Abstract
Objective
The primary purpose of this paper is to investigate the relationship between the microRNA 146a (miR-146a) and the proliferation of cells occurring in glioblastoma multiforme. The secondary purpose of the paper is to investigate abnormalities of expression in miR-146a.
Methods
A real-time PCR assay was used to investigate the abnormal expression of miR-146a in glioma and adjacent tissue. Lipofection was used to transfect a mimic of miR-146a and induce the upregulation of miR-146a. Real-time PCR was used to observe the expression level of miR-146a. A cell viability analysis was conducted using MTT. A luciferase report vector was used to identify potential targets for miR-146a.
Results
The miR-146a component was found to be downregulated in glioma tissue compared with adjacent nontumor tissue (p<0.05). The upregulation of miR-146a in glioma cells through miR-146a mimic transfection led to reduction of cell viability and to an increase in the percentage of apoptosis. Notch1 was the name of the potential targeted gene for miR-146a in glioma.
Conclusions
The study found that the presence of miR-146a potentially affected the proliferation of glioma cells by regulating the rate of Notch1 expression.
Introduction
Glioma is a common disorder affecting the central nervous system. It is a disease of the glia cells which surround and support neurons (1). The disorder accounts for approximately 80% of primary malignant brain tumors (2, 3). Glioma tissues present with different gene expression patterns compared to the surrounding tissue (4, 5). Many of these differentially expressed genes might participate in tumor genesis and tumor development (6). MicroRNAs (miRNAs) are defined as regulatory RNA molecules which play a role in regulating gene expression post-transcription. MiRNAs are thought to play a variety of important roles in physiological and pathological processes not limited to apoptosis, cell differentiation, development, and signal transduction (7). At present, 2,588 miRNAs have been identified by the scientific community (miRBase Release 21). There is an impressive body of evidence showing that miRNAs act as oncogenes or tumor suppressors and can work to regulate cell proliferation, apoptosis, and other forms of biogenesis in cancer (8-9-10).
Alterations of miRNA patterns and functions have been reported in gliomas (11, 12). Among the miRNAs that have been related to glioma, miR-146a has attracted the greatest attention due to its ability to restrict the formation of glioma stem-like cells by regulating the Notch1 pathway (13). Hu et al (14) have reported an association between miR-146a single nucleotide polymorphisms (SNPs) and the increased risk of developing glioma in a northeastern Chinese population.
Clearly, miR-146a might hold great potential in future molecular therapies targeting glioblastoma multiforme (GBM). Previous research has revealed the association of Notch1 and miR-146a in GBM cell lines and mice models, but no direct clinical evidence for miRNA-146a and its functional target genes has been recognized. This paper investigated the role of miR-146a in GBM patients and the possible involved signaling pathway.
Materials and methods
Patients and Tissue Samples
Fifty-one resected GBM tissue samples were collected from July 2010 to July 2013 from Xinxiang Central Hospital, affiliated with Xinxiang Medical University. Samples were taken from 26 males and 25 females, ranging in age from 30 to 72 years (median 51 years). None of the patients sampled had undergone chemotherapy or radiotherapy prior to having surgery (Tab. I). Tissue samples adjacent to target tissue were collected at the same time. Three neuropathologists performed histopathological diagnoses of all tissue samples. This study was granted approval by the ethics committee of Xinxiang Central Hospital. Specimens were handled according to ethical and legal standards in practice in this area. Proof of written informed consent was obtained from all participants.
Demographics of study patients
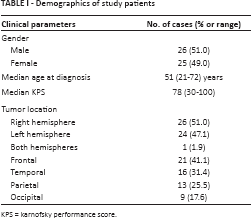
KPS = karnofsky performance score.
Cell Culture
The human GBM cell lines U87, U251, and A172 were purchased from the Japanese Cancer Resources Bank. The human normal brain cell line SVG p12 was purchased from American Type Culture Collection (ATTC). Cells were cultured in Dulbecco's modified Eagle's medium and supplemented with fetal bovine serum. The cells were incubated at 37°C in an environment consisting of 95% air and 5% CO2.
