Abstract
Objectives
Studies in different populations have shown that polymorphisms within the CYP1A1 gene play an important role in determining individual susceptibility to lung cancer. However, the data obtained so far have been contradictory within the same or different populations. Few studies have focused on the synergistic effect of CYP1A1 polymorphisms on the susceptibility to lung cancer overall and to different histological subtypes along with the impact of smoking.
Methods
A total of 704 individuals (353 lung cancer patients and 351 controls) were evaluated for CYP1A1 polymorphisms. CYP1A1 genotyping was done by means of PCR-RFLP.
Results
A CYP1A1 mutant genotype was found to be significantly associated with lung cancer (OR = 3.15; 95% CI = 1.75-5.71; p = 0.0001) and this risk was 4-fold higher in case of squamous cell carcinoma (SCC). The CYP1A1 m2 allelic variant was found to be strongly associated with the risk of SCC and adenocarcinoma. The combined “at risk” genotypes of the CYP1A1 m1 and m2 allelic variants were associated with lung cancer risk and this risk was higher in case of SCC (OR = 2.0; 95% CI = 1.97-3.81; p = 0.028). Furthermore, the lung cancer risk was associated with smoking, especially in heavy smokers carrying CYP1A1 variant genotypes. We also observed that heavy smokers with the mutant m1 genotype were at increased risk of SCC (OR = 5.4; 95% CI = 2.4-11.9; p<0.0001). Furthermore, when stratified for smoking dose and histology, the effect was compounded in heavy smokers and SCC (OR = 7.5; 95% CI = 1.8-30.9; p = 0.004).
Conclusions
Polymorphism in the CYP1A1 gene is an important risk modifier for lung cancer.
Introduction
Lung cancer is the leading cause of cancer-related deaths worldwide and the incidence of lung cancer is increasing in developing countries (1). Approximately 63,000 lung cancer patients are reported each year in India (2). It is postulated that tobacco smoke is the major contributor and risk factor for lung cancer and 90% of all cases are due to smoking (3). However, not all smokers develop lung cancer and it has been estimated that fewer than 20% of smokers develop the disease (4). Tobacco smoke contains an abundance of both known and unknown chemicals which produce highly carcinogenic compounds when activated. The process of carcinogen metabolism is mediated by phase I enzymes referred to as cytochrome P450 (CYP). Enzymes of the CYP family are heme-containing enzymes present in both hepatic and extrahepatic tissues (5). Among the various CYP genes, CYP1A1 located on chromosome 15q22-q24 (6) encodes the enzyme aryl hydrocarbon hydroxylase, which is responsible for the activation of polycyclic aromatic hydrocarbons (PAHs) such as benzo(a)pyrene. Once activated, PAH metabolites act as powerful carcinogens which then bind to DNA to form adducts and produce somatic mutations in either tumor suppressor genes or oncogenes and trigger the process of carcinogenesis and hence cancer initiation. The CYP1A1 gene is polymorphic and has been shown to have 15 different allelic variants (7). Four of these variants, designated as m1, m2, m3 and m4, have been widely studied in Asian and Caucasian populations as risk factors for lung carcinogenesis. The m1 and m2 polymorphisms were found to be involved in lung carcinogenesis (8). The CYP1A1 m1 (rs4646903) polymorphism has been associated with elevated enzyme activity due to a single base point mutation at nucleotide position 3801 that results in transition of thymine to cytosine in the 3’ untranslated region. The CYP1A1 m2 polymorphism (rs1048943) involves a base substitution from adenine (A) to guanine (G) at codon 462 in exon 7, resulting in substitution of isoleucine with valine (Ile462Val), which in turn results in an increase in enzyme activity (8).
Several studies have been conducted to evaluate the role of CYP1A1 polymorphisms in the lung cancer risk in Caucasians and Asian populations, particularly Chinese, Japanese, Koreans and Indians; however, the results were inconsistent (9). Many reports from Japanese and Chinese populations have suggested that the m1 and m2 variants of the CYP1A1 gene are associated with an increased lung cancer risk (10). Furthermore, this risk was compounded in case of squamous cell lung carcinoma (SCC) and tobacco smokers. When evaluated in Caucasians the above findings were not confirmed because the prevalence of both m1 and m2 is low in this population (11). However, similar studies conducted in larger samples of mixed American populations did show a positive correlation between the CYP1A1 m1 polymorphism and lung cancer risk (12). A meta-analysis among Asian and Caucasian populations has suggested that the m2 variant allele is associated with an increased risk of lung cancer, particularly SCC (13). These finding indicated that both polymorphisms in the CYP1A1 gene might be important risk factors for lung cancer development (9). In studies done in Indian populations contradictory results were observed. Studies in the Kashmiri population of north India and also a population of north-central India found a significant association between the CYP1A1 m1 and m2 polymorphisms and elevated risk of lung cancer, especially SCC (7, 14). By contrast, a similar study in a northern Indian population showed that the CYP1A1 m1 variant allele was not associated with lung cancer risk but the m2 variant was significantly associated with this risk (15). Most of the studies done so far in Indian populations have been inconclusive. The present study was therefore undertaken for a couple of reasons. Firstly, we sought to provide substantial evidence of the association between CYP1A1 polymorphisms and lung cancer risk with a much larger sample size. Secondly, very few of the conducted studies so far have looked at the combined effect of both polymorphic variants of the CYP1A1 gene in conjunction with the role of smoking and its synergistic interaction with the CYP1A1 variant alleles. The aim of the present work was to further investigate the role of the CYP1A1 m1 and m2 polymorphic variants as genetic modifiers of the risk of developing lung cancer overall and specific histological subtypes in particular. Furthermore, we planned to study the relationship of the CYP1A1 allelic variants in relation to tobacco smoking with a larger sample size in a northern Indian population. We performed association analysis in this case-control study for the 2 genetic variants of the CYP1A1 gene individually and in different combinations, in order to evaluate their cumulative role in modifying the susceptibility to lung cancer.
