Abstract
In the treatment of lung cancer, radiotherapy has become one of the most important therapies, despite its sometimes unpredictable side effects. As such, identifying lung cancer patients who are at high risk of developing severe radiation-induced damage (mainly radiation pneumonitis and radiation-induced esophageal toxicity) and applying effect intervention or monitoring techniques are important. Although human diversity to a certain amount is explained by clinical and dosimetric factors, the presence of specific genetic determinants also influences the occurrence of radiation-induced damage. Here we summarize the data on mechanisms of radiation pneumonitis and radiation-induced esophageal toxicity supporting the involvement of variances of genes in the evolution of radiation-induced damage. Furthermore, the available evidence from current clinical studies of genetic polymorphisms for the prediction of radiation pneumonitis and radiation-induced esophageal toxicity is discussed. Eventually, this may help to truly individualize radiotherapy, using a personal genetic profile of the most relevant genes for each lung cancer patient.
Keywords
Introduction
Lung cancer, including non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC), represents the leading cause of cancer-related mortality worldwide. A total of 224,210 new cases and 159,260 deaths are projected to occur in the United States in 2014 (1). Although in recent years, significant advances have been made in our understanding of the molecular pathogenesis and treatment of lung cancer, the survival benefits are still limited.
Radiotherapy (RT) is one of the most important treatment mortalities in lung cancer patients. In general, greater than 50% of all lung cancer patients require RT (either for thoracic disease or for extrathoracic metastatic sites), with stereotactic ablative RT considered the preferential option for medically inoperable stage I NSCLC and 3-dimensional conformal radiotherapy (3D-CRT) or intensity-modulated radiotherapy (IMRT) referred to as one of the standard treatment modalities for patients with unresectable or recurrent lung cancer (2–3–4–5). Given the recent seminal advancements in RT, it may be postulated that the need for RT is increasing. However, even with technological advancements in patient immobilization, setup verification and radiotherapy delivery, thoracic RT is commonly accompanied by a high risk of radiation-induced damage, which can not only hamper radiation dose escalation, but also negatively impact quality of life and impose a considerable economic burden, or even influence the prognosis (6). Furthermore, metrics to diagnose and explain the severity of radiation-induced damage are somewhat limited. As such, accurate approaches for identification of lung cancer patients at high risk of developing radiation-induced damage are required for optimizing the quality of RT planning and minimizing the treatment-related side effects, and/or intensive monitoring to optimize the likelihood of timely and successful intervention.
Radiation pneumonitis (RP) and radiation-induced esophageal toxicity (RIET) have been identified as 2 major dose-limiting complications of thoracic RT. For RP, approximately 15% to 45% of patients will develop RP up to 2 months after completion of the RT course (7–8–9–10), with mild dry cough, fever, mild dyspnea and a late phase of pulmonary fibrosis representing major clinical manifestations. Several patient-related factors have been reported to influence susceptibility to RP, including performance status, tumor location, smoking history and concurrent chemotherapy (7, 11). Recently, it has been demonstrated that V20 (volume of total lung receiving 20 Gy or more), V5 (volume of total lung receiving 5 Gy or more) and mean lung dose (MLD) independently predict the risk of RP in lung cancer patients (12–13–14). For RIET, it is found that around 22% patients develop grade 3 acute RIET in NSCLC patients with concurrent chemoradiotherapy (15). This complication, characterized by dysphagia, odynophagia, esophagus stenosis and perforation, to name a few, can necessitate hospitalization, initiation of tube or intravenous feeding and treatment interruptions, or even affect the long-term quality of life. As reviewed in the literature, radiation dose, concurrent chemotherapy regimens and other clinical and dosimetric factors are correlated with risk of ≥ grade 2 acute RIET (15–16–17–18). A recent study has shown that equivalent uniform dose and V76.7 to the esophagus were significantly associated with severe RIET (19). However, there still exists heterogenicity among lung cancer patients exposed to the same dose range or volume of irradiation, suggesting that genetic predisposition may be involved in RP and RIET development. Thus, it would be of significant benefit to identify additional genetic factors that are likely to predict RP and RIET susceptibility.
