Abstract
The genes CYP3A4 and CYP3A5 form part of a cluster of cytochrome P450 genes involved in drug metabolism reactions. The allelic variants of these genes CYP3A4*1B, CYP3A4*3, CYP3A4*17 and CYP3A5*3 have been linked both to the reduced catalytic activity of cytochromes and to prostate cancer risk in whites, though scarce data exist for North African populations. The main objective of this study was to describe CYP3A4*3, CYP3A4*17, CYP3A4*1B and CYP3A5*3 allele frequencies and haplotype variation in Moroccan Berbers and the general Tunisian population. The data obtained for the Tunisian participants were consistent with the European allele frequency ranges described, while Moroccan Berbers showed high frequencies of CYP3A4*17 (1.8%), CYP3A4*3 (8.5%) and the CYP3A4*1B/CYP3A5*3 haplotype (18.4%). This haplotype, linked to an increased risk of prostate cancer, was detected at a much higher frequency compared with the present Tunisian population (8.4%) or with reported frequencies for populations such as whites (0.6%) or African Americans (5.3%).
Introduction
Cytochromes (CYPs) are the main enzymes involved in drug metabolism and bioactivation, and are responsible for around 75% of all metabolic reactions. CYP-regulated drug metabolism shows genetic variability, which can lead to normal, low or null activity levels of a given enzyme. The allelic variants of the genes CYP3A4 (CYP3A4*3 and CYP3A4*17) and CYP3A5 (CYP3A5*3) induce structural differences in the enzymes they encode, causing a >99% reduction in their catalytic activity (1). Some CYP3A4 and CYP3A5 variants have been linked to prostate cancer risk and severity in whites (2). However, so far the literature lacks data on the prevalence of these mutations and haplotypes in North African populations.
Genetic variants of CYP3A4 and CYP3A5, the major forms of CYP450, show marked differences among populations. Some CYP3A4 and CYP3A5 genetic variants have been associated with differential drug metabolism and cancer incidence. The CYP3A4*1B variant features an A to G change in the 5′-flanking region. Whereas the 1B or G allele has been linked to an increased risk of prostate cancer among African populations, this association has not been observed among whites or Asians (3). The CYP3A4*1B allele occurs more frequently in South African and Nigerian than European populations (4). CYP3A4*3 carries a 1331T>C change that gives rise to a Met445Thr substitution, and CYP3A4*17 is a rare 15615T>C variant whose product has a Phe189Ser substitution, purportedly rendering it a poor metabolizer of drugs such as nifedipine (1). Both alleles, CYP3A4*3 and CYP3A4*17, appear at very low frequencies in white groups (5). CYP3A5*3 bearing 6986G>A encodes a nonfunctional allele. The authors of a recent meta-analysis suggested that the CYP3A5*3 polymorphism may play a role in the development of leukemia and colorectal cancer in Asians and whites (6) but not in African individuals. High levels of genetic diversity have been observed for CYP3A5*3 within east African populations (7).
This study was designed to describe CYP3A4*3, CYP3A4*17, CYP3A4*1B and CYP3A5*3 allele frequencies and CYP3A4*1B-CYP3A5*3 haplotype variation in DNA samples from 2 North African populations: one of Moroccan Berbers and the other of the general Tunisian population (Arab-Berber); both these populations have been examined for other polymorphisms (8, 9). North African populations (Moroccans, Algerians, Tunisian and Libyans) have a similar structure. Each one is composed of a general population of Arabic speakers (a mixture mainly of Berbers and Arabs) and Berbers (few in Tunisia, an intermediate number in Libya and abundant in Morocco and Algeria), who often speak both Berber and Arab languages (9).
Materials and Methods
DNA samples
The DNA samples (peripheral blood) examined were obtained from 97 participants from the Khenifra region (Middle Atlas) of Morocco and 102 participants from Tunisia. All participants were healthy (both sexes), unrelated donors aged 18-60 years, who signed an informed consent form approved by the ethics committees of Chouaib Doukkali University (Morocco) and Monastir University (Tunisia).