Material and methods
Sample Collection
The study enrolled 353 lung cancer patients who were recruited from the Department of Pulmonary Medicine, Post Graduate Institute of Medical Education and Research (PGIMER), Chandigarh, India. The study protocol was reviewed and approved by the ethics committee of PGIMER. Informed written consent was obtained from all participants or their representatives. All recruited patients were histopathologically diagnosed as having non-small cell lung carcinoma (NSCLC) or small cell lung carcinoma (SCLC). Patients having a prior history of cancer were excluded from the study. There were no age, gender, smoking, histology or TNM stage restrictions. The control group of the study consisted of 351 unrelated volunteers having no lung cancer history at the time of blood collection; they entered the hospital for health check-ups. The controls were pair-matched for sex, age (±10 years) and smoking parameters in order to avoid any sampling bias. Each participant filled in a detailed questionnaire with the help of a trained interviewer. The questionnaire included information on demographic and smoking characteristics such as smoking of beedies/cigarettes, etc. Individuals having regular smoking habits were classified as smokers. They were further stratified into light and heavy smokers on the basis of pack years (PYs), which were calculated by the formula: [(cigarettes or beedies per day/20)*years smoked]; PY≤25 was defined as light smokers and PY>25 as heavy smokers. Medical information including tumor histology, TNM classification, clinical staging, primary tumor size, involvement of lymph nodes, and metastasis was obtained from the medical records of the hospital.
DNA Extraction and Genotyping
Approximately 3-5 mL of venous blood was drawn from cases and controls. DNA was isolated according to the protocol of Field et al (16) with slight modifications. Briefly, the collected blood was washed 3 times with washing buffer containing 320 mM sucrose, 1.5% Triton X-100, 20 mM Tris-HCl (pH 8.0), and 5 mM MgCl2 to remove red blood cells. The obtained white blood cells were lysed by resuspending them into lysis buffer with 400 mM Tris-HCl (pH 8.0), 200 mM NaCl, 80 mM EDTA, 1% SDS, and 100 μg/mL proteinase K and then incubated overnight at 45°C. The lysed cells were subjected to a deproteinization step using phenol, chloroform and isoamyl alcohol (25:24:1).The aqueous phase was transferred and DNA was precipitated by adding an equal amount of chilled isopropanol and recovered by centrifugation at 10,000 rpm for 5 minutes and then washed with 70% ethanol. The DNA was resuspended in 200 μL of sterile Tris-EDTA buffer and the quality and quantity of DNA were determined by agarose gel electrophoresis and spectrophotometric analysis using a NanoDrop ND-1000 spectrophotometer. DNA was diluted to 100 ng/μL and stored at -20°C for further use as a template.
Genotyping of CYP1A1 m1 Polymorphism by PCR-RFLP
To analyze the T to C polymorphism in the CYP1A1 gene at the m1 site a polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) technique was used followed by digestion with Msp1 restriction enzyme. PCR amplification of 899-bp-long DNA fragments was done in a 25-μL reaction mixture containing 300 ng of DNA template, 0.5 μmol each of forward primer 5-GGCTGAGCAATCTGACCCTA-3 and reverse primer 5-TAGGAGTCTTGTCTCATGCCT-3 (17) with 1.5 mM MgCl2, 0.2 mM dNTPs, 1×PCR buffer and 1 U Taq DNA polymerase. PCR was performed with an initial denaturation step of 5 minutes at 95°C followed by 30 cycles of 1 minute at 94°C, 30 seconds at 60°C as an annealing step, and 45 seconds at 72°C followed by a final extension step of 5 minutes at 72°C. The amplification was confirmed using agarose gel electrophoresis. The PCR product was then subjected to restriction digestion with 5 U Msp1 enzyme (New England Biolabs). After digestion the samples were electrophoresed on a 2.5% agarose gel and visualized in UV light. PCR fragments of 899 bp were digested into 2 fragments of 693 bp and 206 bp in case of a mutant genotype (CC), while an undigested fragment of 899 bp was indicative of a wild-type genotype (TT). The heterozygous group was represented by fragment sizes of 899, 693 and 206 bp, respectively, as shown in Figure 1. To ensure quality control, genotyping was performed without knowledge of the subjects’ case/control status and a 15% random sample of cases and controls was genotyped twice by different persons; the reproducibility was 100%.
