Abstract
Purpose
Colorectal cancer (CRC) is the third most common cancer and fourth leading cause of cancer mortality, and twin studies have shown that approximately 35% of the variation in susceptibility to CRC involves inherited genetic differences. We sought to investigate potential genetic associations between some single nucleotide polymorphisms (SNPs) and the risk of CRC in the Chinese Han population.
Methods
We conducted a case-control study including 269 cases and 309 controls. Sixteen SNPs associated with CRC risk were selected from previous genome-wide association studies and genotyped using Sequenom MassARRAY technology. Odds ratios and 95% confidence intervals (CIs) were calculated by unconditional logistic regression adjusting for age and gender.
Results
Using the chi-squared test we found that rs9365723 was associated with CRC risk (p = 0.012). With genetic model analysis, the genotype A/G-G/G (OR = 1.50; 95% CI 1.02-2.21; p = 0.038) of rs9365723 showed an increased risk of CRC in the dominant model. Furthermore, we found that rs9365723 was associated with an increased risk only for colon cancer but not rectal cancer (p = 0.009 and p = 0.414, respectively).
Conclusions
Our results, combined with previous studies, suggest that rs9365723, located on SYNJ2, is associated with the risk of CRC in a Chinese population. Thus, SYNJ2 may play a role in the development of CRC, especially colon cancer.
Introduction
Colorectal cancer (CRC) is the third most common cancer and fourth leading cause of cancer mortality (1). It affects over one million people each year worldwide (2). During the past few decades, the incidence and mortality of CRC have been increasing rapidly in China: according to a 2007 investigation from 38 registered areas of China, CRC accounted for approximately 391,000 new cases and 186,800 deaths.
Although the recent development of new drugs and more advanced therapeutics have remarkably improved overall survival, and even if the 5-year survival rate is around 90% for patients who are diagnosed at stage I (3), the prognosis of patients with CRC at an advanced stage is still disappointing. CRC is a very complex disease in which environmental and genetic risk factors play a role. Unhealthy dietary habits and lifestyle, including high-fat and low-fiber diets, may contribute to the development of CRC. Genetic factors are also important: twin studies have shown that approximately 35% of the variation in susceptibility to CRC involves inherited genetic differences (4, 5).
Genome-wide association studies using large sets of cases and controls have proven to be an effective strategy to identify common single nucleotide polymorphisms (SNPs) associated with cancer risk (6). For the present study, we selected 16 SNPs from 15 chromosomal regions which were either associated with CRC in previous studies or were located in close proximity to previously reported risk SNPs: rs1912453 (1q23.3), rs11903757 (2q32.3), rs4591517 (3p24.3), rs367615 (5q21.3), rs647161 (5q31.1), rs2057314 (6q22.1), rs9365723 (6q25.3), rs39453 (7p15.3), rs12548021 (8p12), rs10114408 (9q22.32), rs1665650 (10q25.3), rs10774214 (12p13.32), rs3217901 (12p13.32), rs59336 (12q24.21), rs4939827 (18q21.1), and rs961253 (20p12.3) (7-8-9-10-11). Among these variants, rs4939827 and rs961253 have been researched in Chinese (12-13-14), but the results were inconsistent across studies. Moreover, few studies have reported the association between other SNPs and CRC susceptibility in Chinese. This case-control study focused on the risk of 16 SNPs in a Chinese population and the differences between our results and previous association studies in other populations.
Materials and methods
Study Subjects
The present study included 269 CRC cases and 309 controls. All patients were consecutively recruited from the First Affiliated Hospital of Xi'an Jiaotong University, China. The diagnosis of CRC was confirmed by histopathological examination. The control subjects were without a positive personal history of malignancy or chronic disease. Both patients and controls were Han Chinese. We excluded patients who underwent radiotherapy or chemotherapy as well as controls with chronic disease. All participants in the current study were not blood related.
Clinical Data and Demographic Information
We used a standard epidemiological questionnaire to collect personal data through in-person interviews including gender, age, occupation history, tobacco history and other lifestyle factors, as well as clinical and pathological investigations in a custom-designed questionnaire. After the study participants had signed informed consent forms, 5 mL of peripheral venous blood was collected in EDTA-coated tubes and stored at −80°C. The study was approved by the ethics committee of the First Affiliated Hospital of Xi'an Jiaotong University.
SNP Selection and Genotyping
All of the 16 SNPs we selected were previously reported to be most associated with CRC, with minor allele frequencies >5% in the HapMap Chinese Han Beijing population. DNA was extracted from whole-blood samples by means of a GoldMag-Mini whole blood genomic DNA purification kit (GoldMag Co. Ltd.). The extracted DNA was quantified using a NanoDrop 2000 spectrophotometer (Thermo Scientific). The Sequenom MassARRAY Assay Design 3.0 software was used to design primers for amplification and extension reactions (15, 16). Genotyping was performed by Sequenom MassARRAY RS1000 according to the manufacturers’ instructions.
