Abstract
BACKGROUND:
miRNAs play a crucial role in the genesis of cancer, either as tumor suppressor genes or as oncogenes. Single Nucleotide Polymorphisms (SNPs) in the seed region of microRNAs (miRNAs) can dysregulate their levels in the tissues and thereby affect carcinogenesis. The association of SNP in miR-146a (rs2910164) with the risk of oral squamous cell carcinoma (OSCC) has not been understood.
OBJECTIVE:
In the present study, we have determined the association and functional significance of miR-146a (rs2910164) SNP with susceptibility to OSCC predisposition.
METHODS:
In the present case-control study, we enrolled 430 subjects from central India (215 OSCC cases and 215 healthy controls). We performed genotyping by Kompetitive Allele Specific PCR (KASP), and their correlation with OSCC susceptibility was analyzed. miRNA expression profiling in tumor tissues and adjacent normal tissues from six OSCC patients was done by a NanoString n-Counter-based assay. Subsequently, gene ontology and pathway analysis were performed with FunRich version 3.13.
RESULTS:
The CC genotype of rs2910164 miR-146a was significantly associated with the increased risk for OSCC (CC vs GC, OR
CONCLUSION:
Our finding suggests that SNP rs2910164 of miR-146a may be a genetic risk factor for OSCC susceptibility in the Central India population. However, more extensive multicenter studies are required to validate these findings.
Introduction
Oral Squamous Cell Carcinoma (OSCC) is a locally aggressive tumor that arises in the oral cavity and lips and represents the most common type of Head and Neck Cancer (HNC) [1]. According to the Global Cancer Observatory (GCO) data, in 2020, the global incidence of OSCC was 377,713. Asia contributed the most, with the highest number, 248,360, followed by Europe and North America [2]. In India, the estimated incidence of OSCC in 2020 was around 135,929, which accounted for 10.3% of the total cancer burden of the country (GLOBOCAN 2020) [1]. Major risk factors of OSCC are smokeless tobacco, betel-quid chewing, alcohol, smoking, and unhygienic oral condition. These factors work synergistically towards the initiation, development, and progression of OSCC and collectively increase the risk by 35% [3]. Human Papillomavirus (HPV) and Epstein-Barr Virus (EBV) are also accepted as risk factors and are associated with cancer of the oropharynx and nasopharynx [4]. Other than these risk factors, genetic variations like polymorphisms in DNA repair genes and cellular regulatory networks can modify the susceptibility to OSCC. Single nucleotide polymorphisms (SNPs) in microRNA (miRNA) genes plays an active role in establishing susceptibility towards OSCC through their involvement in various cellular proliferative, developmental, metabolic, and immunological mechanisms [5].
MicroRNAs are small non-coding RNA molecules, approximately 18–24 nucleotides long, bind to 3
SNPs are the most common types of genetic variations that can alter target binding affinity, expression pattern, and abnormal maturation. A strong association exists between SNPs in miRNA gene, promoter, and target binding sites with genetic susceptibility to various cancers [5]. Furthermore, SNPs are specific to ethnicity, and exploration in diverse populations will further help in the development of diagnostic, prognostic as well as therapeutic modalities for cancer [5]. miR-146a is known to have an antagonistic effect on inflammation and dampening of adaptive as well as innate immune responses [8]. It has recently gained attention due to its major role in M2 macrophage polarization and NF-kB regulation [9]. rs2910164 (G
Previous studies have shown that rs2910164 in miR-146a is significantly associated with non-small cell lung cancer (NSCLC), gastric cancer, sporadic breast cancer, colorectal cancer, and digestive system cancer [11, 12, 13, 14]. Although previous studies have explored the role of rs2910164 in miR-146a and its significance with a predisposition to multiple cancers, its role in OSCC is not well studied. Therefore, the present study has analyzed the association of rs2910164 of miR-146a with OSCC susceptibility, miR-146a expression levels in tumor tissue of OSCC patients, and in silico functional characterization.