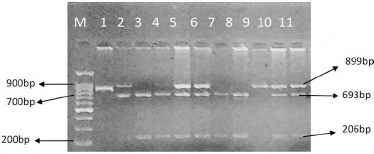
Ethidium–bromide-stained agarose gel showing detection of the single nucleotide polymorphism in the Msp1 polymorphism of the CYP1A1 gene. Lane M shows the 100 bp molecular weight marker, lane 1 shows the 899 bp uncut product, lane 9 represents the 899 bp homozygous wild-type genotype (TT), lanes 2, 4-6, 10 and 11 show the heterozygous genotype (TC) and lanes 3, 7 and 8 represent the mutant genotype (TT) of the Msp1 polymorphism.
CYP1A1 m2 Genotyping by PCR-RFLP
Genotyping of CYP1A1 m2 was performed with the PCR-RFLP method according to the protocol of Oyama et al (8). The product of the amplicon that was generated by PCR was 187 bp in size; it was amplified using the following primer sequences: 5-GAACTGCCACTTCAGCTGTCT-3 and 5-GAAAGACCTCCCAGCGGTCA-3. The PCR conditions were as follows: denaturation at 95°C for 1 minute, annealing at 55°C for 30 seconds, and extension at 72°C for 45 seconds for 30 cycles with final extension at 72°C for 5 minutes. Restriction digestion of the PCR product was done using 5 U of HincII enzymes (New England Biolabs). The digested PCR products were then separated on a 10% PAGE gel and silver staining was done to observe the digested bands. The restriction digestion pattern observed in 10% acrylamide gel was as follows: a wild-type genotype (AA) gave 2 bands of 139 and 48 bp, a heterozygous genotype (AG) gave 4 bands of 139, 120, 48 and 19 bp, and a mutant genotype (CC) gave 3 bands of 120, 48 and 19 bp, as shown in Figure 2.
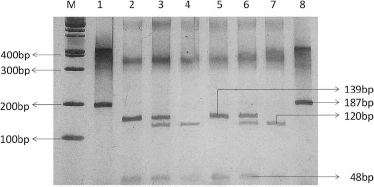
Polyacrylamide gel electrophoresis (PAGE) showing CYP1A1 Ile462Val genotype analysis by PCR-RFLP. Lane M shows the 100 bp molecular weight marker, lanes 1 and 8 represent the uncut 187 bp product, lanes 2 and 5 represent the 139 and 48 bp homozygous wild-type genotype (AA), lanes 3 and 6 represent the 139, 120, 48 and 19 bp heterozygous genotype (AG), lanes 4 and 7 show the 120, 48 and 19 bp homozygous mutant genotype (GG) of the m2 polymorphism.
Statistical Analysis
Differences in the distribution of demographic and genotypic characteristics of cases and controls were evaluated using the MedCalc statistical software, version 15.6.1, and SPSS version 20.0. We used the chi-square test for categorical data and Student's t-test for continuous variables. To calculate the genotyping frequency of both polymorphisms the Hardy-Weinberg equilibrium (HWE) (p2 + + 2pq + q2 = 1; where p is the frequency of the wild-type allele and q is the frequency of the variant allele) was used in both cases and controls by means of the chi-square test. Haplotype analysis was done for the 2 polymorphisms studied and the D’ value and r2 were calculated with the SHEsis software (18). To identify the risk of lung cancer due to the allelic variants, the odds ratios (ORs) along with 95% confidence intervals (CIs) and p values at the level of significance <0.05 were calculated using logistic regression analysis with adjustment for possible confounders such as age and PYs as continuous variables and gender as a nominal variable. In addition to overall association analysis, stratified analysis was performed to estimate the risk for subgroups on the basis of smoking status, tumor histology and gender.
Results
Demographic Characteristics and Clinical Data of Study Participants
The demographic characteristics and clinical data of the study participants are shown in Table I. A total of 353 cases and 351 controls were enrolled, with an age range of 28-86 years and 19-83 years, respectively. There was no significant difference in the distribution of men and women between groups, suggesting adequate matching (p = 0.804). There were more smokers in the case group than the control group. When stratified on the basis of PYs, there was a significant difference in distribution between PYs of cases and controls. Lung cancer cases were subdivided on the basis of histology into SCC, adenocarcinoma (ADC) and SCLC. Cases were further classified according to TNM staging. Tumor size varied from T1 to T4 and lymph node status from N0 to N3. Less than half of the lung cancer patients for whom TNM data were available had distant metastases (Tab. I).
Demographic characteristics and clinical data of study subjects
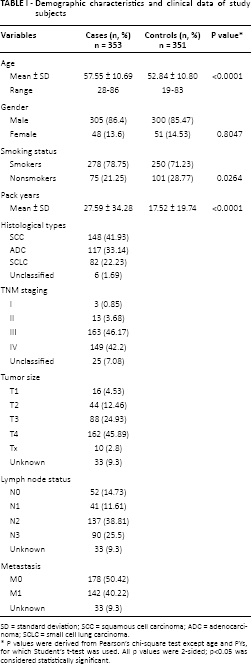
SD = standard deviation; SCC = squamous cell carcinoma; ADC = adenocarcinoma; SCLC = small cell lung carcinoma.