Up to now, analysis of gene expression has not resulted in a satisfied prediction of occurrence in clinical practice. Another active area of research in looking for biomarkers in blood is through the study of single nucleotide polymorphisms (SNPs), which are single-base inheritable variations in a given and defined genetic location and have been well recognized as useful markers in genetic association studies for disease susceptibility or adverse drug reactions. Studies documenting SNPs coincident with cis-regulatory elements as critical regulators in defining human diversity and disease by regulating the nature and timing of gene expression are also available (20–21–22). Given their significance in interindividual differences, research through the study of SNPs into adverse responses of normal tissues is rapidly increasing. These results indicate the significant differential expression and functional role of SNPs in the prediction of radiation-induced toxicities, suggesting that SNPs could potentially serve as their biologic markers (23–24–25–26). However, the search for SNPs as predictive biomarkers has just begun, with no definitive conclusion yet reached as to which patients might benefit from such a strategy. The following review discusses the role of genes involved in the mechanism of radiation-induced damage, especially in RP and RIET. Moreover, a broad overview of the potential value of SNP models to identify lung cancer individuals at risk for RP and RIET during the early stages of RT is presented based on the available evidence.
Current understanding of radiation-induced damage
The mechanism of radiation-induced damage remains obscure; however, one thing is certain, that is, the development of injury to surrounding health tissue is regarded as continuous molecular events proceeding. Indeed, there is evidence that a combination of direct radiation cytotoxicity to surrounding health normal tissue and secondary inflammatory changes may contribute to the development of the major radiation-induced toxicities (RP and RIET) in lung cancer patients (27).
RP
It has been indicated that RP is the result of a dysregulated wound-healing inflammatory reaction (28). An immediate response to thoracic irradiation is the generation of reactive oxygen species (ROS) or nitrogen (RNS) species that can cause oxidative damage to DNA, lipids and proteins, leading to type I and II pneumocyte injury or apoptosis (29). Subsequently, type I cells are thought to be denuded, and the proliferation of type II pneumocytes is thought to be stimulated. Secretion of growth factors and degradation of the extracellular matrix allow the removal of dead cells (30). During the process, the loss of epithelial or endothelial cells or vessel integrity, the reduction of microvessel density and lung perfusion, and hypoxia give rise to secondary inflammatory processes (28). Several cytokines, including interleukins, tumor necrosis factor and transforming growth factor β1 (TGF-β1), to name a few, are critical components for modulating and ameliorating the effects of inflammatory reactions. Meanwhile, more ROS is triggered by hypoxia, leading to chronic oxidative stress, fibrosis via TGF-β1, angiogenesis via vascular endothelial growth factor (VEGF) production and ultimately nonhealing tissue injury in the lung (30).
Riet
RIET is also regarded as a dysregulated wound-healing inflammatory reaction to tissue injury. The initial process may involve direct action of ROS on DNA, which causes DNA breaks and rapidly triggers the production of cytokines and growth factors. Then the production of inflammatory cytokines, including TGF-β1 and cyclooxygenase-2 (COX-2), is the result of an activation of an inflammatory reaction leading to the expression and maintenance of an elevated cytokine cascade. During the process of repair, both activation of inflammation and vascular changes as well as stress response contribute to the development of hypoxia, which further stimulates inflammatory cells recruited to an area of injury, leading consequently to chronic radiation injury.
As described above, pathways of DNA repair, inflammation, stress response and angiogenesis participate in the development of radiation-induced damage. Because genetic factors can extend the wound-healing process or intensify inflammatory reactions, variance of genes may have functional consequences resulting in total loss or diminished expression of the encoding gene, or functional alterations. Substantial clinical efforts have been made regarding the prediction of relevant gene SNPs for RP and RIET in patients with lung cancer.
