Abstract
Background:
Head and neck squamous carcinoma (HNSCC) is caused by different exogenous risk factors including smoking cigarettes, alcohol consumption, and HPV infection. Base excision repair (BER) is the frontline to repair oxidative DNA damage, which is initiated by the DNA
Objective:
Explore whether BER genes’ (
Design:
RNA sequencing (RNA-Seq) and clinical information (age, gender, histological grade, survival status, and stage) of 530 patients of HNSCC were retrieved from the Cancer Genome Atlas. Patients’ data are categorized HPV positive or negative to analyze the tumor data including the tumor stage,
Methods:
RNA-Seq of HNSCC data retrieved and mutation count and aneuploidy score were compared using an unpaired
Results:
Our data analysis reveals that BER genes frequently overexpressed in HNSCC tumors and increase mutation count. In addition, OGG1 and POLB overexpression are associated with low infiltration of immune cells, low immune checkpoint gene expression (PD-1, cytotoxic T-lymphocyte antigen 4, program death ligand 1, and program death ligand 2), and innate immune signaling genes. Furthermore, dysregulated BER factors in
Conclusion:
Our analysis suggests that dysregulation of the BER genes panel might be a potential prognosis marker and/or an attractive target for an immune checkpoint blockade in HNSCC cancers. However, our observation still requires further experimental-based scientific validation studies.
Introduction
Head and neck carcinoma (HNSCC) is a heterogeneous tumor located at the nasopharynx, oropharynx, hypopharynx, larynx, and oral cavity. 1 HNSCC accounts for about 4% of all cancers in the United States and an estimated 66,920 people (49,190 men and 17,730 women) will be diagnosed in 2023. 2 Over 90% of HNSCC are squamous cell carcinomas and are caused mainly by environmental risk factors.3 –6 In addition, alternation of the genetic landscape of DNA damage repair enhances susceptibility to HNSCC.7,8
Interestingly, HPV-positive HNSCCs have distinct clinical, pathological, and molecular features compared to HPV-negative HNSCCs.9,10 HPV-induced HNSCCs are triggered by E6 and E7 onco-viral proteins. E6 promotes the degradation of the cellular proteins that lead to cell proliferation, and E7 promotes genomic instability and resistance to apoptosis. 11 Furthermore, E6 and E7 cause dysregulation of the cell cycle and DNA repair pathways, resulting in a dramatic increase in genomic instability.12–14 HPV-positive HNSCCs harbored oxidative stress associated with DNA damage with impaired DNA repair.12,15
Base excision repair is one of the key pathways to repair oxidative and alkylating agents and dysregulated in the tumor.16,17 In the first step of BER, reactive oxygen species-associated DNA damage, including oxidation of guanine to 8-dihydro-7,8-oxoguanosine (8-oxodG), is processed by OGG1 and apurinic/apyrimidinic endonuclease I. 18 This leads to a DNA gap, which is then filled with one or more nucleotide by DNA polymerase β (Pol β) and the DNA nick is sealed by DNA ligase III. 19 The DNA glycosylase not only recognizes and removes the damaged base but also protects cells from mutagenic and/or cytotoxic effects of DNA base lesions while still maintaining genomic integrity.20,21 Aberrant expression of BER creates an imbalance in the BER pathway, which leads to saturation of downstream repair enzymes and/or accumulation of genotoxic BER intermediates.18,22,23
BER expression is deregulated in several types of cancer, which leads to genomic instability, ultimately impacting treatment outcomes.24–27 To date, many studies have tried to establish prognostic signatures, including gene sets 28 as promising predictors of prognosis for HNSCC. There is a significant need to explore potential prognostic predictors in HNSCC. Therefore, it is crucial to identify new DNA repair markers that predict an antitumor immune response to achieve personalized treatment. In this study, the expression of OGG1 and Pol β in HNSCC tumors and the relationship between overexpression of POLB and OGG1 with genomic instability and tumor immune infiltration were investigated.
Methods
Data acquisition
RNA sequencing (RNA-Seq) and clinical information (age, gender, histological grade, survival status, and stage) of HNSCC (
Exclusion and inclusion criteria
Since BER (POLB and OGG1) is the primary source of interest for this analysis, only individuals with valid RNA Seq V2 RSEM data for BER were included. Individuals with a
Mutation count and aneuploidy score
The mutation counts and aneuploidy score for each individual were extracted from the TCGA data set for HNSCC. Values were grouped according to low and high BER gene expression for a given cancer type and an unpaired
Immune cell infiltration in HNSCC
The TIMER algorithm was used to calculate the tumor abundance of six infiltrating immune cells (CD4+ T cells, CD8+ T cells, B cells, neutrophils, macrophages, and dendritic cells) based on RNA-Seq expression profile data. The correlation between the POLB, OGG1, and immune cells was calculated by Spearman correlation analysis using TIMER 2.0.
Estimation of stromal and immune cells in tumor tissues
The ESTIMATE algorithm-generated matrix, immune scores, and stromal scores were used to estimate the level of infiltrating matrix and immune cells in HNSCC tissue and tumor purity through expression profiles. Results for the tumor-infiltrating immune component were yielded with data extracted from the TCGA database, which was analyzed by the CIBERSORT algorithm.
