analysis for nearly one decade now 2 . Although their proportion in the research market (e.g. expression profiling 3 ) has increased significantly they still play a minor role in clinical and medical diagnostic fields. Main drawbacks for the implementation of BioChips in clinical routine analysis are insufficient reproducibility, lack of standardization as well as expensive chips and detectors. With respect to BioChip production, industrial standards and uniform protocols have not yet been defined. Therefore different BioChip layouts and analytical performance cannot be compared 4 .
This article describes the development and production of DNA/RNA BioChip arrays for diagnostic applications emphasizing in particular quality controlled production processes with high reproducibility of each step. In order to achieve the required analytical performance, many of the currently used components and techniques such as substrate material, coupling chemistry, arraying, probe design, hybridization and high resolution scanning were reevaluated by IMNT and replaced by appropriate new approaches. In addition, TECAN has developed a novel nanopipetting solution based on GENESIS pipetting robot to dispense volumes as small as 0.5 nanoliters (nl).

EXPERIMENTAL
DNA capture probes are modified at the 5′-positon with a hexaethylenglycol spacer and a unique photo reactive group for the coupling process. After spotting the capture probes onto a plastic surface containing an alkyl group, coupling is performed by a one minute irradiation at 365 nm. A preactivation of oligonucleotides and/or the surface is not necessary.
For an ongoing project focusing on expression profiling of disseminated cancer cells in blood 5 several perfect match and mismatch capture probes, derived from the HER2/neu oncogene 6 were designed. To validate the performance of the system, a 10μM oligonucleotide solution of an 18 mer (5′-TAT-TCA-GGC-TGG-GGG-CTG-3′) was spotted in single droplets onto the plastic surface. After photocoupling and washing, the oligonucleotide array was hybridized with a complementary 55 mer Cy5-labelled reporter probe in hybridization buffer.
The GENESIS NPS (NPS = NanoPipetting Solution) is equipped with novel Volume Control Centers (VCC) based on XP3000 Plus dilutors. In the GENESIS NPS used in the experiments, channel 3 is equipped with the nanopipetting capability, encompassing besides electronic components the nanopipetting device “ActiveTip M” (