Statistical Analysis
Statistical analyses were conducted with the SPSS 17.0 statistical package. All p values presented in this study are 2-sided, and a p value <0.05 was considered statistically significant. The differences in the characteristics of the case and control groups were compared using the chi-squared test (for categorical variables) and Student's t-test (for continuous variables). The Hardy-Weinberg equilibrium of all selected SNPs in controls was tested using the chi-squared test (17). The chi-squared test was also used to detect statistical differences in the distributions of genotypes between cases and controls. In addition, we compared allele frequencies in different tumor locations of CRC. The effects of the polymorphisms on the risk of CRC were expressed as odds ratios (ORs) with 95% confidence intervals (95% CIs), computed using unconditional logistic regression analysis with adjustments for age and sex (18).
SNP stats, a website software tool (http://bioinfo.iconcologia.net/SNPstats_web), was used to test the associations between all SNPs and the risk of CRC under different genetic models (codominant, dominant, recessive, overdominant and additive) (19). The most common control homozygote was used as a reference.
Results
This study included 268 CRC cases (101 women, 167 men; mean age 58.4 years) and 309 controls (113 women, 196 men; mean age 49.1 years). The clinical characteristics of cases and controls are summarized in Table I.
General characteristics of the study population
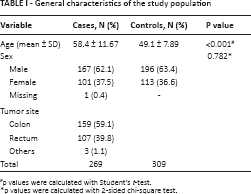
p values were calculated with Student's t-test.
p values were calculated with 2-sided chi-square test.
No significant difference was found between patients and controls in the distribution of sex (p = 0.782). The incidence of CRC was higher in men than women (62.1% and 37.5%, respectively; p<0.01). The case group included a higher proportion of colon cancer than rectum cancer (59.1% and 39.8%, respectively).
Table II shows the basic characteristics of all SNPs in the study population. The minor allele frequency of SNPs among cases and controls was consistent with the genotyping data for the Chinese Han Beijing population in the HapMap database. The SNPs were in Hardy-Weinberg equilibrium in controls (p>0.05), except for rs1077421, which was excluded from subsequent analyses. Our study established that rs9365723 was significantly associated with increased risk of CRC, with an OR of 1.34 (95% CI 1.06-1.70; p = 0.012). No significant associations were detected between CRC risk and the other SNPs.
Basic information of candidate SNPs and associations with colorectal cancer risk
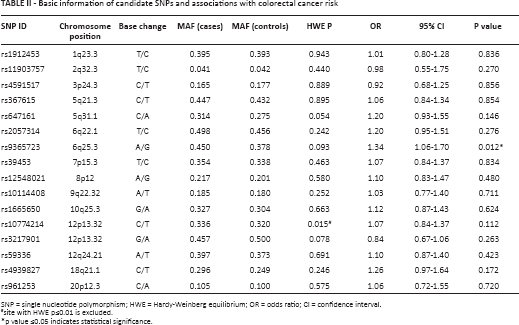
SNP = single nucleotide polymorphism; HWE = Hardy-Weinberg equilibrium; OR = odds ratio; CI = confidence interval.
site with HWE p≤0.01 is excluded.
p value ≤0.05 indicates statistical significance.
We further assessed the association between SNPs and CRC risk using genetic models by unconditional logistic regression analysis with adjustments for age and gender. The results showed that the G allele of rs9365723 increased the CRC risk (Tab. III). In the dominant model, the GG and AG genotypes increased the CRC risk more than 1.5-fold (OR = 1.50; 95% CI 1.02-2.21; p = 0.0038) after adjusting for age and sex.
Genotypic model analysis of relationship between rs9365723 and colorectal cancer risk
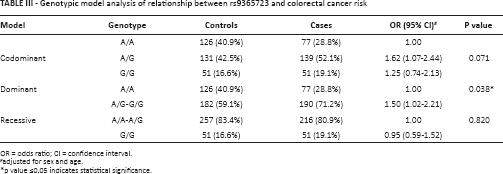
OR = odds ratio; CI = confidence interval.
adjusted for sex and age.
p value ≤0.05 indicates statistical significance.
Given the biological differences between the histological forms of CRC, we stratified by subtype. In this restricted analysis, the statistically significant association between rs9365723 and CRC was stronger among patients with colon cancer (OR = 1.43; 95% CI 1.09-1.88; p = 0.009) than among all patients with CRC (Tab. IV). The results of the analysis by genetic models are listed in Table V. In the dominant model, the GT and GG genotypes of rs9365723 were associated with a 1.70-fold increased colon cancer risk (OR = 1.70; 95% CI 1.06-2.73; p = 0.027).
