Abstract
Background
Many studies have investigated the prognostic value of metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) in human cancers. However, these studies were often limited by small sample sizes. Therefore, we performed this updated meta-analysis to summarize the potential value of MALAT1 as a biomarker for early treatment and to predict survival in various human malignant neoplasms, through the inclusion of the latest literature and improved methodology.
Methods
Twelve eligible articles were systematically obtained from PubMed, Medline, Embase, Web of Science, China National Knowledge Infrastructure and the Cochrane Library, from inception up to June 30, 2015. Survival was assessed using pooled hazard ratios (HRs) and 95% confidence intervals (95% CIs).
Results
By combining the results of 12 studies, we found elevated MALAT1 expression was associated with poor survival in most cancers, with a pooled HR of 1.90 (95% CI, 1.56-2.30) for overall survival (OS) and 3.06 (95% CI, 2.06-4.56) for recurrence-free survival/disease-free survival. Subgroup analyses according to ethnicity, tumor type, assay method, sample size, HR-calculation method and analysis type did not affect the predictive role of MALAT1 for OS in various cancer types. Further, by combining results from studies that used multivariate analyses, we found elevated MALAT1 was an independent prognostic factor for OS (HR = 1.98; 95% CI, 1.58-2.48).
Conclusions
MALAT1 could serve as a potential prognostic biomarker in various cancers and may be a potential therapeutic target for the treatment and early detection of recurrence.
Keywords
Introduction
Malignant tumors, one of the most common public health issues worldwide, have become the primary cause of mortality from clinical diseases (1). Despite therapeutic advances, the 5-year survival of patients with malignant tumors remains unsatisfactory (2). Many genes are known to function as regulatory molecules and to play critical roles in predicting treatment outcome in human cancers (3); nevertheless, there is a dearth of effective molecular prognostic biomarkers. Hence, newer, universally accepted cancer biomarkers are crucial for the detection, treatment and prevention of this fatal disease (4).
Noncoding RNAs (ncRNAs), including microRNAs, PIWI-interacting RNAs and long noncoding RNAs (lncRNAs) (5), have shown great clinical value in cancer prognosis (6). lncRNAs are a type of ncRNA with over 200 nucleotides (7), and many of them play regulatory roles in various biological processes. Their potential use as novel biomarkers of tumor progression, occurrence and prognosis (8-9-10-11) has created new awareness regarding cancer therapeutic strategies (12, 13).
Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1), which has been mapped to human chromosome band 11q13, is a highly abundant nucleus-restricted RNA that localizes to nuclear speckles (14). MALAT1 is abnormally expressed in several types of human cancers, such as liver cancer (15), non-small cell lung cancer (NSCLC) (16), prostate cancer (17) and glioblastoma (18). It promotes tumor growth and metastasis via mechanisms such as regulating cellular levels of phosphorylated forms of SR proteins (14), binding to the active regions of chromosomes (15) and regulating alternative splicing of oncogenic mRNAs.
Several studies have revealed that MALAT1 plays an essential role in many cancers (19-20-21) and interacts with many genes, resulting in cellular migration and tumor development. Hirata et al reported that MALAT1-bound Ezh2 contributed to oncogenesis, while the knock down of MALAT1 inhibited oncogenic function, resulting in the inhibition of epithelial-mesenchymal transition via E-cadherin recovery and beta-catenin down-regulation (21). Vassallo et al showed that MALAT1 transcription was regulated by Wnt signaling, and a novel Wnt/beta-catenin-MALAT1 regulatory axis contributed to endometrial carcinoma development (18). Experimental research has proven that MALAT1 overexpression could accelerate cell proliferation, invasion and migration and could enhance survival in tumor cells in vitro (22-23-24). Conversely, inhibition of MALAT1 expression reduced cellular proliferation and invasion while increasing cancer cell apoptosis (7, 25). Taken together, all of these findings indicate that MALAT1 is substantially involved in cancer pathogenesis.