P values were derived from Pearson's chi-square test except age and PYs, for which Student's t-test was used. All p values were 2-sided; p<0.05 was considered statistically significant.
Distribution of Allele and Genotype Frequencies of CYP1A1 m1 and m2 and their Association with Risk of Lung Cancer Overall and Histological Subtypes
The allele and genotype frequencies were calculated according to HWE analysis. The data showed that the allele frequency of CYP1A1 m1 in cases (χ2 = 0.76; df = 1; p = 0.38) and controls (χ2 = 3.05; df = 1; p = 0.08) was in HWE, suggesting that there was no population stratification and no sample bias. Similarly, in CYP1A1 m2 the allele frequencies of the control group followed HWE (χ2 = 1.81; df = 1; p = 0.17). The minor allele frequency (MAF) of the C allele of the m1 polymorphism was 0.35 in cases and 0.26 in controls, whereas MAF of the G allele of the m2 polymorphism was 0.18 in cases and 0.11 in controls. As shown in Table II the genotype frequencies and ORs of CYP1A1 m1 and m2 were calculated after adjusting for age, sex and smoking status using logistic regression analysis to evaluate the lung cancer risk. In the CYP1A1 m1 polymorphism the frequency of the heterozygous variant allele was the same in patients and controls, while the mutant genotype (CC) was overrepresented in patients (13% in patients vs. 5% in controls). Taking the homozygous wild-type genotype (TT) as the reference, we observed a 3-fold increased risk of lung cancer in individuals carrying the homozygous mutant genotype (OR = 3.15; 95%CI = 1.75-5.71; p = 0.0001). Such an association was not observed in individuals with a heterozygous (TC) genotype. When both the mutant and heterozygous alleles were combined as a single genotype a marginal risk was observed (OR = 1.4; 95% CI = 1.03-1.91; p = 0.02). Similarly, in the CYP1A1 m2 polymorphism it was observed that the frequency of the heterozygous allele was higher in cases than controls (32% vs. 13%). Taking the wild-type genotype (AA) as the reference, it was observed that individuals with the heterozygous genotype (AG) had a 2-fold increased risk of lung cancer (OR = 2.3; 95% CI = 1.6-3.3; p<0.0001) compared to individuals with the wild-type genotype (AA). The mutant genotype was not associated with any tendency towards lung cancer development. Furthermore, as shown in Table II, when patients were stratified according to the different histological types of lung cancer, SCC patients with the mutant genotype (CC) of CYP1A1 m1 had a 4-fold increased risk (OR = 4.41; 95% CI = 2.19-8.86; p = 0.0001), whereas a 2-fold risk was observed in ADC patients, (OR = 2.2; 95% CI = 0.9-5.0; p<0.05). No such association was observed in SCLC. In the CYP1A1 m2 polymorphism, a significantly elevated risk was seen in SCC patients with the heterozygous genotype (AG) (OR = 2.65; 95% CI 1.68-4.18, p = 0.04) as well as ADC patients (OR = 2.5; 95% CI = 1.54-4.06; p = 0.0001). No significant association was observed in SCLC patients for both polymorphic variants.
Distribution of allele and genotype frequencies of CYP1A1 m1 and m2 and their association with risk of lung cancer and histological subtypes
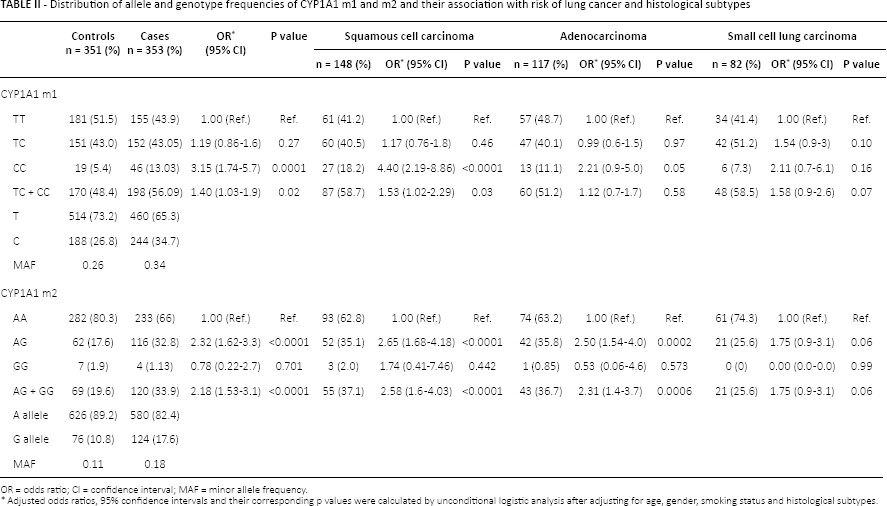
OR = odds ratio; CI = confidence interval; MAF = minor allele frequency.