Methods
Eligible studies were identified by searching the PubMed and Cochrane databases for relevant reports before July 2014, using different combinations of the following search terms: “SNP” and “lung cancer” and “radiation pneumonitis/radiation-induced lung toxicity” or “radiation esophagitis/radiation-induced esophageal toxicity.” Only English-language literature data and the most recent or the most complete studies were used. Meeting abstracts, unpublished reports and review articles were not considered.
Clinical evidence of gene polymorphisms in the prediction of radiation-induced damage in lung cancer patients
Recently much work on promising gene polymorphisms for prediction of radiation-induced toxicities (RP and RIET) has been done in lung cancer patients. As SNPs can be detected more easily in the blood than in neoplastic tissues, genetic analysis in the circulating system has always been used to predict RP and RIET in lung cancer patients.
Predictive SNPs for RP in lung cancer patients
Predictive SNPs of DNA repair genes
DNA repair pathways are divided into single-strand break repair, which includes nucleotide excision repair (NER), base excision repair (BER), as well as mismatch repair and double-strand break (DSB) repair, consisting of homologous recombination repair and nonhomologous end joining (NHEJ) (31). A lot of genes, including ataxia-telangiectasia mutated gene (ATM), RAD51, X-ray repair cross complementing genes (XRCC) and apurinic/apyrimidinic endonuclease 1 (APE 1), are involved in these pathways. Most of these genes have been shown to contribute to interindividual variability of DNA repair in responses to RT in lung cancer patients.
Predictive SNPs of DSB repair genes
ATM
ATM is a critical mediator in the homologous recombination pathway. It has been found that ATM not only participates in detection and repair of double-strand breaks, but also controls cell cycle checkpoints and apoptosis. Recent experiments have shown that ATM can promote autophagy in response to radiation and other genotoxic or oxidative stimulants via the MAPK14 pathway, the mTOR pathway, and Beclin/PI3KIII complexes, which contribute to the effect of ATM on radiosensitivity (32–33–34). The association between ATM variants and RP has been well explored by Zhang et al (35). A total of 253 lung cancer patients receiving thoracic irradiation between 2004 and 2006 were included in this study. In this study, 2 ATM variants from blood lymphocytes, consisting of ATM-111G>A polymorphism and 126713 G>A polymorphism, were demonstrated to be associated with increased RP risk (hazard ratio [HR] = 2.49 and 2.47; 95% confidence interval [95% CI], 1.07-5.80 and 1.16-5.28, respectively). These differences were also verified both in cell lines (p<0.001) and in individual lung tissue samples (p=0.003; Tab. I) (35). Further study from the same group found that the ATM-111G>A and P53 Arg72Pro polymorphisms displayed an combinative effect in predicting the risk of developing RP, with the cross-validation test showing positive results in 63.2% of cases (Tab. I) (36). Xiong et al (37) conducted a comprehensive analysis of ATM SNPs for the prediction of RP. The data from 362 NSCLC patients (72.4% of non-Hispanic whites) receiving definitive radiation therapy identified that patients carrying ATM rs189037 AG/GG or rs228590 TT/CT genotypes or rs189037G/rs228590T /rs1801516G (G-T-G) haplotype had a lower risk of ≥ grade 3 RP (rs189037: GG/AG vs. AA, HR=0.49, 95% CI, 0.29-0.83, p=0.009; rs228590: TT/CT vs. CC, HR=0.57, 95% CI, 0.33-0.97, p=0.036; haplotype: G-T-G vs. A-C-G, HR=0.52, 95% CI, 0.35-0.79, p=0.002; Tab. I). As has been implied, ATM polymorphisms may serve as biomarkers for susceptibility to severe RP in lung cancer patients. However, further study of this topic is needed.