In addition to the role of MALAT1 in malignant cells, Ma et al recently reported that MALAT1 plays a role in maintaining the proliferation potential of hematopoietic cells (26). Furthermore, MALAT1 was down-regulated by the tumor suppressor gene p53 that binds to the MALAT1 core promoter region, thus suppressing proliferation. This latest experimental result provides additional evidence of a close connection between MALAT1 expression and both malignant and normal cell growth. Nevertheless, the precise mechanism through which MALAT1 regulates cell proliferation and promotes tumor development and progression is still poorly understood, and the prognostic value of MALAT1 has been found to be nonsignificant or even inconsistent in current clinical research (20, 27-28-29-30). Therefore, we performed a meta-analysis to reveal the true value of MALAT1 as a prognostic biomarker in various human malignant neoplasms, by developing a more comprehensive retrieval strategy, following a more scientific data-extraction process to ensure all data were accurate and valid, and performing stratified analyses to obtain a stronger, more valid conclusion.
Methods
This study adhered to the guidelines for meta-analyses of tumor biomarkers prognostic studies (31, 32) and the Preferred Reporting Items for Systematic Reviews and Meta-analysis criteria (Supplementary Tab. I, available online at www.biological-markers.com) by Moher et al (33).
We performed an online search using PubMed, Medline, Embase, Web of Science, China National Knowledge Infrastructure and the Cochrane Library to identify studies on the association between MALAT1 and survival of cancer patients (last update: June 30, 2015). The following search words were used: “MALAT1,” “metastasis-associated lung adenocarcinoma transcript 1,” “long noncoding RNA,” “lncRNA,” “cancer,” “neoplasm,” “carcinoma,” “prognostic,” “prognosis,” “mortality,” “survival,” “outcome” and “recurrence.” The reference lists of the included studies were also manually searched to identify related articles. The comprehensive electronic database search was conducted independently by 2 authors (Y. L. and Z. Y.). Disagreements were resolved by consensus.
Inclusion and Exclusion Criteria for Articles
The inclusion criteria were as follows: the study (a) included patients with any type of cancer; (b) investigated the 2 levels of MALAT1 expression: high or low; (c) identified the relationship between MALAT1 expression and patient outcomes; and (d) included a sufficient number of cases to determine effect sizes and 95% confidence intervals (95% CIs). Studies that met the above-mentioned eligibility criteria were further excluded if they (a) were case reports, letters, reviews or animal trails; (b) lacked complete effect quantities; (c) lacked insufficient survival data for meta-analysis; or (d) were non-English or non-Chinese studies. Disagreements on eligibility were resolved through a discussion with the fourth author (J. Z.).
Data Extraction and Qualitative Assessment
The data were independently extracted by 3 investigators (Y. L., Z. Y. and X. W.); these data included author name; publication date; origin of population; ethnicity; age; tumor type; case number; assay method; cutoff values; statistical methods; and hazard ratios (HRs) of MALAT1 expression for overall survival (OS), disease-free survival (DFS) and recurrence-free survival (RFS), as well as 95% CIs and p values. If a study reported the results obtained using both univariate and multivariate analyses, we chose the HRs adjusted using a Cox regression model since it considers confounding factors and is therefore more precise, unless only an unadjusted value was available.
Using the Newcastle-Ottawa scale (NOS) for quality assessment (34), we assessed the quality of eligible studies carefully and objectively. For the evaluation results, we used scores of 0 to 9 to represent the quality of the research, and studies were considered to be of high quality if the score was 6 or more.