Adjusted odds ratios, 95% confidence intervals and their corresponding p values were calculated by unconditional logistic analysis after adjusting for age, gender, smoking status and histological subtypes.
Haplotype and Linkage Disequilibrium Analysis in CYP1A1 m1 and m2 Polymorphisms
We obtained the inferred haplotypes of CYP1A1 by using the SHEsis platform. The haplotype approach revealed 4 possible haplotypes in both patients and controls, as shown in Table III. Haplotypes with frequencies <0.03 among cases and controls were omitted from the analysis. The haplotype T-A was considered to be the reference carrying wild-type alleles. The frequency of the other 3 haplotypes (C-A, C-G, T-G) and their distribution among cases and controls is shown in Table III. Comparison of overall haplotype distribution profiles revealed a statistically significant difference between cases and controls (global test: χ2 = 17.91, df = 3, p = 0.0004). The T-G haplotype was associated with an increased risk of lung cancer (p = 0.046). No associations were observed for any of the other haplotypes. In haplotype analysis for the 2 loci of the CYP1A1 polymorphism, the linkage disequilibrium between m1 and m2 was D’ = 0.592, r2 = 0.131.
Distribution of CYP1A1 haplotypes among lung cancer patients and controls *
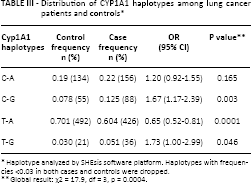
Haplotype analyzed by SHEsis software platform. Haplotypes with frequencies <0.03 in both cases and controls were dropped.
Global result: χ2 = 17.9, df = 3, p = 0.0004.
Association between Combined CYP1A1 m1 and m2 Genotypes and Risk of Lung Cancer Overall and Histological Subtypes
We also evaluated the combined effect of CYP1A1 m1 and m2 on the lung cancer risk, as shown in Table IV. There were 192 cases and 214 controls having a common genotype for both the m1 and m2 sites of the CYP1A1 gene. Among the lung cancer cases, 66.67% carried homozygous wild-type alleles for both m1 and m2, 31.77% had heterozygous alleles for both polymorphic sites, and 1.56% had mutant alleles. Compared to the lung cancer cases the control group had a higher frequency (78.5%) of both homozygous wild-type alleles, whereas the frequency of heterozygous alleles for both CYP1A1 polymorphic sites was lower (20% vs. 31.77%). Individuals having heterozygous alleles for both polymorphic sites were found to have a 2-fold higher risk of lung cancer, which was found to be significant (OR = 1.99; 95% CI = 1.2-3.1; p = 0.004). No significant association was observed in subjects having homozygous mutant alleles for both polymorphisms. Furthermore, when patients were stratified according to histological subtypes, a 2-fold increased risk was found for SCC (OR = 2.0; 95% CI = 1.97-3.81; p = 0.028) as well as SCLC (OR = 2.2; 95% CI = 1.04-4.7; p = 0.037) but there was no significant association with ADC.
Association between combined CYP1A1 m1 and m2 genotypes and risk of lung cancer and different histological subtypes
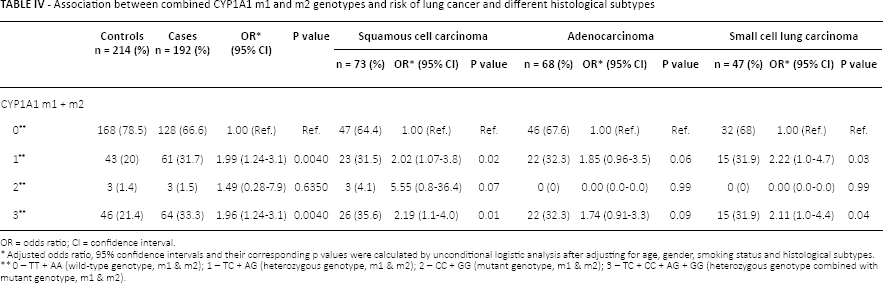
OR = odds ratio; CI = confidence interval.
Adjusted odds ratio, 95% confidence intervals and their corresponding p values were calculated by unconditional logistic analysis after adjusting for age, gender, smoking status and histological subtypes.
0 – TT + AA (wild-type genotype, m1 & m2); 1 – TC + AG (heterozygous genotype, m1 & m2); 2 – CC + GG (mutant genotype, m1 & m2); 3 – TC + CC + AG + GG (heterozygous genotype combined with mutant genotype, m1 & m2).
Interaction of CYP1A1 Genotypes and Smoking Status with Risk of Lung Cancer
The risk of lung cancer in relation to both the m1 and m2 polymorphisms of the CYP1A1 gene was further examined after stratification based upon smoking status and cumulative smoking dose as shown in Table V and supplementary Table S1 (Interaction of CYP1A1 genotypes and smoking status after stratification to pack years. Available online at www.biological-markers.com). It was observed that smokers with the mutant genotype (CC) for CYP1A1 m1 had a 3-fold increased risk of lung cancer (OR = 3.44; 95% CI = 1.66-7.13, p = 0.0009), which was found to be highly significant as compared to smokers with the wild-type genotype (TT), but there was no significant association in nonsmokers. Similarly, in the CYP1A1 m2 polymorphic site, smokers with the heterozygous (Ile/Val) genotype had a 2-fold increased risk of lung cancer (OR = 2.44; 95% CI = 1.60-3.73, p<0.0001), while no such association was observed in nonsmokers.