Studies Addressing Impact Of Snps On Risk Of Rp In Lung Cancer Patients Undergoing Rt
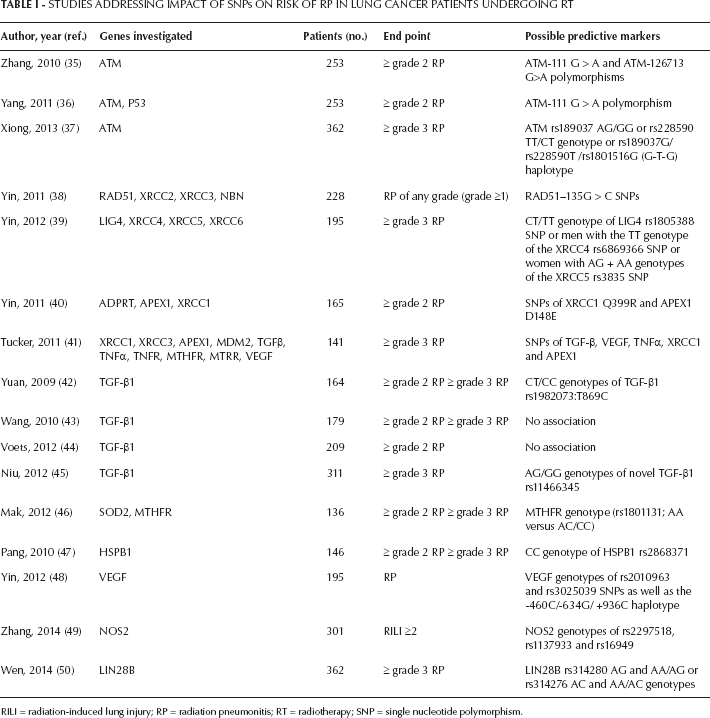
RILI = radiation-induced lung injury; RP = radiation pneumonitis; RT = radiotherapy; SNP = single nucleotide polymorphism.
RAD51
RAD51, another important DSB repair gene in the homologous recombination repair pathway, has been demonstrated to confer radioresistance in previous studies (51, 52). To further investigate its predictive potential as secreted biomarkers for severe RP, its variations were explored. In a retrospective study, clinical data were derived from 196 NSCLC patients with pneumonitis information between 1999 and 2005. Results revealed an increased risk of ≥ grade 3 RP was found in patients with RAD51-135G>C SNPs (adjusted HR=0.52, 95% CI, 0.31-0.86, p = 0.010 for CG/CC vs. GG) (Tab. I) (38). Future studies are warranted to address its relevance in the clinical setting.
Other genes
Some other genes in the NHEJ pathway have also been found to be useful. One such study was published by Yin et al (39), who reported a predictive role of the DNA ligase IV (LIG4) rs1805388 SNPs in ≥ grade 3 RP development (adjusted HR=2.08; 95% CI, 1.04-4.12; p=0.037 for the CT/TT genotype vs. the CC genotype). In addition, men with the TT genotype of the XRCC4 rs6869366 SNP and women with AG+AA genotypes of the XRCC5 rs3835 SNPs were at increased risk of developing severe RP (Tab. I). Future efforts toward better comprehension of DNA repair may help in discovering more potential markers.
Predictive SNPs of single-strand break repair genes
XRCC1
XRCC1 is a critical repair gene involved in BER, a major system responsible for efficient repair of DNA single-strand breaks generated by RT. Data reveal that the XRCC1 mutant plays a novel role in increasing the retention of XRCC1 at DNA damage sites and in DNA alkylation damage repair (53, 54). Its polymorphisms for predicting radiation-induced response have also been verified in a wide range of human malignancies, including lung cancer, breast cancer, nasopharyngeal cancer and rectal cancer (55–56–57–58–59). One such example was presented in a recent study of 165 patients with NSCLC who received definitive chemoradiation therapy (Tab. I) (40). Of the 2 SNPs tested, SNPs of XRCC1 Q399R were shown to have a significant effect on the development of ≥ grade 2 RP (AA vs. GG, adjusted HR=0.48, 95% CI, 0.24-0.97). Subsequently, the predictive roles of XRCC1 SNPs were further verified by the same group in 141 NSCLC patients treated with definitive radiation therapy, with or without chemotherapy (Tab. I) (41).