Statistical Analysis
HRs and 95% CIs were extracted using the method described by Parmar et al (35). The effect sizes were directly obtained from the literature. The second method used to obtain the HRs involved analyzing the sample size and p values. If the above methods failed, we extracted data from the survival curves by using Engauge Digitizer version 4.1 (36, 37). Where necessary, we approached the authors for original data. The chi-square test and I2 statistic were performed to quantify heterogeneity among articles (38). A chi-square test with p<0.10 and I2 statistic with values >50% indicated strong heterogeneity across studies. We used a fixed-effect model for estimating HRs and 95% CIs; if between-study heterogeneity was significant, then the random-effects model was used. Subgroup analyses and meta-regression were further conducted by ethnicity, tumor type, assay method, sample size, HR-calculation method and analysis type to explore between-group differences.
The results for OS and RFS/DFS were summarized in forest plots, and stability was confirmed using a sensitivity analysis for OS. Potential publication bias was assessed using Begg's funnel plots and Egger's tests. Our meta-analysis was conducted using Stata/SE 12.0 (Stata Corp., College Station, TX, USA) for Windows. A p value <0.05 was considered statistically significant.
Results
Included Studies and Characteristics
On the basis of the selection criteria, 326 studies were initially selected (Fig. 1). After reading the title and abstract, we excluded 230 irrelevant or duplicate articles. After reading the full texts of the remaining 96 studies, 81 were eliminated because they were review articles, laboratory studies or animal studies. Fifteen potential candidate studies were further evaluated, and 3 of these (16, 21, 39) were excluded from further analysis because of insufficient survival data (HRs and survival curves). Finally, we included 12 eligible articles that evaluated the potential relationship between MALAT1 expression and patient survival. While we also searched for studies published in Chinese, none of these met the inclusion criteria.
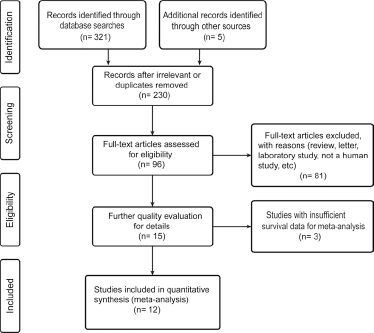
Flow chart for the selection of eligible studies included in this meta-analysis.
The major characteristics and main results of the 12 articles are shown in Table I. In total, 1,042 patients were included in the analysis. While 2 studies investigated whites, 10 studies investigated Asians. The malignant neoplasms assessed in these studies included NSCLC (19, 27), gastric cancer (20), clear cell renal cell carcinoma (40), hepatocellular carcinoma (41), multiple myeloma (29), glioma (42), colorectal carcinoma (43), pancreatic ductal adenocarcinoma (44, 45) and bladder cancer (30). Nine studies used quantitative real-time polymerase chain reaction (qRT-PCR) to assess MALAT1 expression, whereas 2 used in situ hybridization (ISH). All studies provided a cutoff value for the analysis, such as the mean value of MALAT1 expression, receiver operating characteristic (ROC) curve analysis and intensity of ISH hybridization signal. Eleven studies mainly focused on OS, 1 focused on RFS and 2 focused on OS and DFS. HRs and 95% CIs were extracted from each study. Eight studies directly reported HRs and 95% CIs, but for the remaining 4, survival data had to be extracted from the text.