Interaction of CYP1A1 genotypes and smoking status with risk of lung cancer
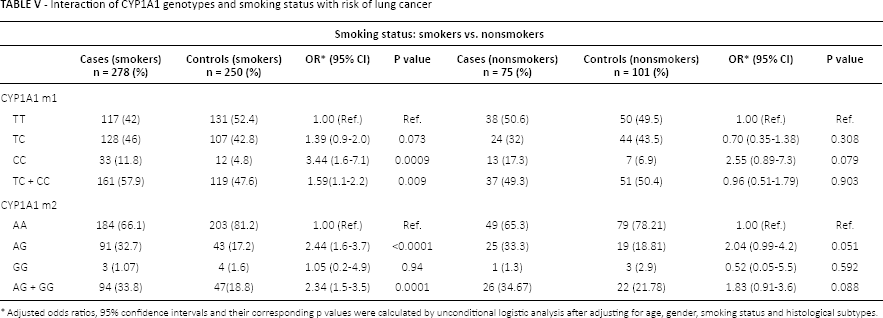
Adjusted odds ratios, 95% confidence intervals and their corresponding p values were calculated by unconditional logistic analysis after adjusting for age, gender, smoking status and histological subtypes.
Further stratifying the smokers into light and heavy according to PYs (as shown in supplementary Table S1) we found that in light smokers with ≤25 PYs and carrying the mutant genotype (CC) for CYP1A1 m1 the OR for lung cancer was 2.6 (95% CI = 1.3-5.2; p = 0.004) and was statistically significant. The OR increased to 5.6 (95% CI = 1.5-20.9; p = 0.009) in heavy smokers who carried the homozygous mutant genotype (CC). Similarly, when analyzed for the CYP1A1 m2 polymorphism, light smokers who carried the heterozygous genotype (Ile/Val) showed an approximately 2-fold increased risk of lung cancer (OR = 1.9; 95% CI = 1.28-3.1; p = 0.002), while among heavy smokers a 3-fold increased lung cancer risk was observed for individuals carrying the heterozygous genotype (OR = 3.0; 95% CI = 1.6-5.9; p = 0.0008).
Interaction of CYP1A1 Genotypes and Tobacco Smoke According to Histological Subtypes of Lung Cancer
The interaction of the CYP1A1 polymorphisms and tobacco smoke was also assessed separately for SCC, ADC and SCLC, as shown in supplementary Table S2 (Interaction of CYP1A1 genotypes and tobacco smoke according to histological subtypes of lung cancer. Available online at www.biological-markers.com). Compared with individuals with the CYP1A1 m1 wild-type genotype (TT) who were smokers, it was observed that SCC patients carrying the mutant genotype (CC) for m1 had an elevated risk of lung cancer (OR = 5.4; 95% CI = 2.4-11.9; p<0.0001) that was found to be significant, whereas no such association was observed in either ADC or SCLC. In the case of the CYP1A1 m2 polymorphism, SCC patients who were smokers showed an approximately 3-fold increased lung cancer risk with the heterozygous genotype (AG) (OR = 2.8; 95% CI = 1.72-4.6; p<0.0001). Similar results were observed in ADC (OR = 2.2; 95% CI = 1.2-4.2; p = 0.009) and SCLC (OR = 2.1; 95% CI = 1.15-4.0; p = 0.015), but the effect was less pronounced than in SCC patients. In case of nonsmokers no significant association with risk was observed in any histological subtypes with the variant CYP1A1 m1 genotype. However, for the CYP1A1 m2 polymorphism ADC patients having the heterozygous allele (Ile/Val) showed an approximately 3-fold increased risk (OR = 2.9; 95% CI = 1.3-6.5; p = 0.008) (supplementary Table S3 - Interaction of CYP1A1 genotypes and nonsmokers risk towards lung cancer risk on the basis of histology. Available online at www.biological-markers.com).
In the present study we also evaluated the role of combined polymorphic sites of CYP1A1 along with smoking status (supplementary Table S4 - Interaction of combined CYP1A1 genotypes and its association with lung cancer risk stratified according to smoking status and pack years. Available online at www.biological-markers.com). It was observed that individuals who were smokers and carried the variant allele of each polymorphic site had an elevated risk of lung cancer (OR = 2.3; 95% CI = 1.31-4.01; p = 0.002) compared to individuals with the same genotype who were nonsmokers (OR = 1.3; 95% CI = 0.51-3.4; p = 0.554). Furthermore, the smokers were stratified into light and heavy smokers according to PYs. Heavy smokers had a higher lung cancer risk when both variant alleles of the 2 polymorphic sites of CYP1A1 were combined as a single genotype and compared with the combined wild-type genotype (OR = 2.8; 95% CI = 1.24-6.5, p = 0.013).