Methodology
Study design
The present study is a hospital-based case-control study. A total of 430 subjects, patients with OSCC (
SNP genotyping
Blood samples were collected in EDTA vials, and genomic DNA from all cases and controls was isolated by the conventional phenol-chloroform method [15]. Genotyping of rs2910164 in miR-146a was performed using Kompetitive Allele Specific PCR (KASP) genotyping assay, and the post-run data were analyzed using the SNPviewer software, LGC Biosearch Technologies [16]. All assays were performed on 96-well plates. Primers for each tag SNP were designed using Kraken
RNA Extraction and expression profiling
A total of six biopsy-proven newly diagnosed adult patients (cases) of OSCC were included for mRNA expression study, and independent diagnosis, and histological examinations of each patient was made by physicians and pathologists following standard criterion (Table 3). Three patients had tumor in the buccal mucosa and three in the tongue. None of the patients received curative treatment (surgery, chemo- or radiotherapy) before their enrollment. Tumor tissue (approx. 2 mm
Target prediction and functional annotations
Target genes for miR-146a-5p were predicted using the multiMiR package of R [19]. The multiMiR data-base represents data from fourteen external databases consisting of miRecords, TarBase, and miRTarBase (validated miRNA-Target database), DIANA-microT, MicroCosm, miRanda, ElMMo, miRDB, PITA, PicTar, TargetScan (Predicted miRNA-Target databases), miR2Disease, PhenomiR, and Pharmaco-miR (disease-/drug-related miRNA databases. Subsequently, the impact of SNP in miR-146a was predicted for any change in the miR-146a target site, like disruption or creation of a new target site with miRSNPV3, MicroSNiPer, multiMiR (R) [10, 20].