A confocal fluorescence scanner with a resolution < 3 μm was used for quality assurance and signal detection. The dynamic range of the used fluorescence scanner covers four orders of magnitude. It can be extended by the use of reduction filters, mounted on an automated filter wheel. The scanner is optimized for the detection of Cy5.
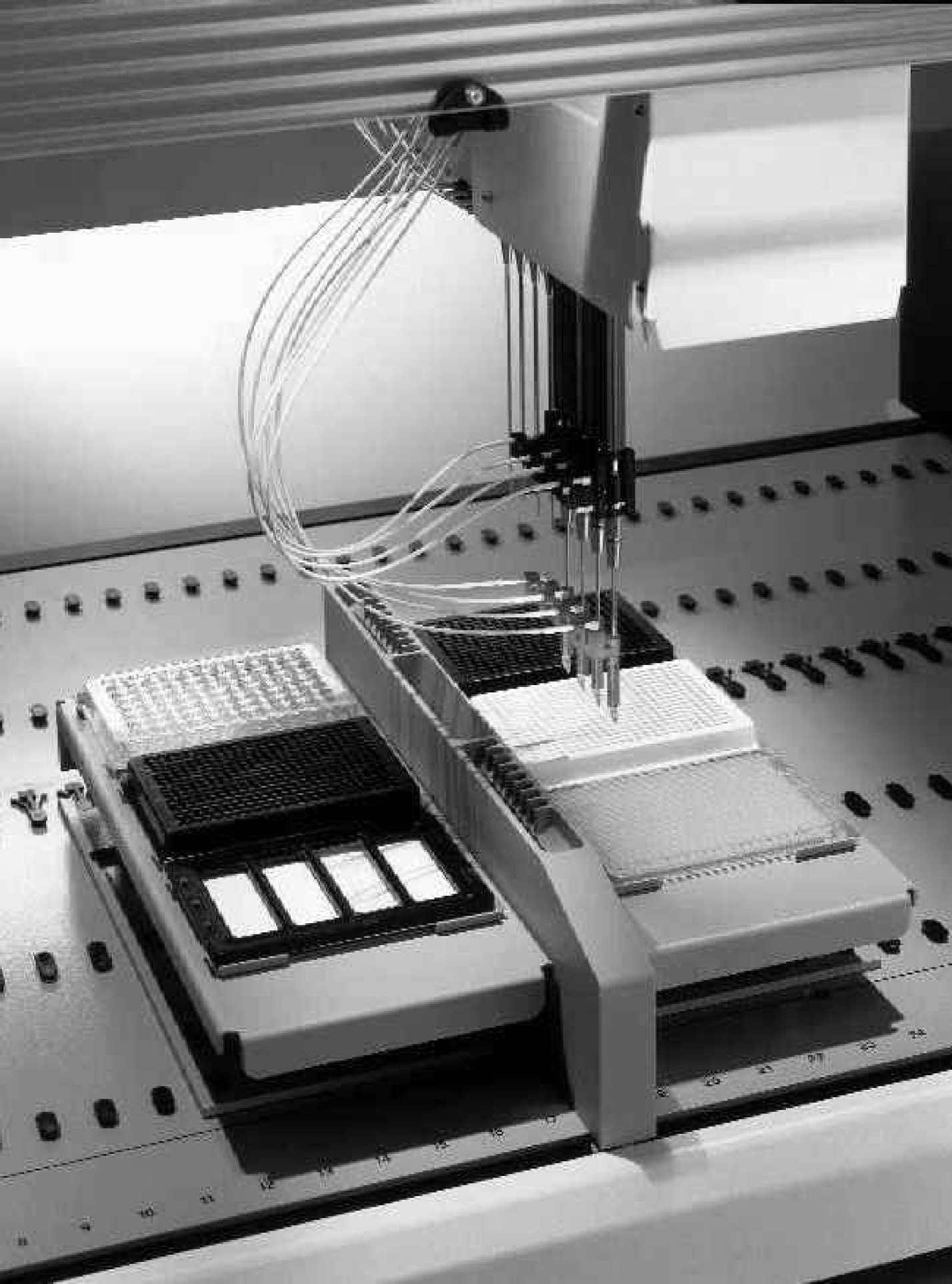

Worktable of a GENESIS NPS 100/8, equipped with 4 “ActiveTip M” and 4 standard tips for the transfer of DNA capture probes from microplates to slides.
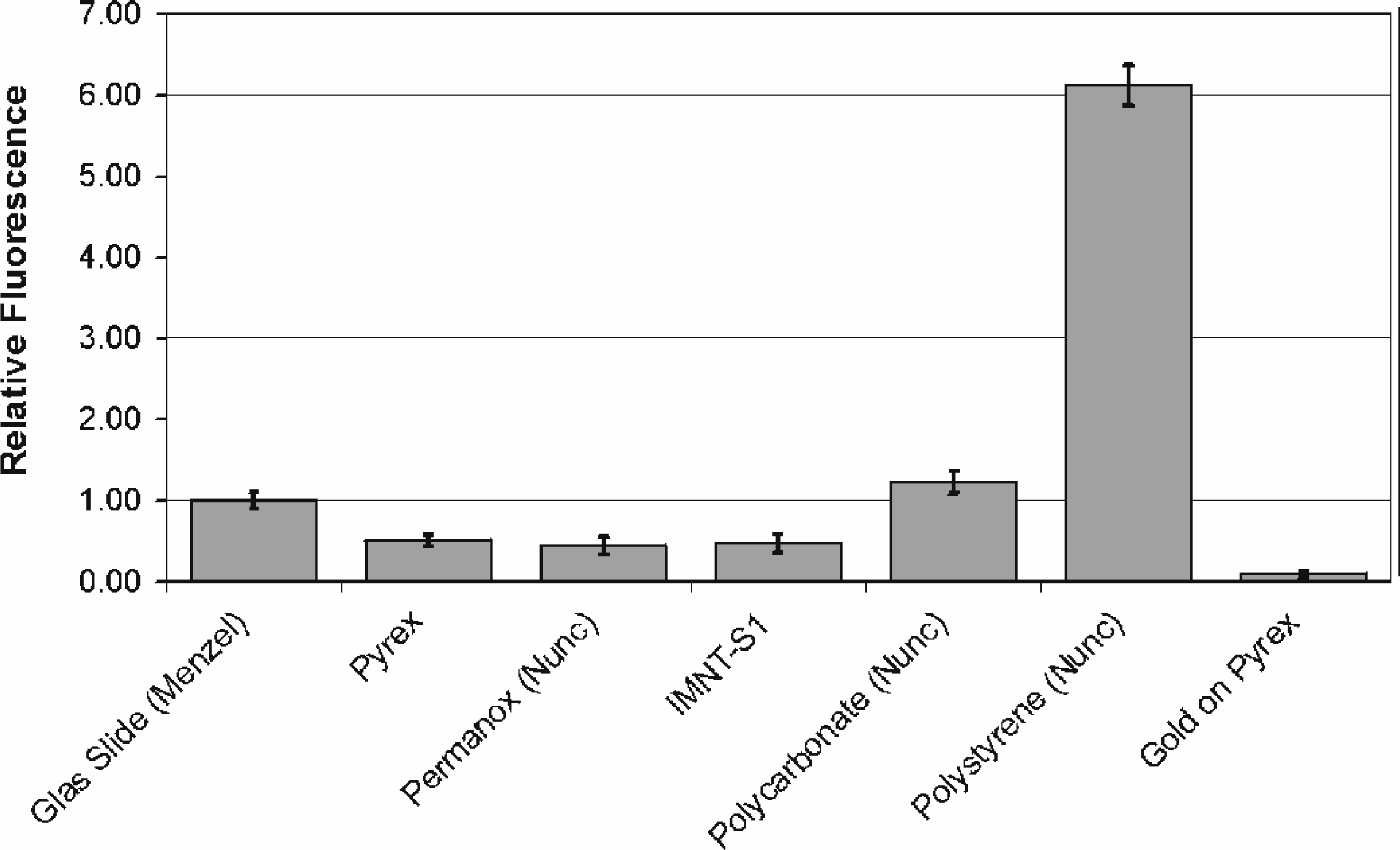
Background fluorescence of different support materials (n=3, 1s).
RESULTS AND DISCUSSION
The sensitivity and the reliability of DNA micro-arrays is strongly influenced by the properties of the solid support used for the coupling of oligonucleotides. We have examined several support materials for their suitability in BioChip production. As demonstrated in Figure 3, the materials differ significantly in their background fluorescence. The particular plastic material IMNT-S1 showed the highest contact angles with water (>90°), an optical transparency comparable to Pyrex glass and a low tendency for nonspecific binding of oligonucleotides. In addition it enables ultimately cheap production of tailored devices by injection molding or hot embossing.