Transient Transfection of miR-146a Mimics or Inhibitors
GenePharma (Shanghai, China) was used as the source for miR-146a mimics, inhibitors, and negative miR controls. Lipofectamine 2000 (Invitrogen) was used to perform transfections. Cells were transfected with miR-146a inhibitors, controls, or mimics into a final 50-nM solution.
Luciferase Assay
The instructions described previously (13) provided a template for the construction of luciferase reporter vectors for the miR-146a target, Notch1. The cells in question were transfected with 0.8 µg of luciferase reporter vector including 100 nM of miR-146a mimic, inhibitor, or miR control (Invitrogen). Luciferase assays were performed using the dual luciferase reporter assay system developed by Abcam (USA).
Cell Proliferation Assay
Cell proliferation was tested with a commercial MTT assay kit (Thermo Fisher Scientific) according to the manufacturer's instructions. Cells were seeded briefly at a density of 1 × 104 per well across 96-well plates. These cells were then transfected with miR-146a inhibitors, control, or mimics. The proliferation of cells was measured 48 hours later by cell counting.
Real-Time PCR
Total RNA was extracted with the help of the TRIzol reagent (Invitrogen) and stored at -80°C. Quantitative real-time PCR (qRT-PCR) was conducted in a 2-step fashion as per the manufacturer's instructions (Takara). The testing of real-time PCR required the use of the ABI 7900 fast sequence detection system (Applied Biosystems) as well as the SYBR green fluorescent dye. The following PCR was used: 95°C for 10 minutes, 40 cycles of 95°C for 15 seconds, 60°C for 1 minute, and finally an elongation step at 72°C for 30 seconds. To control for variations in the total quantity of miRNA in each sample, an internal reference (the U6 snRNA) was used. The differential expression was evaluated with the 2-(∆∆Ct) method. Each RNA sample was analyzed 3 times with the following primers: 1) human Notch1 primers: forward 5′-GAGGCGTGGCAGACTATGC-3′; reverse 5′-CTTGTACTCCGTCAGCGTGA-3′; 2) human miR-146a primers: forward 5′-TGTAACCAGAGAGCGGGATGT-3′; reverse 5′-TTTTGGCATAACTAAGGCCGAA-3′; 3) human U6 snRNA primers: forward 5′- CTCGCTTCGGCAGCACA-3′; reverse 5′- AACGCTTCACGAATTTGCGT-3′.
Measurement of Apoptosis by Flow Cytometry
Apoptosis kits were purchased from Thermo Fisher Scientific (Thermo Fisher Scientific, Boston, MA, USA). Single cell suspensions were collected from GBM and control tissue. Harvested cells were washed in cold PBS and then treated with annexin binding buffer. Afterwards, the suspension cells were fixed in 70% cold ethanol and treated with 10 g/L RNase. Subsequently, 5 μL of Alexa Fluor 488 annexin V and 1 μL of 100 μg/mL working solution were added to each 100 μL of cell suspension. After 15 minutes of incubation at room temperature, 400 μL of annexin binding buffer was added to the mix and mixed in gently with the rest of the sample. The sample was kept on ice for further testing. For this experiment, the CellQuest software was used to quantify and analyze both cells with early apoptotic signals (stained with annexin V) and cells with late death signals (stained with PI).
Statistical Analysis
SPSS 13.0 (SPSS Inc.) was used for all data analyses. We used Student's t-test to determine the difference between patient and control samples. Pearson's correlation analysis was performed for the association study. Each experiment was repeated at least twice or was performed in triplicate. A p value of >0.05 was adopted for statistical significance.
Results
Demographics of Study Population
All 51 patients underwent surgical treatment with total resection or partial resection for recurrent tumors. As shown in Table I, a variety of clinical evaluations were recorded by the certified health provider to assess patients’ clinical features.