The second level of validation was applied to further select significant miRNA targets at the expression level in oral cancer. A list of downregulated genes in OSCC was downloaded from the TCGA databases (
Pathway analysis
The molecular mechanism and significant pathway associated with miR-146a-5p dysregulation were found using FunRich version 3.1.3 with Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) based on p-value. Further, the protein-protein interaction (PPI network) was made with the STRING database (
Distribution of selected demographic variables of OSCC Patients and Healthy subjects
Distribution of selected demographic variables of OSCC Patients and Healthy subjects
The distribution of selected demographic factors between cases and controls was examined using the Wilcoxon rank-sum test, Pearson’s Chi-square test, and Fisher’s exact test. The goodness-of-fit test was performed to assess any deviation of genotypic frequency in the controls from the Hardy-Weinberg equilibrium (HWE). The odds ratio (ORs), 95% confidence interval (CI), and p-value were used for the analysis and performed using the Chi-square test with five different models; Allele, Co-dominant, Dominant, Recessive, and Heterozygous. All statistical tests were carried out using SPSS-25 software (SPSS, Inc.) and the gtsummary package of R 4.2.1 tool. All p-values were two-sided, and p-value less than 0.05 was considered statistically significant.
Results
Study population and genotyping
The present study had 215 OSCC cases and 215 healthy controls. The distribution of selected demographic variables is summarized in Table 1. The mean age of subjects was 46.2
Genotypic distribution of miR-146a-5p rs2910164 in OSCC patients and healthy control subjects with various genotyping models
Genotypic distribution of miR-146a-5p rs2910164 in OSCC patients and healthy control subjects with various genotyping models