APE 1
APE1 is another key gene in the BER pathway. In addition to its DNA repair function, APE1 maintains a number of transcriptional factors including p53 in their reduced and active state, thereby regulating their DNA-binding activity, influencing gene expression and maintaining genomic stability (60). Recently, most analyses have shown that a decrease in APE1 levels leads to enhanced cell sensitization to ionizing radiation in lung cancer, colorectal cancer and hepatocellular carcinoma, as well as human osteogenic sarcoma cells in vitro and in vivo (61–62–63–64). In RP induced by thoracic RT, it has been documented that APEX1 SNP is also involved in RP risk. For example, Yin et al (40) conducted a retrospective study with 165 NSCLC patients receiving definitive chemoradiation therapy. He found that SNP of APEX1 D148E had a significant effect on the development of ≥ grade 2 RP (APEX1: GG vs. TT, adjusted HR=3.61, 95% CI, 1.64-7.93) in an allele-dose response manner (trend tests: p=0.001) (Tab. I). Subsequently, the same group reached a similar result by verifying APEX1 SNP in 141 NSCLC patients treated with definitive radiation therapy, with or without chemotherapy (Tab. I) (41). In all, these studies support a role of single-strand break repair genes in the prediction of severe RP. However, further evidence is still pending.
Predictive SNPs of cytokine genes
There have been significant research efforts to identify specific cytokines as risk-predicting biomarkers, among which TGF-β1 has been the most frequently studied. Preclinical studies demonstrate that TGF-β1 could regulate tissue morphogenesis, inhibit the proliferation of many cell types and stimulate the proliferation of fibroblasts and endothelial cells, suggesting that it may play an important role in the pathogenesis of RP (65–66–67). Clinical studies have further shown that changes in plasma TGF-β1 levels are associated with the development of RP (68–69–70). The predictive value of plasma TGF-β1, however, is confounded by the tumor effect, since lung cancer always produces TGF-β1. Additionally, there is concern that numerous factors, including improper handling of blood samples or inadequate centrifugation conditions, can falsely increase the calculated level of circulating TGF-β1 (71).
In recent years, TGF-β1 SNPs have not only been associated with radiation sensitivity, but have also been implicated as markers of radiation-induced toxicity risk in lung cancer, although studies have been somewhat confusing (72–73–74–75–76–77–78, 42–43–44). As for TGF-β1 SNPs in NSCLC, Yuan et al (42) tested 3 SNPs of TGF-β1 (rs1800469:C-509T, rs1800471:G915C and rs1982073:T869C) in 164 patients of mixed ethnicity with TGF-β1 genotype distributions similar to white patients in general. Compared with the TT homozygotes, CT or CC genotypes of rs1982073:T869C were associated with a statistically significantly lower risk of ≥ grade 2 RP (HR=0.489; 95% CI, 0.227-0.861; p=0.013) and ≥ grade 3 RP (HR=0.390; 95% CI, 0.197-0.774; p=0.007), respectively (Tab. I). No such associations were observed in the Chinese and European population (43, 44). Further study suggested these polymorphisms were inconsistent with data from plasma TGF-β1 levels (43). As such, Niu et al (45) conducted a more comprehensive analysis of TGF-β1 SNPs for predicting RP in Chinese NSCLC patients. Eight SNPs of the TGF-β1 gene (rs1800469, rs1800471, rs1982073, rs4803455, rs11466345, rs12983047, rs10417924 and rs10980942) were genotyped by the direct DNA sequencing method (Tab. I). Data showed that only the AG/GG genotypes of TGFβ1 rs11466345 were associated with a higher risk of RP in patients treated with definitive RT, after adjustment for age, smoking status and dosimetric parameters (for MLD, adjusted HR=2.295, 95% CI, 1.101-4.783, p=0.027; for V20, adjusted HR=2.142, 95% CI, 1.033-4.441, p=0.041) (Tab. I). Since SNPs have a significant ethnic variation, which may account for differences in individual disease susceptibility, we propose that significant interethnic variations exist in the TGF-β1 gene. Therefore, validation of TGF-β1 SNP associations found in different ethnic populations may be an important consideration in RP risk evaluation.
