Abstract

In 1997, a group of scientists and innovators recognized the need for a global repository of human genomic DNA samples. Such a repository was envisioned to comprise up to a million samples covering a wide spectrum of diseases across a multitude of ethnicities. The samples would be coupled to extensive disease-specific phenotypic data gathered from diagnostic assays and questionnaires. To achieve this goal, the repository would have to be organized on an industrial scale with highly automated storage and retrieval systems and computer interfaces. Genomics Collaborative was established to build such a repository. The DNA and phenotypic data from these samples will be made available to researchers throughout the world to promote research and facilitate finding cures for diseases represented in the repository.
CREATING THE REPOSITORY
To create the repository, whole blood is collected and processed to extract DNA. After extraction, the DNA is normalized to a standard concentration, checked for RNA or protein contamination, examined for possible cross-contamination and to ensure that the DNA can be amplified. The DNA is then frozen for long-term storage, with some of the DNA kept in liquid form for ready access. These samples are maintained in the Genomics Collaborative facility in Cambridge, MA, and at an alternate site for risk management. The DNA samples will be offered in liquid form in flexible increments.
When the Genomics Collaborative first began operation, the crucial but tedious normalization step created a significant bottleneck. A technician prepared a UV specific 96 well microtiter plate with each sample in triplicate. The plate was then read by spectrophotometer to determine DNA concentration and purity. The technician would then calculate the dilution factor necessary to reach the desired concentration, and then hand-dilute each sample.
MOVING TO AUTOMATION
Management wanted to ramp up the output of the normalization process, which at that point was the step that constrained the speed at which the repository could be created. They were also concerned that the tedium of the job might lead to dissatisfaction among some members of the team, about the possibility of repetitive stress injury and high likely error rate. Several technical personnel were assigned to investigate robotic workstations that might be appropriate for this task. They were unanimous in recommending the GENESIS robotic sample processor (RSP), 150 from Tecan, Research Triangle Park, North Carolina. Their primary reason for recommending this instrument is its unique programming flexibility that makes it possible for virtually every function to be controlled by programs in popular languages such as JAVA and Visual Basic that can access external databases and prompt the user to set parameters for the current process.
Management agreed to the purchase of the Tecan robotic workstation and several staff members were assigned the task of creating programs needed to automate the normalization process. For typical laboratory tasks, the workstation can be visually programmed by graphically defining the procedure on a screen that is designed to mimic the appearance of the instrument. This application was much more complicated because it required reading the results of the concentration measurements and then calculating the amount of buffer that needed to be dispensed into the sample tube in order to achieve normalization. The team decided to incorporate the Tecan robotic workstation into GCI bioinformatics framework, which guaranteed data sharing with the rest of automation components. Tecan could also produce mapping data that was to be used by other devices as well as to use concentration results from the spectrophotometer.
PROGRAM SEQUENCE
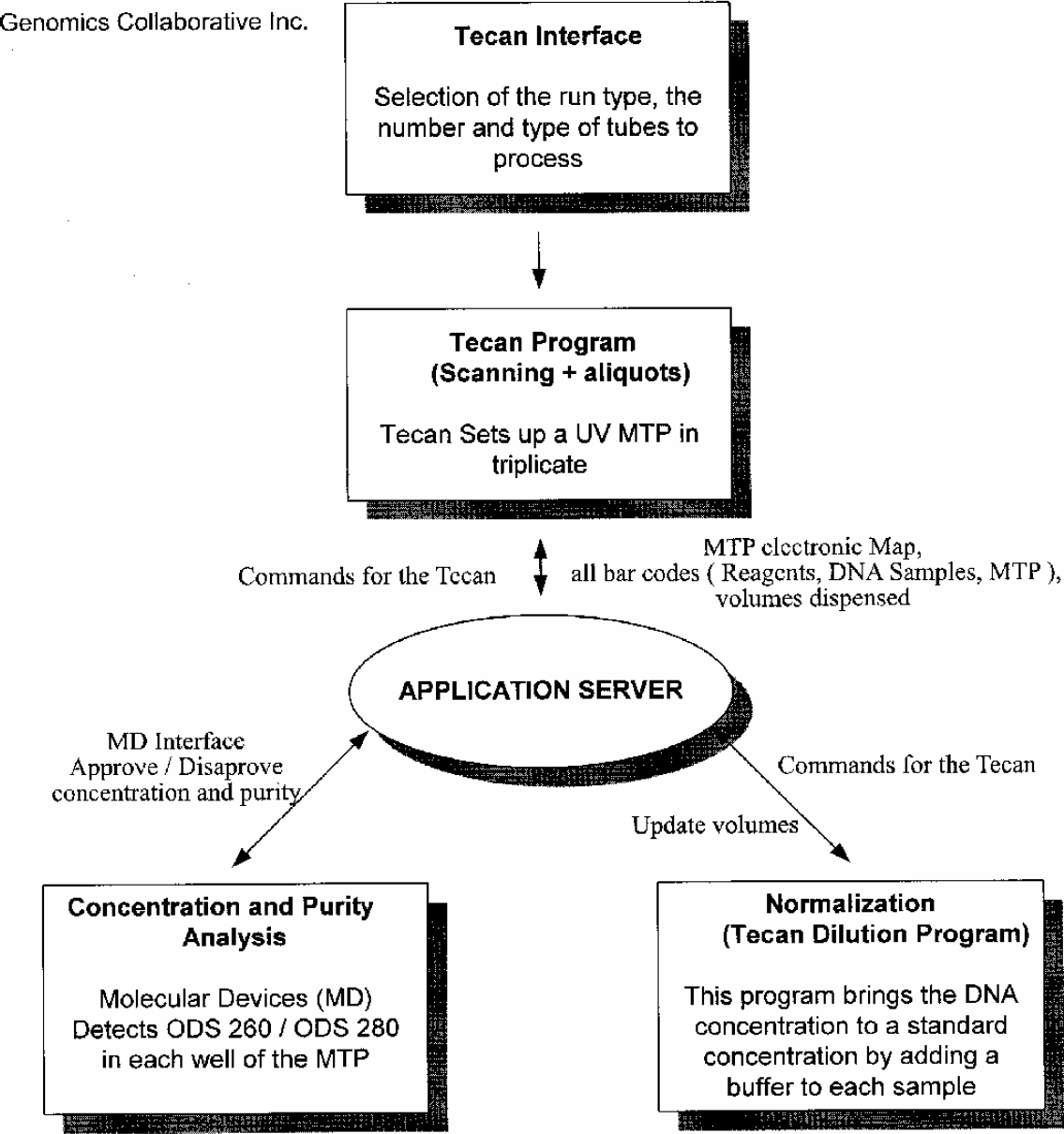
The program developed by the team works as follows. The user is asked to select the type of operation desired such as the normalization procedure described above or several others written by Genomics Collaborative programmers (such as genotyping, quality gel analysis preparation, etc.). Next, the program asks the user to select the types of tubes in which the samples are stored allowing flexibility of the system. Finally, the program prompts the user to select the number of racks of tubes that will be processed. From this point on, the instrument is mainly on its own.