Demographic and clinicopathological details of OSCC patients included in miR-146a expression profiling
The observed genotype frequencies for the rs2910164 were in agreement with that expected under the HWE in the controls (
A) miR-146a expression profile in OSCC tumor and adjacent normal tissue. Figure shows the upregulation of miR-146a in tumor tissue by 1.42-fold in comparison to normal tissue. B) The ENCORI Pan-Cancer analysis of miR-146A expression in Head and Neck Squamous Cell Carcinoma and Overall survival with LogRank 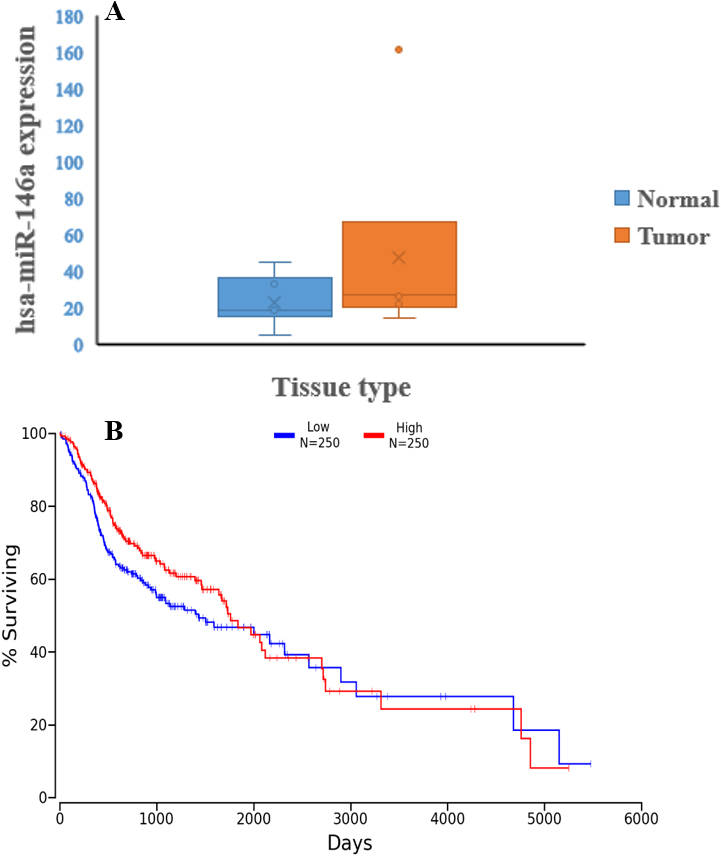
Twenty potential target genes were gained by rs2910164 of miR-146a. The genes were selected as potential targets based on their expression levels in oral cancer.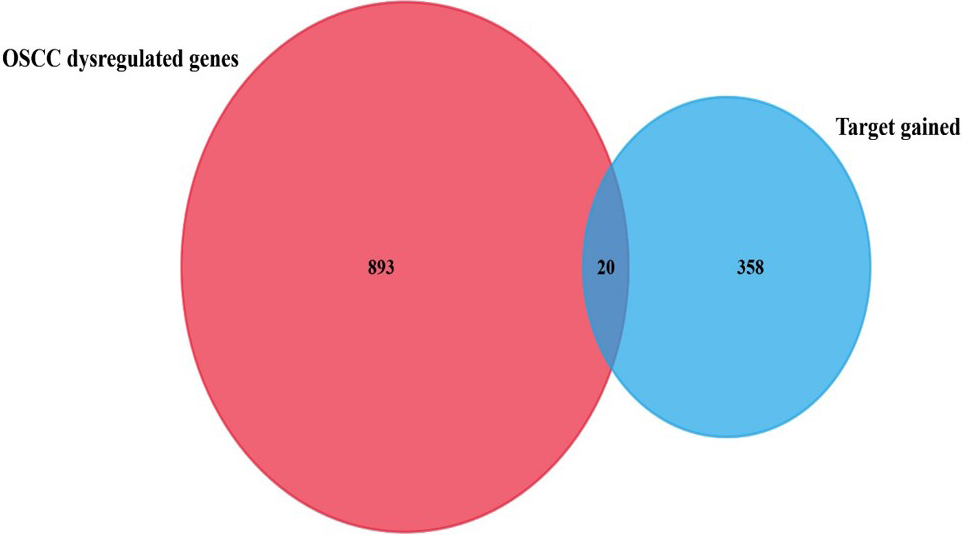
rs2910164 of miR-146A correlation with clinical features of OSCC patients
rs2910164 of miR-146A correlation with clinical features of OSCC patients
The FunRich version 3.1.3 based Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis for target genes gained by miR-146A with rs2910164. A) Activated pathway: Pyrimidine metabolism, Transcriptional activation of p53 and p21, Activation of PUMA and NOXA; B) Transcription factors, C) Biological processes, D) COSMIC.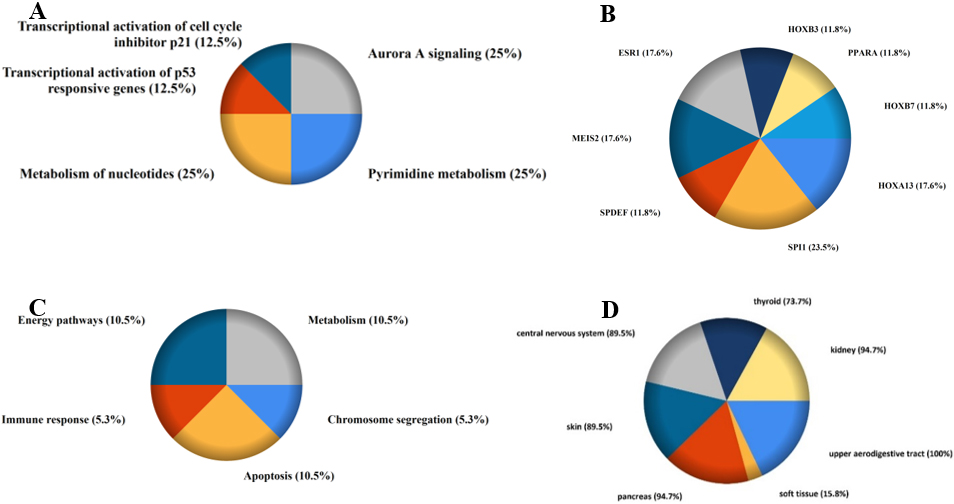
We found that the expression of miR-146a-5p was upregulated in tumor tissue by 1.42-fold compared to normal tissue (Fig. 1A). The survival analysis with univariate Cox proportional hazard regression model showed no significance with up-regulation of miR-146a expression (
The ENCORI Pan-Cancer analysis for final target genes gained by miR-146a with rs2910164. Box plot of gene expression profile in Head and Neck Squamous Cell Carcinoma FBXW7 (A), TP53 (B), SYC (C); Overall survival for FBXW7 in Head and Neck Squamous Cell Carcinoma (D).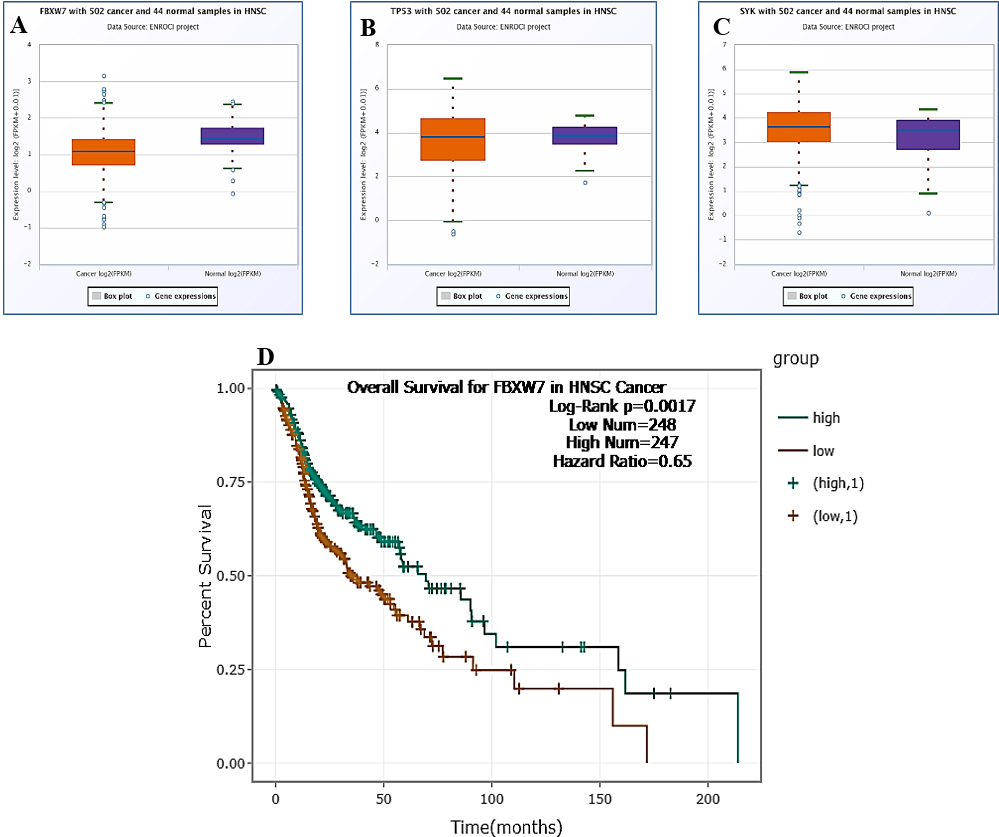