Predictive SNPs of oxidative stress response genes
There is strong support in the literature that elevated oxidative stress and related response pathways can be associated with the generation of radiation-induced healthy tissue damage (79). Thus, Mak et al (46) conducted a retrospective study of superoxide dismutase 2 (SOD 2) and methylene tetrahydrofolate reductase (MTHFR) from oxidative stress pathways, in a sample of 136 lung cancer patients. Data showed that after adjusting for clinical factors and lung dosimetric parameters only the MTHFR genotype (rs1801131; AA versus AC/CC), a key regulator of folate, homocysteine, thiol, methylation and thymidine metabolism, was significantly associated with risk of ≥ grade 2 RP (HR=0.37; 95% CI, 0.18-0.76; p=0.006, corrected p=0.018) and ≥ grade 3 RP (HR=0.21; 95% CI, 0.06-0.70; p=0.01; corrected p=0.03) (Tab. I). Another prime candidate for stress response induced by ionizing radiation is heat-shock protein (HSP), among which HSP27 is highly inducible in the lung in response to oxidative stress and inflammatory mediators such as tumor necrosis factor-α (TNFα). The large oligomeric form of HSP27 is also believed to have HSP27-mediated resistance to radiation, and alteration of normal HSP27 oligomerization may be a good strategy for abolishing HSP27-mediated radioresistance (80). As plasma concentrations of HSP27 protein are predominantly under genetic control by HSPB1, Pang et al (47) tested 2 functional SNPs of HSPB1 (rs2868370 and rs2868371) in NSCLC patients and evaluated the association of SNPs with RP risk. The data from 271 patients treated with chemoradiation at 1 institution identified serum CC of HSPB1 rs2868371 had a higher risk of ≥ grade 3 RP than CG/GG after adjustment for sex, age, performance status and MLD (Tab. I). These results provide clinical evidence supporting a role of stress response pathways in RP. However, published data are far too limited to draw any firm conclusions on their predictive value.
Predictive SNPs of the VEGF gene
It is well established that RT can exert an antiangiogenic effect and induce apoptosis of vascular endothelial cells, which can be observed in both malignant and normal tissues (48). As such, it is likely that a combination of RT with antiangiogenic agents can enhance the radiosensitivity of tumors and their surrounding normal tissues, which may conversely lead to increased radiation-induced toxicities. This has been verified in some lung cancer patients (81). Recently, SNPs of VEGF, a major driver of angiogenesis, have been shown to have an predictive impact upon RP. Such a function was proposed by Yin and colleagues (82) who reported the VEGF genotypes of rs2010963 and rs3025039 SNPs as well as the -460C/ -634G/+936C haplotype were predictors of RP development in 195 NSCLC patients from the University of Texas MD Anderson Cancer Center, United States, with definitive RT (adjusted HR=2.33, 95% CI, 1.01-5.37, p=0.047 for CC vs. GG genotypes; adjusted HR=28.13, 95% CI, 5.24-151.02, p<0.001 for TT vs. CC genotypes; and adjusted HR=2.51, 95% CI, 1.27-4.98, p=0.008 for T-C-T vs. C-G-C haplotypes) (Tab. I). These results correspond with those of Tucker et al (41). Hence, SNPs of VEGF gene may be used as a biomarker for the prediction of RP.