Blood samples from Berbers were collected in Khénifra (32.93°N, 5.67°E) in the Middle Atlas region. Most Berbers in this region belong to the Zayane confederation (a group of neighbor tribes). Khénifra is one of the main Berber towns in Morocco. The Middle Atlas region covers an area of 13,140 km2 in the center-south of the country, and has 3 cities (219, 168 inhabitants) and 35 rural communities (245,893 inhabitants). The Moroccan Berber participants were Tamazight speakers. Tunisian individuals were Arab speakers representative of the country's general population. Samples were collected from 84 participants in the central-northern region of Tunisia and from 18 individuals in the southern Tunisian regions of Gabès, Gbelli and Mednine.
Genotyping
Genomic DNA was extracted from peripheral blood using standard phenol-chloroform procedures. Two CYP3A4 single nucleotide polymorphisms (SNPs) (CYP3A4*3 -rs4986910- and CYP3A4*17 -rs4987161-) (1, 5) and 1 CYP3A5 SNP (CYP3A5*3 -rs776746-) (10) were genotyped by PCR–restriction fragment length polymorphism (RFLP) according to procedures described for each (1, 5, 10). The CYP3A4*1B A-G polymorphism was detected using the Q-PCR standard SDS allelic discrimination assay protocol (11).
Statistical analysis
For each polymorphism, allele frequencies were estimated through direct gene counts, and Hardy-Weinberg equilibrium was assessed using an exact test (12). Maximum likelihood haplotype frequencies were computed using the expectation–maximization (EM) algorithm. Gene and haplotype diversities were estimated according to Nei's equation (13). Populations were compared using the exact test of population differentiation. All statistical tests were performed using the Genepop 3.3 (14) and Arlequin software packages (15). Significance was set at a p value <0.05.
Results and Discussion
Allele frequencies for CYP3A4 and CYP3A5 SNPs are reported in Table I for the general Tunisian and the Moroccan Berber population samples. Fifteen percent of Moroccan samples could not be genotyped due to technical problems. Both sets of data fulfilled Hardy-Weinberg equilibrium proportions for all polymorphisms except CYP3A4*17 in Tunisia (due to lack of polymorphism) and CYP3A4*3 in Morocco (p<0.001). Moroccan Berbers returned considerably higher diversity values as compared with the general Tunisian population: heterozygosity means (±SD) for the 4 SNPs were 0.178 ± 0.144 vs. 0.107 ± 0.113; excluding CYP3A4*17, means were 0.225 ± 0.127 vs. 0.143 ± 0.107. Population comparisons revealed significant differences between Moroccan Berbers and Tunisians in allele frequencies obtained for the CYP3A4*1B and CYP3A4*3 SNPs (Fisher exact probabilities <0.001 in both cases).
CYP3A4 and CYP3A5 Allele Frequencies in Moroccans and Tunisians
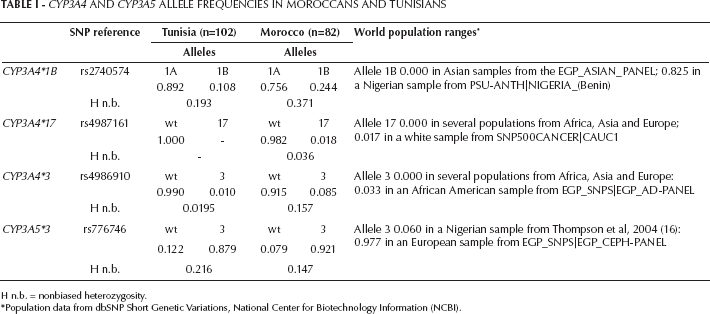

H n.b. = nonbiased heterozygosity.
Population data from dbSNP Short Genetic Variations, National Center for Biotechnology Information (NCBI).