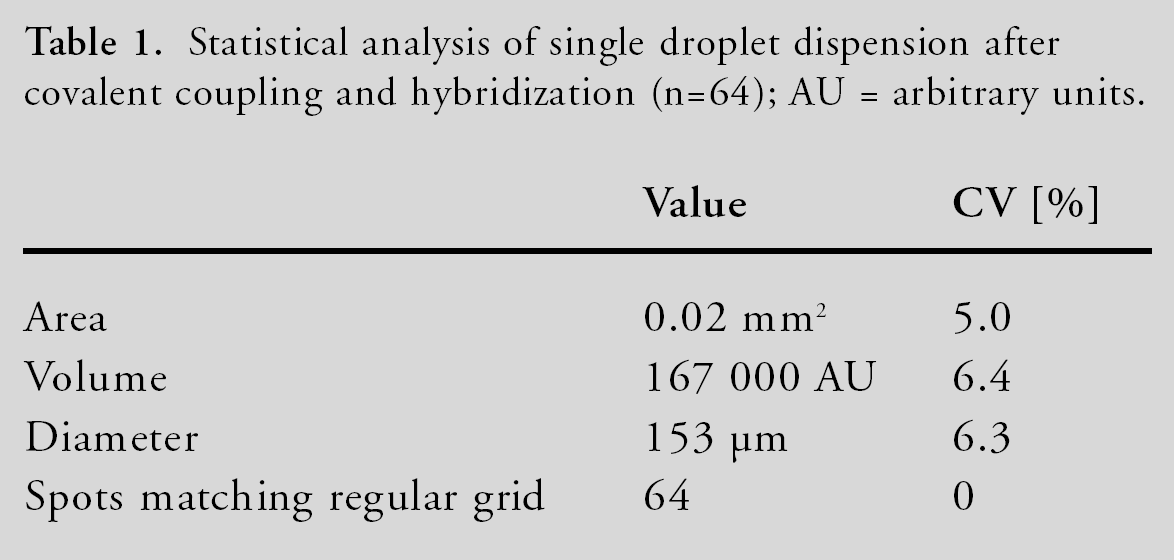
For the evaluation of the novel NanoPipetting Solution, single droplets of a 10μM oligonucleotide solution were dispensed with a 140 μs, 55 V pulse onto the plastic surface. The center to center distance of the spots is 300 μm. All spots were positioned within the given regular grid and no overlays or satellite spots were observed (
Statistical analysis of single droplet dispension after covalent coupling and hybridization (n=64); AU = arbitrary units

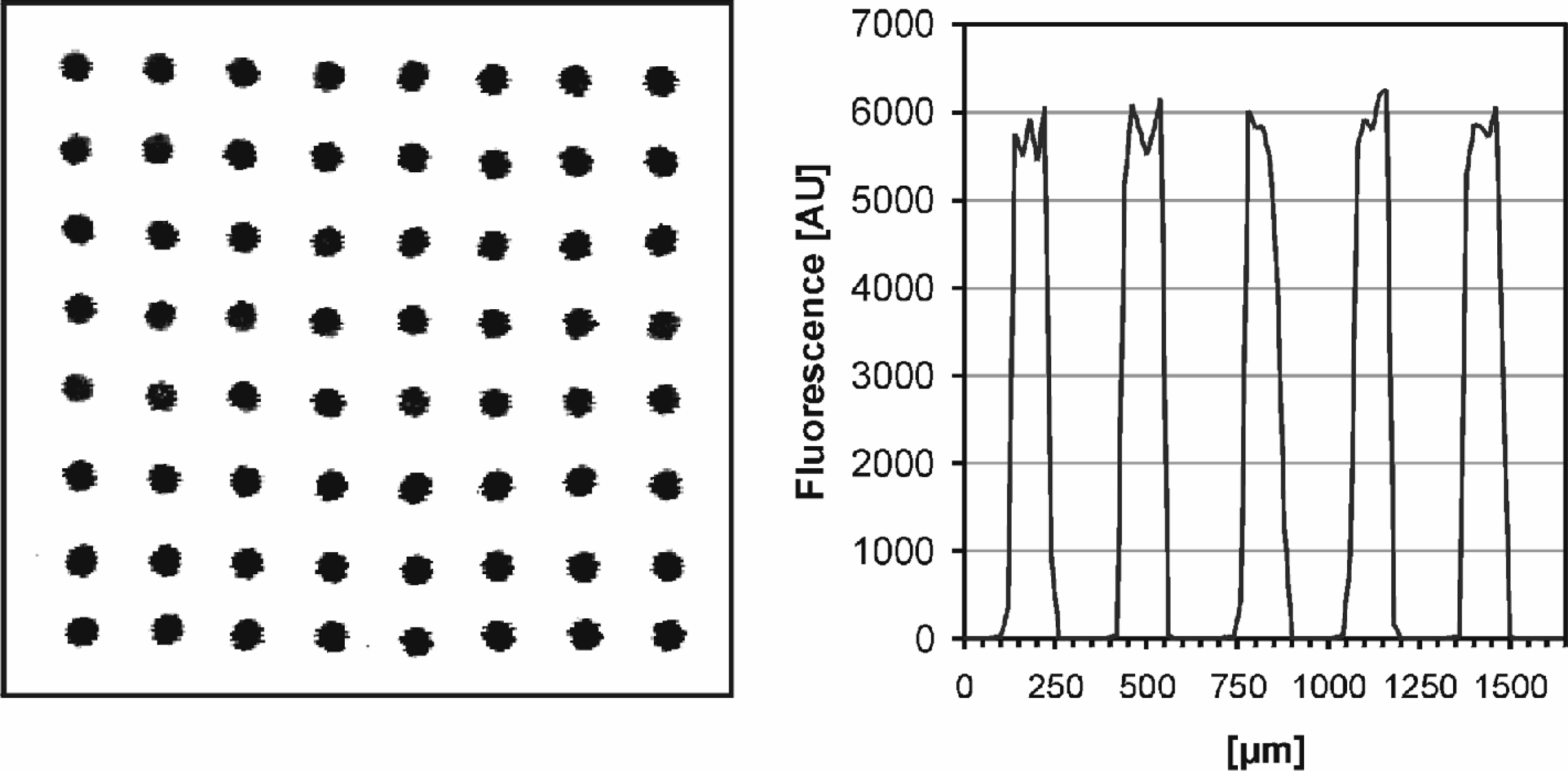
Fluorescence scan of a spotted array with 10 μm resolution (left). UV coupling of capture probes on IMNT-S1 plastic material results in the formation of homogeneous spots with even signal distribution (right).
The adapted coupling chemistry has several advantages: evaporation of the small droplets after dispensing does not influence the coupling efficiency, a complete saturation of the surface is achieved and the oligonucleotides are immobilized with uniform orientation. As demonstrated in