MiR-146a is Frequently Downregulated in Glioma Tissues and GBM Cell Lines
To explore the effect that miR-146a has throughout the course of glioma, we used qRT-PCR to analyze the expression of miR-146a in tumor tissues and across glioma cancer cells. With a total of 51 paired glioma and adjacent nontumor glioma tissue samples, we found that on average the presence of miR-146a was significantly downregulated in tumor tissue (Fig. 1A, p<0.05). In vitro, miR-146a was universally decreased in all tested GBM cell lines (0.77 for U87, 0.57 for A172, 0.65 for U251) compared to normal control (SVG p12). Cell behavior may be controlled by miRNA expression derived from clinical samples and cell lines, which suggests that miR-146a may act as an onco-miR by regulating oncogenic signals to control cell behavior. Overexpression experiments were performed to test this hypothesis and to determine whether exogenous miR-146a could further enhance proliferation and tumorigenesis of GBM. Since A172 cells showed the highest reduction of miR-146a expression among all tested cell lines, they were chosen for the next overexpression experiment (Fig. 1B, p<0.05).
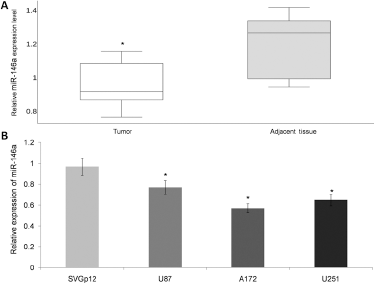
Expression of miR-146a in clinical tissue samples and GBM cell lines. (
Overexpression of miR-146a Increased the Rate of Apoptosis and Decreased the Rate of Cell Proliferation
We studied the impact of miR-146a on tumor cell proliferation. We transfected A172 cells with either miR-146a mimics, inhibitors, or control samples and waited 48 hours. The expression of miR-146a was validated by qRT-PCR. It was found that the level of miR-146a had increased significantly among cells treated with the mimic (Fig. 2A). The MTT assay was used to evaluate the impact of miR-146a on cell proliferation. A statistically significant decrease in proliferation was observed after transfection with the microRNA (Fig. 2B). The data in these figures suggests that miR-146a has the potential to inhibit cell proliferation in vitro. We then investigated the effect of miR-146a on the apoptosis of glioma cells after cisplatin treatment. Cisplatin is widely used to treat several types of tumors including newly diagnosed GBM by promoting tumor cell apoptosis. In the present study, we attempted to prove the promotion effect of miR-146a on cisplatin-induced apoptosis. (Fig. 2C). The results showed that miR-146a clearly increased apoptosis compared with control, while inhibition of miR-146a lowered the apoptosis effect (Fig. 2D).
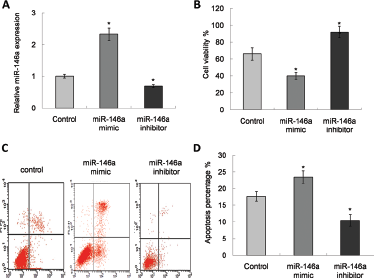
Regulation of miRNA in cell proliferation and apoptosis. (
Notch1 Reacted as a Direct Target of miR-146a
This study investigated whether the Notch1 gene was a potential target for miR-146a in glioma. A luciferase reporter vector containing Notch1 3’ UTR with miR-146a binding sites (Fig. 3A) was generated to address the question of whether or not miR-146a binds to the 3’ UTR of the target miRNA. A mutant reporter vector was also generated which contained Notch1 3’ UTR with a mutation at the putative binding site for miR-146a. As shown in Figure 3B, luciferase activity in miR-146a-mimic-transfected cells significantly decreased compared with control cells (p<0.05). Simultaneously, no luciferase alteration was observed in cells that had been transfected with the mutant 3’ UTR constructs. In order to further assess the correlation between miR-146a and Notch1 expression in glioma tissue, we looked at miR-146a and Notch1 mRNA in all glioma specimens and their matched normal adjacent tissue. Pearson's correlation analysis showed a significant negative correlation between the presence of Notch1 mRNA and miR-146a (Fig. 3C, r = - 0.826, p<0.05).