World population ranges for these SNPs are indicated in Table I. Although both data sets fell within the ranges described for white individuals, our Moroccan Berber sample showed high frequencies of the derived alleles CYP3A4*17 and CYP3A4*3. Thus, the CYP3A4*17 allele in Morocco was detected at a frequency of 0.018, comparable to the highest value reported for a white sample (0.017; see population references in Table I). The CYP3A4*3 variant in Morocco showed a frequency of 0.085. This figure is considerably higher than that reported in African Americans (from sub-Saharan Africa) showing the highest frequency (0.033), or the frequency observed here for the general Tunisian population (0.010) (see Tab. I).
Haplotype diversities for the 2 populations examined are shown in Table II. Haplotypes were calculated by including the 4 SNPs and considering only CYP3A4*1B and CYP3A5*3 because information about the combination of these 2 variants was available for some populations (2). Moroccan Berbers displayed higher haplotype diversities than the general Tunisian population (0.178 vs. 0.107 for haplotypes including 4 SNPs, and 0.259 vs. 0.205 for haplotypes with 2 SNPs, respectively). Population comparisons indicated significant differences (p<0.001) in both cases.
CYP3A4 and CYP3A5 Haplotype Frequencies in Moroccans and Tunisians
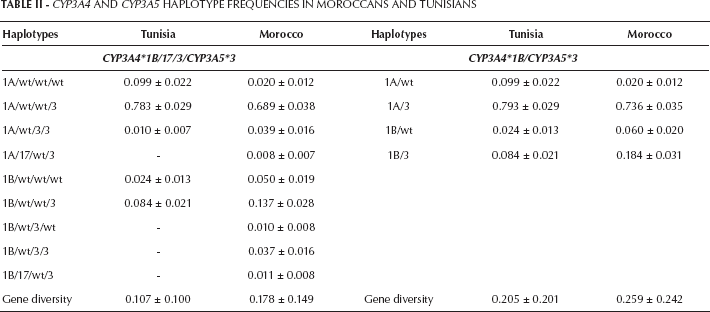
The CYP3A4*1B/CYP3A5*3 haplotype has been linked to more severe forms of prostate cancer among European American and African American men (2). The authors of this last study detected the high-risk CYP3A4*1B/CYP3A5*3 haplotype at a higher frequency in African Americans than in European American individuals (5.3% vs. 0.6%). In our study, this haplotype was found at considerably higher frequencies than reported by Plummer et al (2): 8.4% in the Tunisian population and 18.4% in the Moroccan Berbers. The 2 variants giving rise to this haplotype, CYP3A4*1B and CYP3A5*3, have been related to several cancers such as prostate and colorectal cancer, and chronic and acute leukemia (3, 6). These cancers show different risk values depending on ethnic group, with Asians and whites in 1 such group and sub-Saharan Africans in another group. Whereas the CYP3A5*3 variant produces a nonfunctional allele, the CYP3A4*1B mutation affecting the 5′-flanking region has no apparent repercussions on the final gene product. Some authors (2, 3) have suggested that other functional variant(s) in the haplotype could underlie the links reported.
In this survey, we examined the general population of Tunisia and a group of Berber individuals from the Middle Atlas region of Morocco in terms of their CYP3A4/CYP3A5 variability. Given the ethnic and geographic peculiarities of North African populations and the scarce data available for these genes in North African individuals, we compared the allele frequencies obtained here with those reported for other populations. Thus, our data for the Tunisian population sample falls within the ranges described for several European samples (see Tab. I), whereas the Moroccan Berbers showed high frequencies of the CYP3A4*17 (1.8%) and CYP3A4*3 (8.5%) variants, and CYP3A4*1B/CYP3A5*3 haplotype (18.4%). The CYP3A4*1B/CYP3A5*3 high-risk haplotype appearing at a considerable frequency in the Moroccan population has implications for its use as a biomarker for prostate cancer. Our findings prompt further studies designed to examine cytochrome gene variability in North African individuals and patients with prostate cancer or other cancers associated with CYP3A4 and CYP3A5.
Footnotes
Acknowledgements
The authors thank Drs. Wifak El Moncer, Hassen Chaabani and Pedro Moral for their help with the scientific assistance and blood sample collection process.
