Abstract
Background
The human X-ray repair cross-complementing protein 1 (
Methods
A total of 89 HCC patients and 99 randomly selected healthy controls were enrolled. Genotyping of XRCC1 rs25487 was performed by high-resolution melting analysis and Sanger sequencing.
Results
On univariate analysis, a statistically significant association was found between risk of HCC and
Conclusions
To our knowledge, this is the first study reporting an association between BER SNP and HCC risk in a population of central-southern Italy.
Introduction
The majority of hepatocellular carcinoma (HCC) cases result from DNA damage caused by hepatitis viruses that are the main potential risk factors for the development of HCC (1). Damage due to endogenous or exogenous exposure may be repaired by enzymes encoded by the DNA repair pathways, such as base excision repair (BER), nucleotide excision repair (NER), mismatch repair (MMR), homologous recombination repair (HRR) and nonhomologous end joining (NHRR). Polymorphisms in DNA repair genes may influence the individual DNA repair capacity, which is crucial for preventing genomic instability and, in turn, may be associated with higher risk of cancer (2). This variable DNA repair capacity correlates with peculiar characteristics such as those expected for cancer susceptibility genes: The proteins encoded by these alleles exhibit reduced rather than absence of function, and this impairment may increase the disease risk. The genes involved in DNA repair can carry some polymorphisms, and the different alleles exhibit incomplete penetrance (3). Among the various DNA repair pathways, BER plays a key role in the removing of DNA damage deriving from exposure to various endogenous and exogenous carcinogens. The BER pathway removes alterations of a single oxidized, reduced or methylated base (4). A potential role for DNA repair genes in hepatocarcinogenesis has been underlined in a recent study showing that transcription of most of these genes was up-regulated in cirrhotic liver of HCC-bearing patients (5).
The X-ray repair cross-complementing group 1 (XRCC1) protein plays a central role in DNA repair pathways (6).
Several polymorphisms in the
The aim of our hospital-based case-control study involving a central-southern Italian population was to clarify the effect of the
Materials and Methods
Participants
Study participants were recruited among patients admitted to the Agostino Gemelli teaching hospital of the Catholic University of Rome (Italy), and eligibility was restricted to white individuals born in Italy. Cases were selected from patients with diagnosis of HCC, admitted to the outpatient liver unit of our hospital. Criteria for exclusion were secondary or recurrent tumors. We reviewed clinicopathological features such as Child-Pugh class, Barcelona Clinic Liver Cancer (BCLC) stage and etiology from medical records. The control group was randomly selected among patients admitted to the same hospital during the same period with a broad range of diagnoses, but without malignancy, chronic diseases, digestive diseases or any prior history of malignancy. Written informed consent was obtained from all study participants, after which each participant provided a venous blood sample. This study was performed according to the Declaration of Helsinki and was approved by the ethics committee of our university.
DNA samples
Genomic DNA was isolated from peripheral blood samples, collected by venipuncture, using High Pure PCR Template Preparation kits (Roche Diagnostic, USA). Extracted DNA samples were quantified by spectrophotometer at 260l group was Few stu20ing High Purcessing. DNA integrity was electrophoretically tested.
Genotyping
Genotyping of XRCC1 rs25487 was performed by high-resolution melting analysis using a LightCycler 480 system (Roche). The reaction was performed in 20 μL final volume including 10 μL of 2× Light-cycler 480 High Resolution Melting Master mix; 2.5 mM final concentration of MgCl2. The primer pairs used were forward, 5′-TAAGGAGTGGGTGCTGGACT-3′, and reverse, 5′-ATTGCCCAGCACAGGATAAG-3′. All primers were designed with Primer3 software and synthesized by Eurofins MWG Operon. Overall samples were amplified on the Roche LightCycler 480 using a touchdown polymerase chain reaction (PCR) program: 95 aC for 10 minutes, followed by 45 cycles of denaturation at 95 WC for 10 seconds, annealing starting at 62 C for 15 seconds and decreasing until 5362 denaturation at 95WG C for 1 minutes. Following the amplification steps, PCR products were denatured at 95°C for 1 minute and cooled to 40 mC so as to form heteroduplexes.
These steps were followed by a high-resolution melting program consisting of a first-step heating at 95 hC and a melting program with temperatures ranging from 65 rC to 95°C. Melting data were analyzed using the Gene Scanning module of the LightCycler 480 software (version 1.3), incorporating normalization of premelt and postmelt regions, temperature shift adjustment and calculation of difference curves. To assess how well our protocol was able to discriminate the mutant from the normal allele, the melting curve of each amplicon, deriving from the heterozygous DNA sample, was compared with that derived from same amplicon from a normal DNA sample (previously sequenced by Sanger method, in triplicate). A randomly selected pool of HRM prescreened samples (1 pool per genotype) was also sequenced for further genotype confirmation.