Main characteristics and results of the 12 included studies
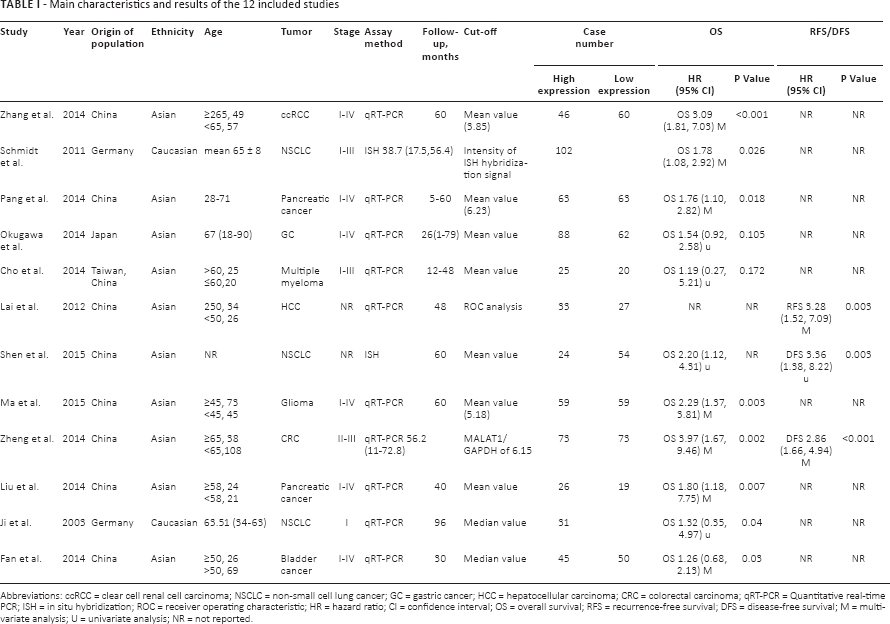
Abbreviations: ccRCC = clear cell renal cell carcinoma; NSCLC = non-small cell lung cancer; GC = gastric cancer; HCC = hepatocellular carcinoma; CRC = colorectal carcinoma; qRT-PCR = Quantitative real-time PCR; ISH = in situ hybridization; ROC = receiver operating characteristic; HR = hazard ratio; CI = confidence interval; OS = overall survival; RFS = recurrence-free survival; DFS = disease-free survival; M = multivariate analysis; U = univariate analysis; NR = not reported.
Qualitative Assessment
The quality of the 12 eligible studies was evaluated according to the NOS. The quality of all qualified articles varied from 6 to 9 (Supplementary Tab. II, available online at www.biological-markers.com). A higher score indicated a better research methodology. Thus, all articles could be analyzed further.
Meta-Analysis Results of the Prognostic value of MALAT1
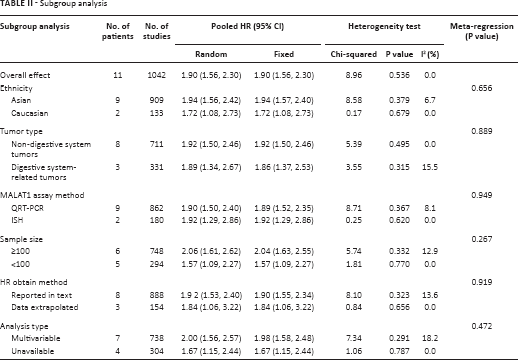
For studies evaluating OS, no heterogeneity was noted in the 11 studies used for OS analysis (χ2 = 8.96, p = 0.536, I2 = 0.0%). Therefore, the fixed-effects model was used for data analysis. We found that patients with elevated MALAT1 expression had significantly unfavorable OS (HR = 1.90; 95% CI, 1.56-2.30; Tab. II) in various tumors. Moreover, subgroup analysis was performed based on the ethnicity, tumor type, assay method, sample size, HR-calculation method and analysis type despite the absence of heterogeneity across articles (Tab. II). There was no evidence of important heterogeneity across studies in the above subgroup analyses. Furthermore, we performed meta-regression analysis by including covariates such as ethnicity, tumor type, assay method, sample size, HR-calculation method and analysis type. Similar to the subgroup analysis, those factors did not result in interstudy heterogeneity. The leave-1-out sensitivity analysis was performed to assess the robustness of the results by removing each study and re-performing the analysis. This analysis showed that the research results were not significantly altered (Fig. 2A), thereby validating the reliability and stability of our data. Ultimately, in the meta-analysis for OS, there was no obvious asymmetry according to the Begg's funnel plot (Fig. 2B). Similarly, Egger's test showed a p value of 0.820, indicating no publication bias in our study.