Predictive SNPs of other relevant genes
Nitric oxide synthase (NOS2) involved in cytotoxicity, inflammation and fibrosis encodes a protein called nitric oxide (NO) which has been found to be associated with radiation-induced lung injury (RILI) in lung cancer patients (49). It has been demonstrated that 3 SNPs (rs2297518, rs1137933 and rs16949) in NOS2 genotyped from a cohort of 301 Chinese patients who received definitive radio(chemo)therapy were significantly associated with risk of RILI ≥2 (p=0.001, p=0.000092 and p=0.001, respectively) after adjusting for other covariates (Tab. I) (49). Further haplotype analysis indicated that the ATC haplotype of the above 3 SNPs is associated with reduced risk of RILI. LIN28 is an evolutionarily conversed RNA-binding protein that can bind to the terminal loops of let-7 family microRNA precursors and block their processing to maturation (83). It has been reported that LIN28 is not only highly expressed in several subsets of tumors that carry poor prognoses, but is also involved in tissue inflammatory response (83–84–85–86). After genotyping 8 potentially functional SNPs of LIN28A and LIN28B in 362 patients with NSCLC, Wen et al (50) found patients carrying LIN28B rs314280 AG and AA/AG or rs314276 AC and AA/AC genotypes had a higher risk of ≥ grade 3 RP (for rs314280 AG and AA/AG vs. GG, adjusted HR=2.97 and HR=2.23, 95% CI, 1.32-6.72 and 1.01-4.94, p=0.009 and p=0.048, respectively; for rs314276 AC and AA/AC vs. CC, adjusted HR=2.30 and HR=2.00, 95% CI, 1.24-4.28 and 1.11-3.62, and p=0.008 and p=0.022, respectively) (Tab. I) – that is, genetic variants of LIN28B may be biomarkers for susceptibility to severe RP in NSCLC patients. However, large, prospective studies are needed to confirm the above 2 findings.
Predictive SNPs for RIET
Since very few potential SNPs have been identified in RIET besides SNPs of TGF-β1, XPD and HSPB1, we focus on SNPs of these genes in this section.
Predictive SNPs of TGF-β1
Aside from particular gene polymorphisms for RP, it has been shown that the CT/TT genotype of TGF-β1 rs1800469:C-509T may be a predictive biomarker for RIET in lung cancer patients. In a retrospective study of 97 NSCLC patients with IMRT and 101 patients with 3D-CRT, univariate and multivariate analysis both demonstrated a statistically higher risk of ≥ grade 3 RIET in patients with the CT/TT genotypes of TGF-β1 rs1800469:C-509T when compared with the CC genotype (p=0.026 and p=0.045, respectively) (Tab. II) (87). Another such study of 213 stage III Chinese lung cancer patients receiving RT showed that the CT/TT genotypes of TGF-β1 C-509T were associated with a significantly higher ≥ grade 2 RIET risk (adjusted HR=2.47; 95% CI, 1.17-5.24; p=0.018, and HR=3.86; 95% CI, 1.50-9.92; p=0.005, respectively) compared with the CC genotype (Tab. II) (88). It may be that differences in the mechanism between RP and RIET is the reason for this discrepancy.
Studies Addressing Impact Of Snps On Risk Of Riet In Lung Cancer Patients Undergoing Rt
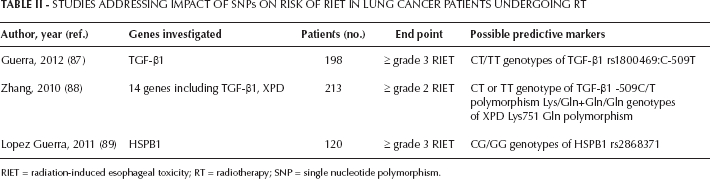
RIET = radiation-induced esophageal toxicity; RT = radiotherapy; SNP = single nucleotide polymorphism.
Predictive SNPs of XPD
There is evidence that polymorphisms of the gene XPD, which codes for an ATP-dependent helicase that unwinds DNA around lesions, influences the efficiency of DNA repair in lymphocytes and HCT116 (colon carcinoma) cells (90). Furthermore, Zhang et al (88) also reported that Lys/Gln+Gln/Gln genotypes of the XPD Lys751Gln polymorphism were associated with a significantly decreased RIET risk (adjusted HR=0.55; 95% CI, 0.32-0.96; p=0.030) (Tab. II). This finding is significant because the variant 751Gln/Gln genotype was associated with less optimal DNA repair capability when the host cells were exposed to B(a)P, a major constituent of tobacco smoke (91). Further study is pending.