We also stratified light and heavy smokers on the basis of histological subtypes (supplementary Table S5 - Interaction of CYP1A1 m1 and m2 genotypes for light smokers after stratifying on basis of lung cancer histology. Available online at www.biological-markers.com). In light smokers we observed that, as compared to the CYP1A1 m1 wild-type genotype, the odds of SCC for the mutant CYP1A1 genotype was 3.9 (95% CI = 1.7-8.8, p = 0.001), whereas no significant association was seen for the other histological subtypes. Similarly, in the case of the m2 polymorphism, ADC patients who were classified as light smokers based on PYs had a high lung cancer risk (OR = 2.5; 95%CI = 1.49-4.4; p = 0.0007) compared to subjects with the same smoking exposure but a wild-type (Ile/Ile) genotype. In case of heavy smokers (supplementary Table S6 - Interaction of CYP1A1 m1 and m2 genotypes for heavy smokers after stratifying on basis of lung cancer histology. Available online at www.biological-markers.com), a strong positive association was observed for SCC and SCLC patients in whom the smoking dose was PY>25 and who carried the mutant genotype of the CYP1A1 gene (CC) (OR = 7.5; 95% CI = 1.8-30.9; p = 0.004) as compared to SCC subjects with the wild-type genotype (TT). In case of the CYP1A1 m2 polymorphism, among heavy smokers, SCC patients showed a 3-fold increased lung cancer risk with the heterozygous allele (Ile/Val) (OR = 3.5; 95% CI = 1.69-7.5; p = 0.0008), whereas no association was observed in ADC and SCLC.
Discussion
Lung cancer initiation can be mediated by exposure to carcinogens such as PAHs and aromatic amines that are present in cigarette or beedi smoke. The CYP1A1 gene is polymorphic in nature and might play an important role in lung carcinogenesis and hence lung cancer. Several studies have been conducted both in Asian and Caucasian populations to assess whether genetic variability in the CYP1A1 gene influences the susceptibility to lung cancer. These studies have shown different associations between the polymorphic genotypes of CYP1A1 and the lung cancer risk. However, the results so far have been inconsistent between different populations and also within the same populations. This discrepancy might be due to varying factors: for example, studies were carried out in different ethnic populations with different heredity backgrounds, lifestyles, and smoking habits. The different histological subtypes of lung cancer might also account for inconsistency in the results.
In our study we observed a significant difference in the frequency distribution of the mutant m1 genotype of CYP1A1 between cases and controls. Furthermore, our data clearly demonstrated a significant association between the m1 genotype and lung cancer risk, and this was more pronounced in SCC. The present findings are consistent with previous studies conducted in various ethnic groups. Studies in northern Indian populations have shown an elevated risk of lung cancer in carriers of a CYP1A1 m1 (TC) variant allele (14, 15, 19). Similarly, Sreeja et al (20) reported a high frequency of a variant CYP1A1 m1 genotype in lung cancer cases in a southern Indian population (20). Thus, the data of this study revealed an association between genetic polymorphisms in the CYP1A1 locus and lung cancer risk among Indians. Also previous studies conducted in other Asian populations such as Chinese (10), Japanese (21-22-23), Mongolians (24) and Koreans (25) showed a positive association between m1 polymorphic sites and lung cancer risk. However, some studies in Asian populations have reported a lack of association between CYP1A1 m1 genotype and lung cancer risk (26-27-28-29). The results obtained in the current study are inconsistent with those reported by studies in Caucasians; this might be attributed to a very low frequency of variant CYP1A1 alleles (30, 31).
In contrast to the significant association of the CYP1A1 mutant m2 genotype with lung cancer previously found in Asians (32-33-34), no significant association was observed in the present study. It has been reported that the frequency of the mutant (Val/Val) genotype is high in Japanese and Chinese compared to Indians (13). In our study we found that the heterozygous genotype for CYP1A1 m2 (Ile/Val) was positively associated with lung cancer risk. The finding is consistent with other Indian studies that also reported an association of the heterozygous genotype of CYP1A1 m2 with lung cancer risk, as was also studied in an Egyptian population (7, 14, 20, 35). However, Sobti et al reported a high frequency of the mutant genotype (15). It should be noted that in Caucasians the frequency of the mutant genotype for CYP1A1 m2 was found to be lower than in East Asians (36).
A plethora of data points to the fact that, since lung cancer is histologically heterogeneous in nature, it is imperative that the different pathological subtypes of lung cancer should be studied separately in relation to the polymorphisms present within in the CYP1A1 gene. When lung cancer was subgrouped according to histology, it was observed that the CYP1A1 m1 variant polymorphism was significantly associated with SCC and not ADC or SCLC. Similarly, our data suggests a significant role of the CYP1A1 m2 heterozygous genotype in the risk of SCC. These results are consistent with those obtained by previous studies (12, 14, 37, 38) that found a significant association between CYP1A1 polymorphisms and an elevated risk of SCC but not ADC of the lung. No significant association was found between the CYP1A1 polymorphisms and SCLC. Some previous studies indicated no significant risk for SCC related to the CYP1A1 m1 polymorphism in Caucasian populations, but one study found that the m1 polymorphism was associated with ADC (39). A study conducted in a Spanish population revealed that the m2 polymorphism contributes to an increased risk of susceptibility to SCC and SCLC but not ADC (40).