Subgroup analysis
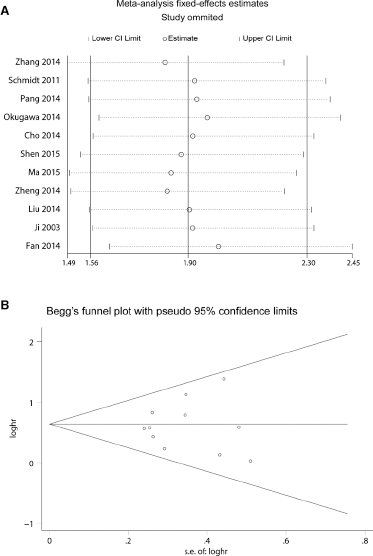
Forest plots showing overall survival after leave-1-out sensitivity analysis (
Meanwhile, there was no significant heterogeneity in RFS/DFS among the 3 studies (χ2 = 0.13, p = 0.936, I2 = 0.0%); therefore, fixed effects were applied to calculate the pooled HR. Similarly, increased MALAT1 expression was significantly correlated with RFS/DFS (HR = 3.06; 95% CI, 2.06-4.56; figure not shown).
Independent Prognostic value of MALAT1
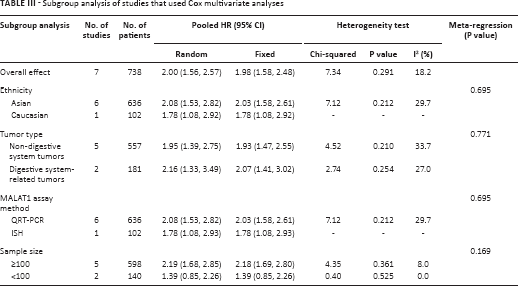
To illustrate whether MALAT1 was an independent prognostic factor in cancer patients, 7 studies that used Cox multivariate analyses were included in further research. By combining HRs, the analysis results indicated that elevated MALAT1 was an independent contributor to the OS of cancer patients (HR = 1.98; 95% CI, 1.58-2.48, Tab. III). Stratified analysis suggested ethnicity, tumor type, assay method and sample size did not change the outcome (Fig. 3). Additionally, slight heterogeneity in the subgroup analysis of studies is acceptable. In accordance with the outcomes of subset analysis, all of the investigated factors did not account for the between-study heterogeneity based on meta-regression analysis (Tab. III). No obvious publication bias was found by assessment with both Begg's test (figure not shown) and Egger's test (p = 0.296).
Subgroup analysis of studies that used Cox multivariate analyses
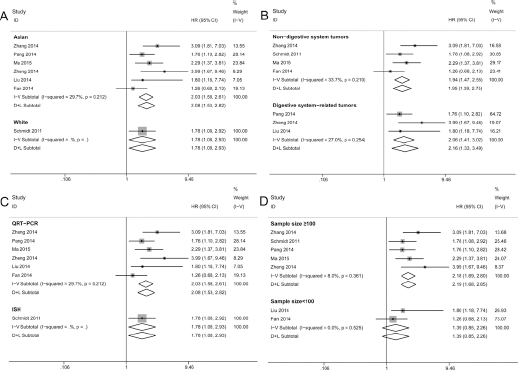
Forest plots showing subgroup analysis according to ethnicity (
Discussion
MALAT1 has been widely studied and is aberrantly expressed in most human cancers. It was the first lncRNA whose role was studied in tumorigenesis and was originally identified as a prognostic factor for metastasis and survival in patients with NSCLC (27). Schwarzenbach et al (6) systematically analyzed the potential clinical use of noncoding RNAs as diagnostic and prognostic tumor markers. Previous meta-analyses have shown that elevated MALAT1 might be an unfavorable prognostic predictor in cancer patients. Zhang et al (46) and Xue et al (47) both found that overexpression of MALAT1 increased the risk to the patient's OS, and there was a significant relationship between MALAT1 and OS in both NSCLC and pancreatic cancer. By subgroup analysis, Wu et al (48) and Wei et al (49) further detected a significant association between MALAT1 and the OS of cancer patients according to the ethnicity, tumor type and assay method. In particular, Zhang et al performed a secondary analysis of Kaplan-Meier survival curves to show the level of MALAT1 expression correlated with OS and DFS. Zhu et al (50) analyzed the relationship between MALAT1 and lymph node metastasis and distant metastasis, confirming the role of MALAT1 in tumor metastasis.