Basic information of candidate SNPs and associations with rectum and colon cancer
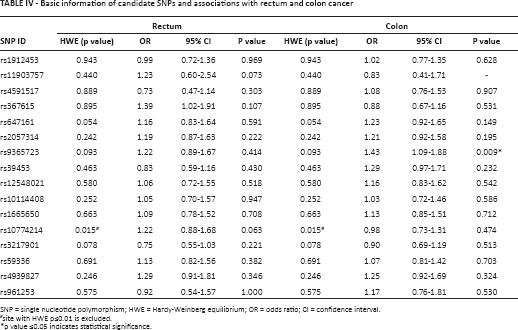
SNP = single nucleotide polymorphism; HWE = Hardy-Weinberg equilibrium; OR = odds ratio; CI = confidence interval.
site with HWE p≤0.01 is excluded.
p value ≤0.05 indicates statistical significance.
Genotypic model analysis of relationship between rs9365723 and colon cancer risk
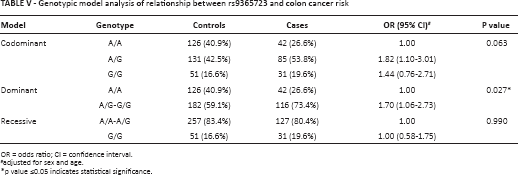
OR = odds ratio; CI = confidence interval.
adjusted for sex and age.
p value ≤0.05 indicates statistical significance.
Discussion
In this study, we investigated the association between 16 SNPs and the risk of CRC in a Chinese population. We found for the first time that rs9365723 was associated with a significantly increased risk of CRC. Stratified analyses indicated that rs9365723 was more likely to increase the risk of colon cancer.
The rs9365723 SNP is located in synaptojanin 2 (SYNJ2), an important member of the inositol polyphosphate phosphatase family (20). SYNJ2 removes the 5-position phosphate from phosphoinositides, thought to be involved in membrane trafficking and signal transduction pathways. Using a causative mutation in the SYNJ2 gene, Manji et al. identified SYNJ2 as a critical regulator of hair cell survival that is essential for maintaining normal hearing in mice (21). Recent studies demonstrated that SYNJ2 SNPs were associated with agreeableness and symptoms of depression in the elderly (22) and with cognitive abilities (23). Although the functions of the SYNJ2 gene were poorly understood, we discovered that SYNJ2 may provide a potential mechanism of colorectal tumorigenesis. Additionally, we observed that rs9365723 was associated with an increased risk of colon cancer but not rectal cancer, and it could therefore be hypothesized that SYNJ2 plays a more important role in the development of colon cancer than rectal cancer. Therefore, the biological functions of SYNJ2 are of great interest and need further investigation.
Our results indicated that rs4939827, from chromosome 18q21.1, did not increase the risk of CRC in Chinese. Previous evidence relates rs4939827 to the risk of CRC. For example, several genome-wide association studies showed that rs4939827 conferred an increased risk of CRC (24, 25). Several studies showed similar results in the Chinese population (12-13-14). However, rs4939827 was associated with a significantly decreased risk of CRC in Korean (26) and Croatian (27) populations. These conflicting results might be attributed to ethnic differences among current studies. Therefore, these findings need to be validated in further large and more ethnically diverse population-based studies.
We also found no correlation between rs961253 (located in proximity to bone morphogenetic protein [BMP] genes) and CRC risk. Associations between rs961253 and CRC risk have also been inconsistent across studies (13, 28-29-30). For example, several studies identified this association in Chinese (13, 28), southern Europeans (29) and African Americans (30), but Thean et al. failed to demonstrate the association in Singapore Chinese (31). It is currently believed that BMP signaling is related to CRC development through an increase in stem cell numbers near colorectal crypt bases, thus elevating the number of cells susceptible to tumor-causing mutations (32).
These conflicting results might be attributable to ethnic differences among the current studies and differences in the number of specimens. Moreover, certain dietary and lifestyle factors are undoubtedly major risk factors for CRC. Therefore, additional studies involving a larger number of cases are needed to assess the predictive value of rs961253 and rs4939827 for CRC. More importantly, functional study of the polymorphism in addition to epidemiological replication is crucial to clarify whether the association is causative.
In conclusion, to our knowledge this was the first attempt to provide powerful new evidence for the relationship between rs9365723 and susceptibility to CRC, especially colon cancer. Our findings suggest that SYNJ2 variants may play an important role in the development of CRC.
Footnotes
Acknowledgment
We are grateful to all patients and individuals in the study who made this work possible. We would also like to thank the clinicians and other hospital staff who contributed to the data collection for this study.
Financial support: This work is supported by the National 863 High-Technology Research and Development Program (No.2012AA02A519).
Conflict of interest: The authors have no conflicts of interest to report.