Correlation between miR-146a expression and clinical features of OSCC patients. A) Correlation between miR-146a expression and OSCC staging, B) Correlation between miR-146a expression and histology grade of OSCC, C) Correlation between miR-146a expression and OSCC tumor volume, D) Correlation between miR-146a expression and TNM staging.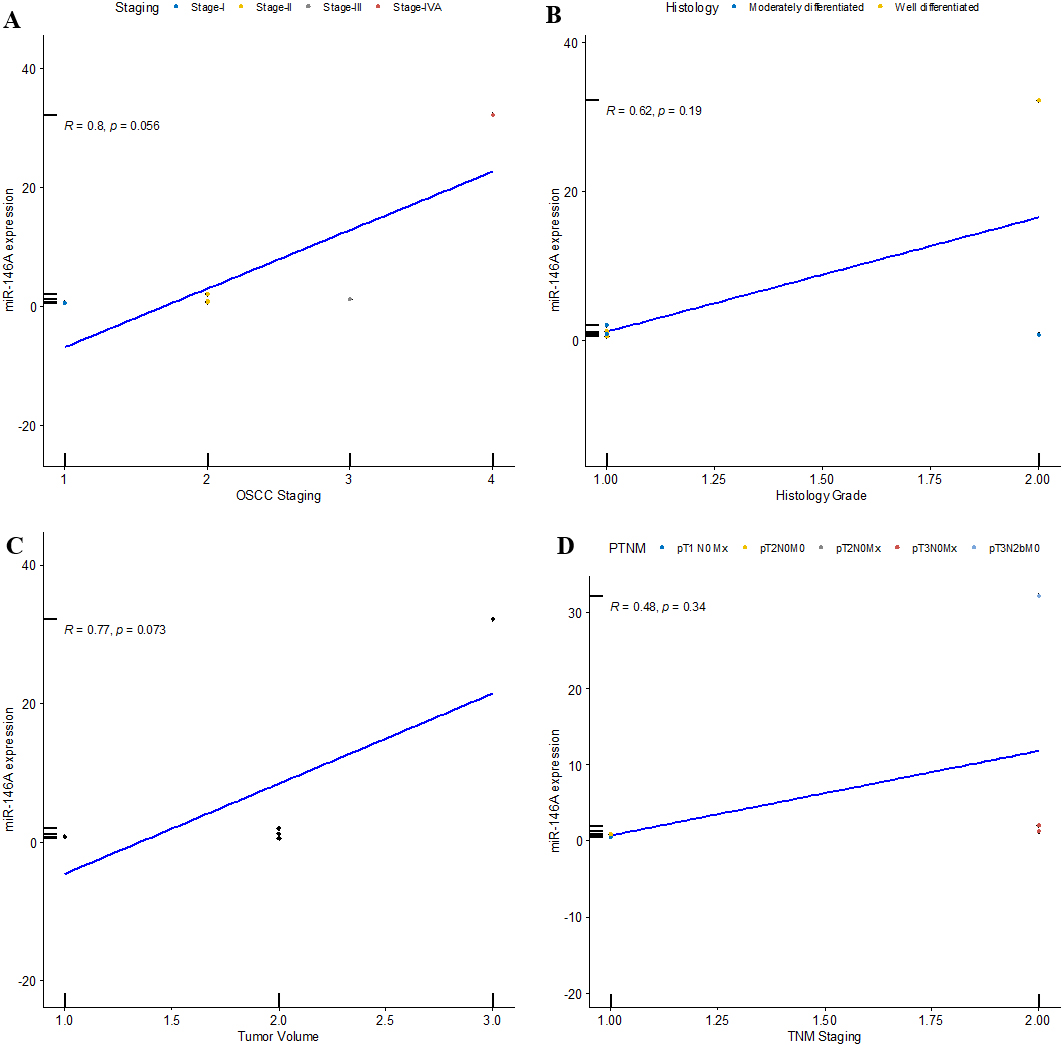
In selected six OSCC patients, there was a strong positive correlation (
In the present study, the prominent lesion site was the Tongue (40%) followed by the Buccal mucosa (37%) in OSCC patients (Table 1). We found that rs2910164 of miR-146a was significantly associated with OSCC risk in the central India population. Specifically, the frequency of the CC genotype was significantly increased in patients with OSCC population, and it was positively correlated with OSCC risk. Conversely, the GC genotype was protective and negatively associated with OSCC risk. The ‘G’ allele was the major allele in both OSCC and healthy populations, and ‘C’ was the alternate allele. In addition, we observed 1.42-fold miR-146a upregulation in tumor tissue, possibly due to rs2910164 SNP in miR-146a. Concurrently, MIRNA SNP DISEASE Database (MSDD) data further supports our findings and suggest that rs2910164 may affect the expression of miR-146a. The elevated miR-146a expression has been reported previously in tumor tissue from OSCC and other cancer types, including breast cancer, hepatocellular carcinoma, ovarian cancer, and lung cancer [21, 22, 23]. Twenty target genes gained by SNP rs2910164 of miR-146a have an essential role in the cell cycle, nucleotide metabolism, and NOTCH pathway, and might directly correlate with tumor development and progression.