Tecan Genesis sets up a UV MTP in triplicate.
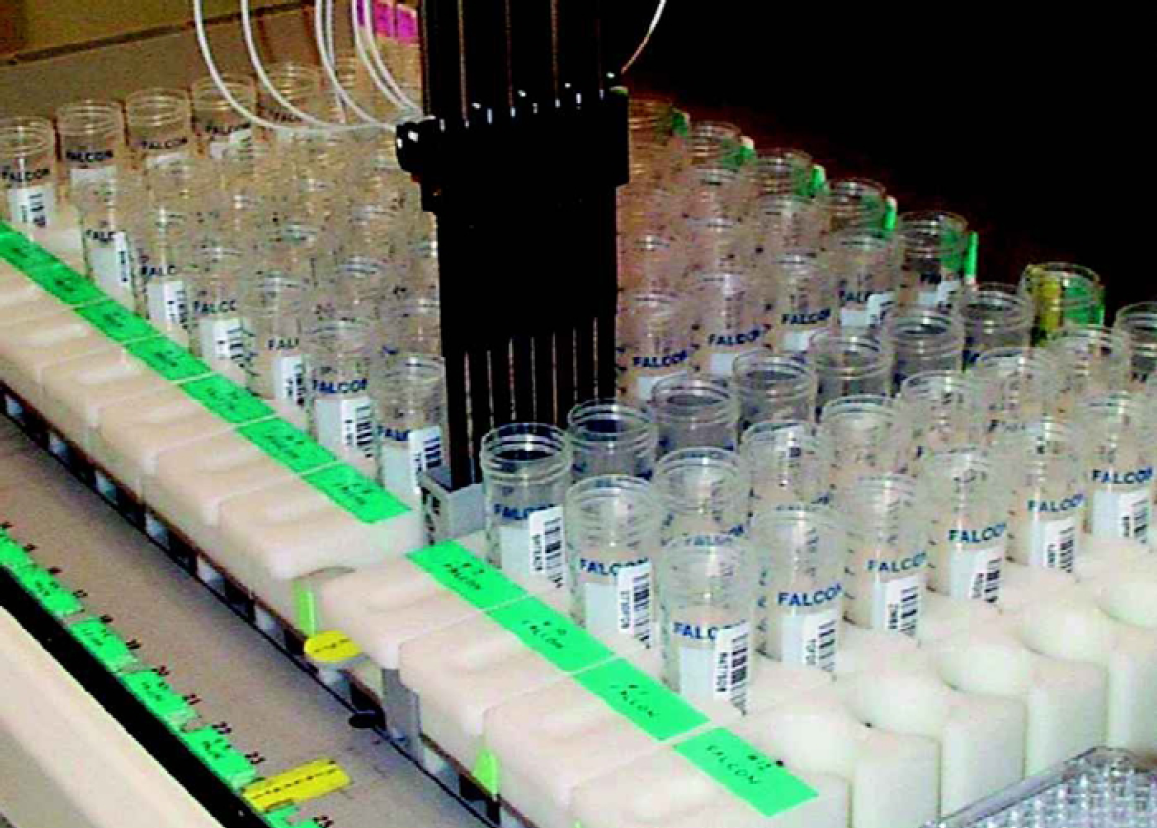

After normalization, the DNA samples are tested again to confirm their concentration by repeating steps 1 through 4. The program will then update the database with the new concentration for each sample as well as the total volume.
DNA Library Normalization Process
THROUGHPUT GAINS
Through automation, GCI has been able to normalize ten times faster than with manual methods. The technician responsible for operating the instrument has to load and unload the samples, add microplates, tips and reagents when necessary, run the initial set-up program and intervene at each reader step. The program tracks all samples, calculates concentrations and dilution factors, updates volume of each sample and keeps historical records of all transactions.
The net result of GCI's automation effort has been improved throughput, decreased hands-on time by the technical staff, decreased risk of repetitive motion injuries, decreased risk of human errors, improved accuracy, traceability and a happier staff by eliminating a tedious, often painful manual task.
For further information contact Tecan US, Inc., PO Box 13953, Research Triangle Park, North Carolina 27709. Phone: 919-361-5200 Fax: 919-361-5201.
Laurent Jacotot and Mark Pytlik with the Tecan Genesis 150