Predictive SNPs of HSPB1
Little is known about whether HSPB1 genotypes are associated with RIET risk among NSCLC patients with RT. Lopez Guerra and colleagues conducted a comprehensive study in 120 NSCLC patients treated with concurrent chemoradiation at the MD Anderson Cancer Center, United States, between 2005 and 2009 (89). They used severe (≥ grade 3) RIET as the end point and found rs2868371 CC genotype was associated with a significantly increased risk of RIET compared with the CG/GG genotype (univariate analysis HR=0.30; 95% CI, 0.10-0.91; p=0.033; multivariate analysis HR=0.29; 95% CI, 0.09-0.97; p=0.045) (Tab. II) (89). This finding corresponds with the relationship between HSPB1 rs2868371 CC genotype and the risk of RP. As has been implied, HSPB1 rs2868371 CC genotype may be valuable for an accurate prediction of radiation-induced toxicities in lung cancer patients. Future multicenter trials including a larger number of cases are needed to study other stress response genes and to validate this promising finding.
Future Directions
Radiation-induced damage is a common problem that may result in considerable morbidity and limited radiation dose during the treatment of lung cancer. Although progress has been made in understanding its underlying mechanisms, more work is still needed to develop safe and effective means for preventing and treating this condition. Recently, increasing numbers of genotype analyses in lung cancer are providing a strong insight into the field of SNP-based prediction for radiation-based toxicities. The well-established expression of DNA repair, inflammation, angiogenesis and stress response pathways and their central roles in the microenvironment of radiation-based damage position them as functionally important factors in radiation-induced damage. The differential expression of SNPs associated with radiation-induced damage have been identified as possible predictive biomarkers. The data reviewed herein show some promising associations and support the idea that polymorphisms of certain genes are useful for the prediction of RP or RIET in lung cancer patients. In addition, there are potential advantages of promoting SNPs as predictive markers for radiation-induced damage. First, SNPs allow for relatively inexpensive and convenient analysis. Second, human diversity and susceptibility of disease regulated by SNPs provides the information that SNPs may be more specific biomarkers for prediction of radiation-induced damage. However, currently none of them can predict occurrence of radiation-induced damage alone and many challenges remain for their translation into practice. For example, first, we have been unable to unravel exact molecular mechanisms through which the above SNPs are involved in RP or RIET. Second, we have used the common candidate SNP method, which does not cover all representative SNPs in the entire gene. Some important but rare SNPs may have been missed, or the observed associations may have been due to genetic linkages with other untyped SNPs. Additional investigations of the tagging SNPs are therefore warranted and will be the subject of future studies. Third, since it is well established that SNPs have a significant ethnic variation, which may account for differences in individual disease susceptibility, validation of genotype associations found in different ethnic populations will be an important consideration in risk evaluation. Fourth, it should be noted that there is a question of whether the above SNPs have similar effects on NSCLC and SCLC patients. Furthermore, one characteristic of the above clinical studies was that investigations were performed on a rather small percentage of the patients included in a large series, which raises questions regarding the representativeness of the findings. Therefore, future efforts should be directed toward better comprehension of gene polymorphisms in the radiation-induced damage microenvironment, along with large prospective studies to confirm the above findings. Additionally, improvements in biotechnology are to be encouraged.
In conclusion, currently, the advantages of such strategies may be limited to patients treated in academic settings. To truly individualize radiation treatment, efforts to develop and implement feasible phenotyping and genotyping strategies along with better comprehension of molecular mechanisms of radiation-related damage are expected to assist clinicians in stratifying lung cancer patients at high risk for radiation-induced toxicities.