Especially for mutation analysis
8
and SNP detection, it is very important to discriminate between the wild type or perfect match and a target having only one base exchanged within the complementary sequence of the capture probe. The higher the discrimination ratio, the higher the analytical value and reliability of the obtained data. To optimize stringency conditions, the match/mismatch discrimination ratio was determined in this model for a perfect match probe (PM) and two mismatch probes (MM) with different base exchanges and positions within the 18mer oligo (MM GA9: C at position 9 replaced by A; MM CC7: G at position 7 replaced by C). After 1h hybridization with 5nM 55mer Cy5 reporter probe, chips were washed twice with 20% formamide in 0.2 × SSP-buffer. A PM/MM discrimination ratio of 3.8 for MM GA9 and of 16.4 for MM CC7 was obtained (
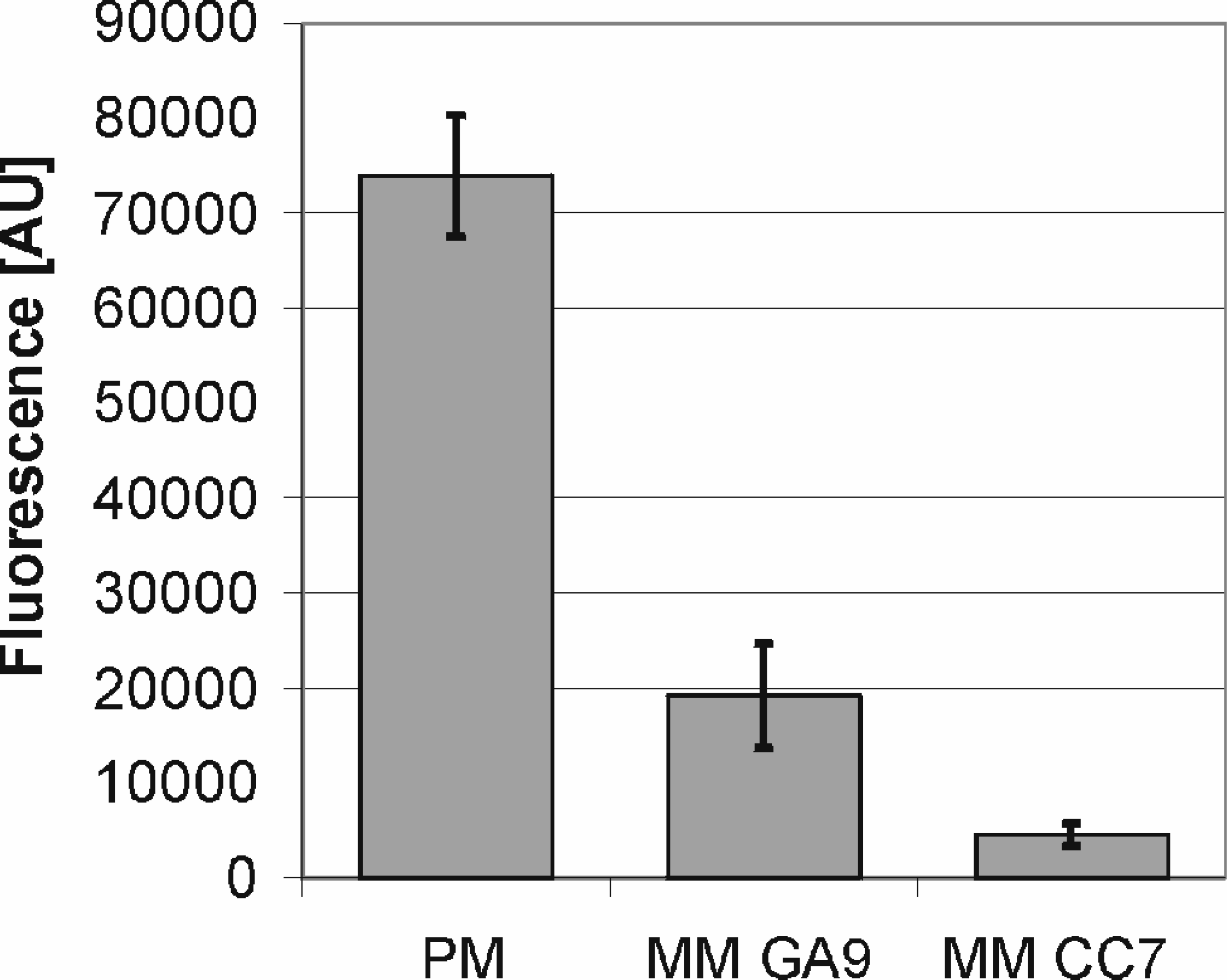
PM/MM discrimination with different base exchanges and positions (n=3, 1s).
Target concentrations differ significantly with respect to the biological material investigated. This concentration difference is enhanced when PCR based techniques for the amplification of the target sequences are applied. The higher the dynamic range for hybridizing BioChips the better the detection of target concentration differences and the applicability of the micro-arrays. The dynamic range for hybridization was determined by incubating the BioChip with a concentration series of 10μL 55mer Cy5 reporter probe under low stringency conditions. Absolute probe amounts from the attomole to the picomole range (>104) can be measured with the presented setup as shown in
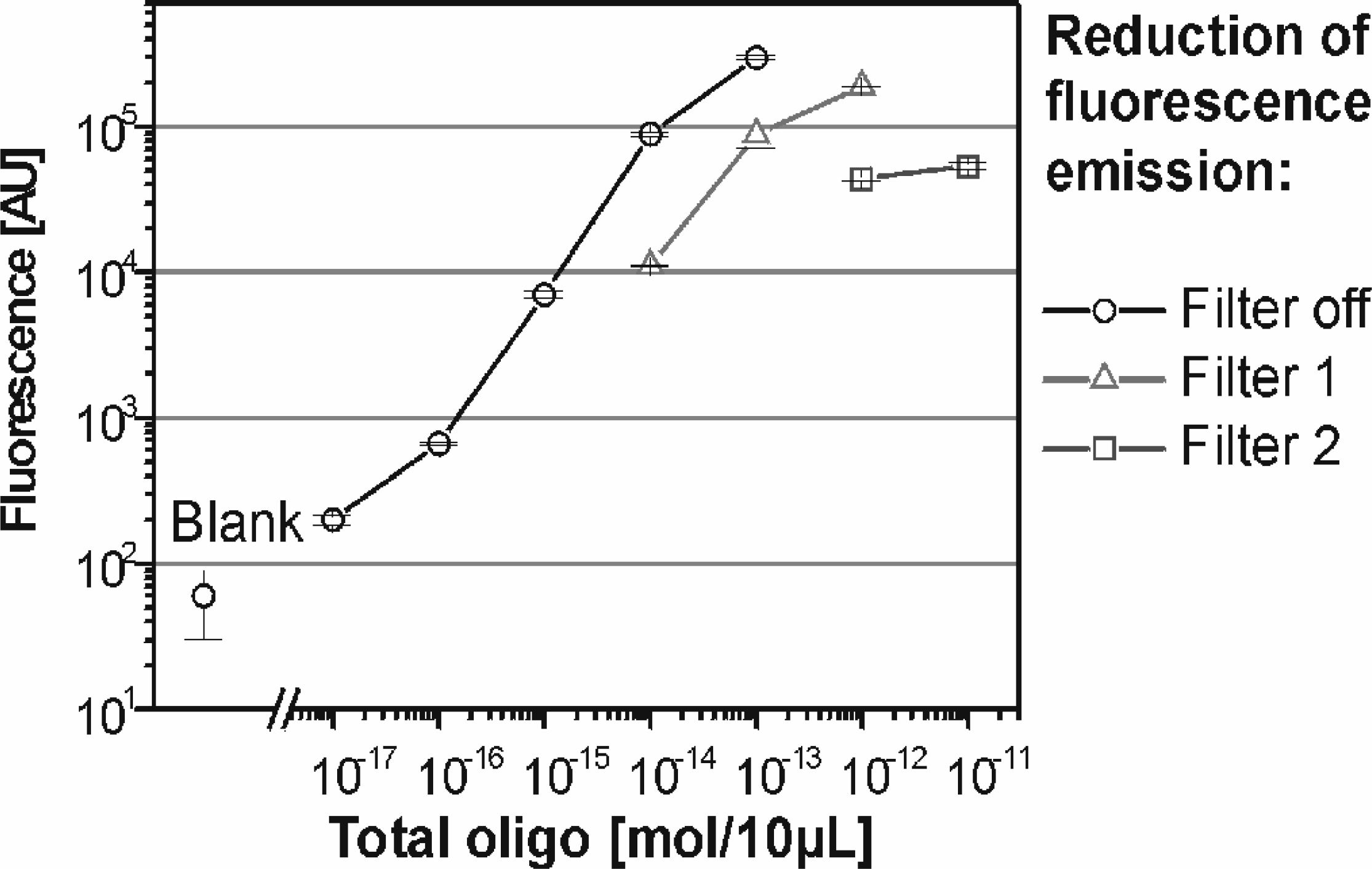
A dynamic range for hybridization of 105 is achieved for the IMNT BioChip (n=3, 1s).
CONCLUSION
Using the novel NanoPipetting Solution GENESIS NPS, the production of oligo-nucleotide arrays with respect to accuracy, reproducibility, carry over and avoidance of satellite spots was significantly improved. Combining the new support material, the novel coupling chemistry and the unique performance of the NanoPipetting Solution, major milestones towards production of quality controlled BioChip arrays have been achieved. The integration of the nanopipetting capability into the GENESIS platform enables adjustment of the capture probe working solutions (dilutions etc.) and subsequent microarraying with one instrument. A fully automated production of medium number BioChip series can be realized.
High fidelity DNA micro-arrays for mutation analysis and for expression profiling for clinical routine analysis will be available from IMNT by the end of the year. For read out of the BioChips a new low cost, high performance chemiluminescence detector will be implemented.