miR-146a has shown a crucial role in the expression of epidermal growth factor receptor (EGFR), and TNF receptor-associated factor 6 (TRAF6). It has also been shown to regulate NF-
On the contrary, rs2910164 has also been associated with decreased miR-146a expression in some cancers like papillary thyroid carcinoma, esophageal squamous cell carcinoma, colorectal cancer, and diffuse large B cell lymphoma [32, 33, 34]. An article reviewed the existing data available with the functional role of rs29010164 G/C with miR-146a expression and provided a plausible explanation for the contradictory findings in colorectal cancer [28]. This study found the involvement of unknown mRNA targets and differences in selected individual races are the key factors for contradictory results, and it enhances the need for SNP-based study in a diverse population [28].
We observed that the expression profile of target genes TP53, SYK, and FBXW7 gained by rs2910164 of miR-146a varies in HNSCC (Fig. 4). Pan-Cancer analysis showed that FBXW7 was downregulated in HNSCC cases and significantly correlated with reduced overall survival. FBXW7 is an important transcription factor for reprogramming induced pluripotent stem cells (iPSCs), and its loss of expression correlates with the prediction of recurrence in colorectal liver metastasis [35, 36]. Additionally, regulatory factor as miR-146a might impact FBXW7 expression and function.
Conclusion
In conclusion, the present study shows that the rs 2910164 of miR-146a polymorphism is associated with the OSCC risk. Notably, CC genotype of rs2910164of increases the risk of OSCC in the central Indian population, and the GC genotype may protect the individuals. Furthermore, this SNP can dysregulate miR-146a expression in tumors and results in additional target yield, which affects downstream metabolic pathways, biological processes, and transcription factors involved in OSCC development and progression, especially FBXW7. Therefore, miR-146a might play a crucial role in the pathogenesis of OSCC either by initiating the new lesions or by accelerating the transformation of oral potentially malignant lesions into a malignant disease. Thus, it might help in the screening of individuals at risk for OSCC. Furthermore, future studies are warranted to understand the functional role of rs2910164 in the OSCC pathogenesis at cellular and in vivo levels.
Funding
This work was supported by the Department of Health and Research, New Delhi, to ST (R. 12013/11/2021-HR/E-office: 8111586). In addition, AK would like to thank the Indian Council of Medical Research (ICMR 5/13/93/2020/NCD-III) for the Grant-In-Aid.
Author’s contributions
Conception: Shikha Tiwari, Ritu Pandey, and Ashok Kumar.
Investigation: Ritu Pandey, Supriya Vishwakarma, Tulasi Sindhuja, and Sana Hasmi.
Interpretation or analysis of data: Shikha Tiwari and Ashok Kumar.
Preparation of the manuscript: Shikha Tiwari.
Manuscript editing: Ashok Kumar.
Revision for important intellectual content: Vinay Kumar, Saikat Das, and Ashok Kumar.
Supervision: Vikas Gupta, Rajeev Nema, and Ashok Kumar.
Supplementary data
The supplementary files are available to download from
sj-pdf-1-cbm-10.3233_CBM-230064.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-230064.pdf
Footnotes
Conflict of interest
Authors declare that there is no conflict of interest.