Statistical analysis
Adjusted odds ratios (ORs) and relative 95% confidence interval (95% CI) estimates were calculated using a conditional multiple regression model to measure the association between HCC and putative risk factors. The exact logistic regression model was also used where appropriate (22). We also examined the possible confounding effect of alcohol and chronic infection with HBV and/or HCV. However, models including these covariates yielded very similar results; thus, given the small numbers in some strata, only the age- and sex-adjusted estimates were presented. A chi-square test of Hardy-Weinberg equilibrium (HWE) for the 3
Results
Our study population included a total of 89 HCC patients and 99 controls. We consecutively contacted 120 healthy individuals, who were asked to participate in this study: Only 99 (82.5%) agreed to participate, while 17.5% of them were not interested or did not have enough time to be interviewed and informed regarding the study. The clinical characteristics of our HCC patients are reported in Table I. Significant differences were found both for sex (p<0.001) and for age (p<0.001). The percentage of males was higher among HCC patients than controls (78.6% vs. 48.4%). The mean age was 66.28 years (±10.53) in HCC patients, and 83.76 years (±3.60) in controls. The age differences between the 2 groups were expected because the control group was deliberately chosen from a pool of individuals whose age excluded them from the risk of developing HCC. In fact, besides the ascertainable possible causes, HCC can also develop for reasons defined as cryptogenic. Considering that development and progression of HCC are very slow (about 20 years), it was unlikely that the individuals chosen as controls in the study would develop HCC.
Clinical features of HCC patients
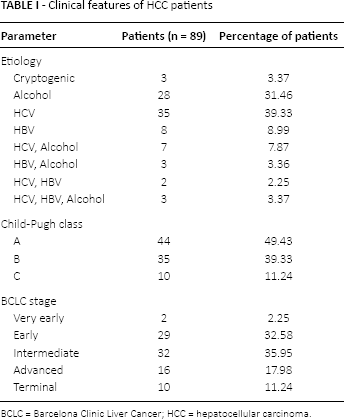
BCLC = Barcelona Clinic Liver Cancer; HCC = hepatocellular carcinoma.
Almost 50% of the HCC patients had Child-Pugh class A HCC. Among the HCC patients, 52.8% were HCV-positive, and 36% were in the intermediate BCLC stage. Statistical analysis did not find any significant association between the genotype 399Arg/Gln and susceptibility to HCC in patients with HCV/HBV or other factors (data not shown).
Table II shows the distribution of the genotypes
Frequencies of
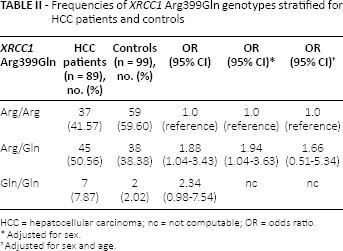
HCC = hepatocellular carcinoma; nc = not computable; OR = odds ratio.
Adjusted for sex.
Adjusted for sex and age.
Out of 89 patients who were treated and received chemotherapy after surgery or radiotherapy with or without chemotherapy in the Agostino Gemelli hospital, 25 died within a few years from enrollment or during the treatment. Monitoring over time was possible for the remaining 64 individuals for whom we calculated that the median survival period was 20 months (SD = 21.3). Although not significant, Kaplan-Meier analysis showed a decreased median survival in Arg/Gln genotype carriers in comparison with Arg/Arg carriers (Tab. III). These findings are also shown in Figure 1.
Median survival obtained by Kaplan-Meier analysis
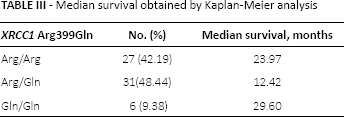
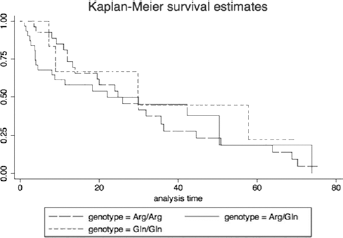
Kaplan-Meier survival curves.
There were no statistically significant differences among the 3 genotypes with regard to survival (log rank test, χ2 = 0.47, p = 0.79).
Discussion
In this study we tried to verify the possible association between
Lately, some molecular epidemiological studies have been conducted to evaluate the role of this
We underline the fact that limited reports on the role of
Since this is preliminary report, our study is affected by the following limitations: (a) although there are 3 common single-nucleotide polymorphism (SNP) sites in the
In conclusion, despite all of the limitations reported above, our study suggests that the 399Arg/Gln genotype in the
Footnotes
List of abbreviations
Financial support: Funding for this study was received from the Erasmus Mundus Western Balkans Fellowship (to B.N.).
Conflict of interest: The authors declare they have no conflict of interest.