Statistical analysis
Group comparisons for continuous data were conducted using
Results
Patient data analysis and clinical characteristics of BER factors in HNSCC patients
Data were accessed from cBioPortal (www.cBioPortal.org). TCGA PanCancer RNA-seq data and corresponding clinic information for HNSCC types were retrieved for validation. TCGA pan-cancer data sets were analyzed based on the expression of OGG1 and POLB from 523 HNSCC patients. The mRNA expression of OGG1 and POLB was significantly high in tumor

Overexpression of OGG1 and POLB in HNSC cancer. (a) The comparison of expression of mRNA in normal tissue adjacent to tumor
BER overexpression correlated with overall survival of HNSCC patients
To investigate whether the aberrant expression of the BER in tumors alters patient survival, Kaplan–Meier survival curves were generated based upon the designation of individuals as either HPV negative or HPV positive with a high level of BER expression (

Kaplan–Meier survival analysis of POLB and OGG1 expression. Time-dependent analysis of OGG1 and POLB gene expression (low and high groups) with and without HPV infection. (a) Overall survival of patients with Stage III/IV in OGG1 gene expression in tumor with high
Aberrant BER gene expression is associated with aberrant genomic instability in HNSCC tumor
BER plays a significant role in maintaining the genomic stability of cells. To examine whether aberrant BER gene expression is associated with genomic instability of HNSCC, we examined whether POLB and OGG1 expression changes in HNSCC with and without HPV infection are associated with mutation and chromosomal aberration. We found that the overexpression of OGG1 and POLB overexpression was significantly associated with the increased mutation count of the HNSCC tumors [Figure 3(a)–(d)]. Particularly, mutation count significantly increased in OGG1 and/or POLB overexpressed tumors [Figure 3(a) and (c); *

BER overexpressed tumors accumulate genomic instability. (a) Mutation counts in OGG1 high expressed tumor (
Aberrant expression of BER genes is associated with changes in immune cell infiltration
Increasing evidence suggests that cancer progression is strongly influenced by the host’s immune response, which is represented by immune cell infiltrates. We compared changes in immune and stromal scores between high expression

Landscape of tumor immune microenvironment impact on HNSCC histological tumor grades. (a) Immune, stromal, and ESTIMATE score of the tumor with high
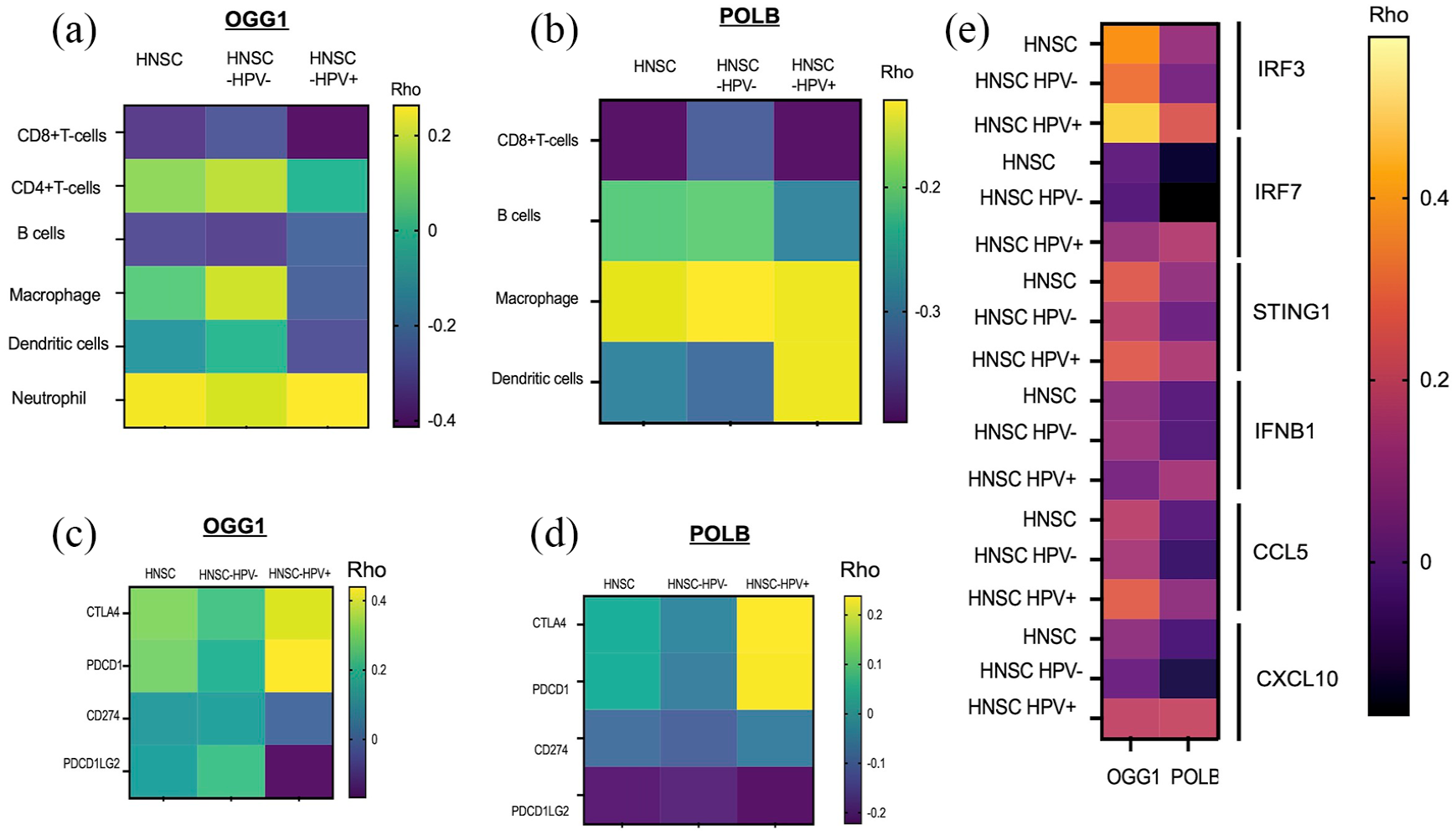
Overexpression of OGG1 and POLB is associated with a low level of infiltration of immune cells to the tumor microenvironment. (a) The expression of OGG1 mRNA
Association of BER with immune checkpoint genes and innate immune signaling landscape in HNSCC
We also analyzed the correlation between
Discussion
POLB and OGG1 are key enzymes for the protection of the genome against DNA damage
Notably, high expression levels of cellular BER factors (OGG1 and POLB) could be a protective response that occurs in cancer to manage HPV oncogenes-dependent DNA damage.
33
However, BER genes commonly mutated or overexpressed in cancer lead to the accumulation of mutations or chromosomal instability.
34
Our