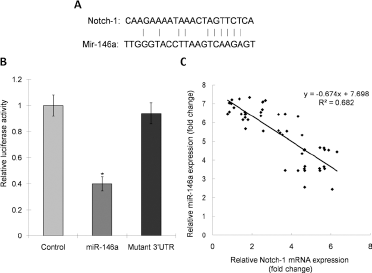
Identification of Notch1 as miR-146a target. (
Discussion
An increasing body of evidence has shown that the dysregulation of miR-146a is related to the development of certain cancers. The function of this miRNA has been controversial due to its contradictory effects across different kinds of cancers. For example, elevated levels of miR-146a are found in cervical cancer (15), whereas downregulation is observed in prostate (16) and papillary thyroid carcinomas (17, 18). Additionally, there is some evidence that miR-146a functions as a tumor suppressor gene in prostate and breast cancers by inhibiting cellular invasion and metastasis (19, 20). Considering the different characteristics of various types of cancers, miR-146a may have multiple regulation roles. Also, genetic polymorphisms that occur in miRNAs may contribute to the previous controversial results. Thus far, we know little about the diverse roles that miR-146a plays in different types of cancers. The current study has looked at the expression of miR-146a in 51 pairs of GBM tumor and adjacent nontumor tissues. The study has also addressed the expression of miR-146a in glioma cell lines. The results show clear downregulation of miR-146a expression in glioma cancer tissues compared with the control group. Li et al (21) obtained results consistent with these, which show that miR-146a inhibits the migration and invasion of glioma cells.
Recent research has discovered several new targets for miR-146a including the interleukin 1 receptor associated kinase 1 (IRAK1), the TNF receptor-associated factor 6 (TRAF6) (22), toll-like receptor 4 (TLR4) (23), signal transducer and activator transcription 1 (Stat 1) (24), the Tata binding protein (25), and the chemokine CCL8/MCP2 (26). For this study, we felt that Notch1 was a good target gene that might be associated with cancer development. Firstly, this was due to the fact that miR-146a has been shown to significantly reduce the activity of a luciferase reporter gene which contains the 3’ UTR sequence of Notch1. Secondly, this was because miR-146a expression is inversely correlated with Notch1 expression in glioma tumors. Stem cell maintenance, cell survival as well as determining whether a cell becomes a neural cell versus a glial cell in the developing nervous system are critically dependent on Notch signaling. Therefore, this Notch signaling pathway, unsurprisingly, plays a key role in brain tumors including GBM (2). Previous research has found that the activation of Notch1 contributes to glioma cell growth, survival and ras-induced transformation (27). Notch1 has been shown to promote radioresistance in glioma stem cells (28). Furthermore, the epidermal growth factor receptor gene (EGFR) has been reported to be overexpressed in GBM through the Notch1-regulated TP53 signaling pathway (29), and knockdown of Notch1 induces apoptosis and inhibits proliferation (30). Many Notch1 signaling pathway component gene expressions increased accordingly alongside EGFR amplification in GBM (31, 32). Taken together, this evidence points to the idea that Notch1 signaling represents an activator in the major GBM development pathway (33, 34). The Notch signaling pathways in the development of brain tumors are regulated by specific miRNAs (35, 36). The pathogenesis and progression of GBM is associated with miRNA deregulation and as a result targeting these specific miRNAs shows great potential for therapeutic intervention.
Our research indicates that miR-146a may negatively regulate the expression of the Notch1 gene. Thus, dysregulation of miR-146a may be another mechanism that leads to downregulation or loss of Notch1 in glioma.
In conclusion, we have shown that miR-146a is significantly downregulated in glioma tumor tissue. We have also shown that the overexpression of miR-146a can effectively inhibit cell proliferation and enhance the apoptosis of glioma cells in vitro. MiR-146a can negatively regulate the expression of the Notch1 gene. These results point to the potential role that miR-146a has for novel cancer therapies.
Footnotes
Acknowledgment
We want to acknowledge the evaluators, research assistants and patients in this study.
Financial support: None.
Conflict of interest: All authors declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