Our study has also evaluated the combined effect of both polymorphic variants of the CYP1A1 m1 and m2 genotypes on the susceptibility to lung cancer. We inferred from the results that individuals who carried the variant alleles of both m1 and m2 were more susceptible to lung cancer and this risk was found to be 2-fold in cases of SCC. Our data are consistent with those of other studies (11, 41). Since CYP1A1 is a key enzyme involved in the xenobiotic metabolism, evidence from expression studies has shown that the variant allele of the CYP1A1 m1 site might affect the inducibility of the enzyme by affecting the transcriptional control elements. Furthermore, the m2 polymorphism might result in increased activity of the enzyme because of an isoleucine-to-valine substitution in the heme-binding region, which then enhances the enzyme activity (42). Individuals having more risk alleles therefore have an enhanced ability to convert these procarcinogens into carcinogens.
We found that the CYP1A1 m1 polymorphism was significantly related to the risk of lung cancer especially in individuals having the mutant genotype (p = 0.0009) and a smoking habit, while patients who were lifetime smokers and carried the heterozygous variant allele of the CYP1A1 m2 gene also had a high lung cancer risk (p<0.0001). These findings are consistent with previous studies where a strong association of lung cancer with variant CYP1A1 genotypes was found in smokers (7, 43-44-45) but not in nonsmokers. Biological and epidemiological data suggest that among smokers the different lung cancer histologies should be analyzed separately (46). When stratifying by histological subtypes of lung cancer, we observed that patients who were smokers and had variant genotypes for both CYP1A1 polymorphic sites had a markedly increased susceptibility to SCC, while no association with SCC was observed for nonsmokers. By contrast, we observed that patients with either the mutant CYP1A1 m1 or heterozygous m2 genotype who were lifetime nonsmokers were at the highest risk of developing ADC. It is plausible that other carcinogenic factors may be involved in the development of ADC among nonsmokers; these factors probably may also be substrates for CYP1A1. The findings were in line with those in a Taiwanese population (47), where it was observed that an elevated risk for SCC and not ADC was significantly associated with polymorphisms in the CYP1A1 and epoxide hydroxylase genes. Our current findings corroborate the evidence of previous studies (10, 14, 15, 48) that CYP1A1 polymorphisms are important determinants of the susceptibility to tobacco-induced lung cancer. We also stratified the smoking status on the basis of PYs, where we included both light smokers (PY≤25) and heavy smokers (PY>25). We observed a 5-fold higher lung cancer risk for individuals who were heavy smokers and carried the homozygous mutant genotype in case of the CYP1A1 m1 polymorphism, whereas heavy smokers who carried the heterozygous variant allele for CYP1A1 m2 had a 3-fold increased risk, which was significant (p = 0.0008). We also observed that in heavy smokers either the mutant m1 or the Ile/Val m2 genotype was strongly associated with the risk of SCC compared to ADC or SCLC. These findings are in agreement with those of Song et al (10) and Dong et al (49) in Chinese populations. However, Sugimura et al (39) reported that light smokers were at an increased risk of lung cancer when they carried the mutant m2 genotype of the CYP1A1 gene. Garcia-Closas et al (50) could not find substantial evidence of the effect of tobacco PYs on the lung cancer risk in relation to the CYP1A1 m1 genotypes (50).
In the combined m1 and m2 polymorphism an increased risk of lung cancer was present in heterozygous allelic variants in case of heavy smokers only, whereas no association was observed in light smokers. This result was consistent with findings in a Kashmiri population (14). It was also observed that patients who were heavy smokers and carried the variant alleles either for m1 or m2 were at higher risk of developing SCC than light smokers.
A limitation of our study was sample size: although the sample was larger than that of previous studies in Indian populations, the number of subjects in some of the subcategories was rather small, which can be considered a drawback. Secondly, the findings of this study were only from a northern Indian population, and it is uncertain whether they are generalizable to the population of other areas of India. Thirdly, the controls were recruited from a health center, which could entail a selection bias compared to a population-based sample. However, potential confounding factors may have been minimized by matching the controls to the cases for age, sex and area of residence, and by further adjustment for the confounding factors in data analyses.
Conclusion
Our study demonstrated that the CYP1A1 m1 (rs4646903) and m2 (rs1048943) polymorphisms are important factors contributing to increased susceptibility to and pathological development of lung cancer in a northern Indian population. Validation of these findings with functional evaluation and larger studies with more rigorous study designs are needed. Studies are also required to assess the carcinogenic role of passive smoking, diet and cooking, pollution, occupational exposure and other environmental factors in addition to genetic factors, so as to establish a risk profile for each individual or subgroup in affirmation of the prevention scope.
Footnotes
Abbreviations
Financial support: This work was supported by a grant from the Indian Council of Medical Research, New Delhi, India (grant No. 5/13/126/2011/NCD-III).
Conflict of interest: None declared.