However, through careful analysis, we found that those systematic reviews still had some disadvantages. The sample size of the above studies was relatively small, and inconsistent results existed in the cancer type subgroups (46, 48). In addition, the effect sizes extracted from the included studies (27, 29, 30) have obvious differences among the previous systematic reviews, and some original research was not included; therefore, the final pooled results were unreliable and required further validation.
By combining the results of 7 articles that used multivariate analyses, we found MALAT1 to be a significant predictor of poor survival of patients (HR = 1.98; 95% CI, 1.58-2.48). Subgroup analyses by ethnicity showed that pooled HRs for Asians were greater than those for whites. Notably, when the multivariate covariates were considered in the subgroup analyses based on tumor type, our study revealed that MALAT1 expression might be more relevant as a marker for patients with digestive system carcinomas than for patients with carcinomas not related to the digestive system for predicting OS; this result differs from the preliminary subgroup analysis and a previous systematic review (49). In addition, subgroup analysis showed that the MALAT1 assay method did not alter the pooled effect significantly. Because the results may be affected by the number of cases in the included studies, we conducted a stratified analysis by sample size and found that high expression of MALAT1 was a significant predictor of poor OS in the studies with more than 100 cases, while it showed no statistical significance in those with less than 100 cases. This conclusion indicated that studies with relatively small sample sizes may fail to detect the prognostic role of MALAT1 in various human cancers, which is also different from previous research (49). Furthermore, through sensitivity analysis and meta-regression, the stability of pooled results was enhanced.
Similarly, increased MALAT1 expression was significantly related to RFS/DFS (HR = 3.06; 95% CI, 2.06-4.56), indicating that MALAT1 expression level was linked closely to the progression of patients with various human cancers. However, the results should be interpreted with caution because only 3 studies were included in the meta-analysis; thus, larger sample size studies are needed in the future to verify our results. Further, considering the limited number of studies, sensitivity analysis and publication bias were not assessed for RFS/DFS to avoid reducing statistical power, which may also affect the stability of the results.
This meta-analysis suggested that elevated MALAT1 expression was predictive of poor survival in patients with various cancer types. However, the association between MALAT1 and human cancers should be considered carefully for several reasons. First, the sample size of the study was still relatively small; thus, we could not evaluate subgroups for a particular type of cancer. Second, the main ethnicity of the patients in the current analyses was Asian. Thus, our results might not be applicable to other populations. Third, the cutoff value of MALAT1 expression was inconsistent in the included articles. Because the amount of MALAT1 expression data was insufficient, it was difficult to set a standard cutoff defining high and low MALAT1 expressions. Therefore, a different, accurate value could affect the effectiveness of MALAT1 as a prognostic marker in human cancers. Fourth, although there was no significant evidence of publication bias in the OS analysis, some unpublished articles or articles published in other languages were not included in this study. Thus, our results might be overestimated.
This meta-analysis indicated that increased MALAT1 expression is significantly associated with poor survival in patients with various human cancers. The development of applicable biomarkers is pivotal for predicting patients’ survival. Despite the limitations described above, our results suggest that MALAT1 could serve as a potential prognostic biomarker in various cancers and may represent a potential therapeutic target for the treatment and early detection of recurrence. Future studies with better design may help validate our results and lead to the wider clinical application of MALAT1.
Footnotes
Financial support: This work was funded by the Natural Science Foundation of China (NSFC; Grant No. 81360351) and the Department of Science and Technology of Guizhou Province (Grant No. Qian Ke He J Zi [2014] 2186).
Conflict of interest: The authors declare that they have no conflict of interests regarding the publication of this article.