It is known that the tumor immune microenvironment has great implications for cancer progression and susceptibility to immunotherapy. 37 Recently, we have shown that defective BER induces innate immune signaling and Type I interferon response cytokines. 38 Moreover, sustained signaling through interferon-induced pathways has also been associated with resistance to immune checkpoint inhibitors through upregulation of program death ligand 1 (PD-L1), program death ligand 2 (PD-L2), CTLA-4, CIITA, IDO1, CXCL12, and nitric oxide production in tumor cells.39–42 We found that antitumor cells including CD8+ T cells and CD4+ T cells were less abundant in the tumor samples, while several types of immunosuppressive cells were more abundant in the low-immune score patients. Interestingly, overexpression of OGG1 and POLB is associated with low immune cell infiltration and low expression of Type I interferon response genes that are responsible for cytokine-driven immune cell recruitment and inflammation. Several studies have shown that Type I interferon responses result in upregulation of immune checkpoint proteins such as PD-L1, 43 resulting in inhibition of T-cell-mediated cancer cell killing.44–46 Furthermore, CTLA-4 (CD152) is a receptor expressed by both CD4+ T and CD8+ T cells, which inhibit T-cell activation, positively correlated in OGG1 and POLB overexpression. PDCD1 expression, which is a cell surface receptor on T cells and B cells that has a role in regulating the immune system positively correlated with OGG1 and POLB overexpressed tumors. By contrast, the tumors had low expression of PD-L1 and PD-L2 associated with OGG1 and POLB overexpressed tumors suggesting that the BER overexpression likely contributes to immunosuppressive tumor microenvironments. The data demonstrate that OGG1 and POLB negatively correlated with innate immune gene expression and immune cell infiltration, suggesting that OGG1 and POLB may be involved in regulating an antitumor immune response.
Conclusion
Taken together, our silico-data-driven observations suggest that tumors with dysregulated BER factors may enhance tumor immunogenicity that likely enhances immune checkpoint blockade therapy response. In addition, using POLB and OGG1 overexpression may likely contribute to identifying reliable biomarkers to stratify patients for appropriate immune-based treatment. Nevertheless, there is a clear need to understand the mechanisms through which aberrant BER contributes to the immunogenicity of the tumor either through increased tumor mutation burden or concurrent activation of the innate immune signaling
Supplemental Material
sj-jpg-1-tam-10.1177_17588359241248330 – Supplemental material for Dysregulation of base excision repair factors associated with low tumor immunogenicity in head and neck cancer: implication for immunotherapy
Supplemental material, sj-jpg-1-tam-10.1177_17588359241248330 for Dysregulation of base excision repair factors associated with low tumor immunogenicity in head and neck cancer: implication for immunotherapy by Zackary Shpilman and Dawit Kidane in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-jpg-2-tam-10.1177_17588359241248330 – Supplemental material for Dysregulation of base excision repair factors associated with low tumor immunogenicity in head and neck cancer: implication for immunotherapy
Supplemental material, sj-jpg-2-tam-10.1177_17588359241248330 for Dysregulation of base excision repair factors associated with low tumor immunogenicity in head and neck cancer: implication for immunotherapy by Zackary Shpilman and Dawit Kidane in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
