Abstract
Background
Currently, no reliable blood-based assay for early detection of pancreatic ductal adenocarcinoma (PDAC) is available. Cell-free DNA (cfDNA) quantitation in patients’ plasma has been recently applied in monitoring several cancer types. This study evaluates the diagnostic potential of cfDNA in PDAC patients.
Methods
Plasma cfDNA levels and integrity ratio were assayed using quantitative real-time PCR of Alu-repeat amplicons in patients with pancreatic ductal adenocarcinoma (n=50), pancreatic neuroendocrine tumor (n=23), and chronic pancreatitis (n=20), as well as in healthy volunteers without evidence of pancreatic disease (n=23).
Results
The total load of cfDNA, obtained by Alu83 quantitation, was the highest in PDAC patients than in any of the other patient groups (Welch t test; p<0.001) and was an average predictor of PDAC disease (AUC=0.664; CI, 0.56-0.77). A nonlinear association between Alu83 levels and subjects’ age was detected (Spearman's rho=0.35; p<0.001) in the overall population, as well as within the PDAC patients’ group (Spearman's rho=0.47; p<0.001). Necrosis-derived cfDNA fragments, quantitated with the Alu244 amplicon, were barely detectable in any of the samples and, in that respect, comparable between the different subject groups. CfDNA integrity estimation (Alu244/Alu83 ratio) was significantly affected by the limited detectability of plasma Alu244 levels.
Conclusion
The lack of detectable levels of necrosis-derived cfDNA in pancreatic pathologies considerably affects the clinical use of such biomarker in PDAC patients. Different methods of analysis should be applied in the evaluation of the cfDNA diagnostic value in pancreas pathology.
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is the fourth leading cause of cancer-related death in the Western world, and only 20% of patients present localized, surgically resectable disease (1). Pancreatectomy remains the single most effective treatment modality, and offers the only potential for cure. Even if surgical resection is possible, the 5-year survival is only 15%-40% depending on the size of the tumor (2). Symptoms are often vague and easily confused with other gastrointestinal diseases.
Despite much progress concerning the molecular characterization of pancreatic cancer, no reliable blood-based assay for its screening and early detection is clinically available (3). Measurement of the CA19-9 antigen does not represent an efficient biomarker for PDAC due to the high, positive readout in patients with benign diseases as well as the lack of expression in a subset of patients (3). Attempts to use genetic markers, such as KRAS mutations, have been unsuccessful: although present in 70% of the primary tumors in one study, they were not further detected in matched blood samples (4).
As a consequence, at present, clinical screening for asymptomatic individuals at risk relies only on imaging techniques (5). However, this approach has a limited capacity to detect pancreatic cancer at an early or pre-invasive stage, such as that of a pancreatic intra-epithelial neoplasia (PanIN) (5).
Tumor-related cell-free DNA (cfDNA) circulating in blood is a promising candidate biomarker for malignant tumor detection and prognosis (6, 7). In the blood of a healthy individual, low-abundance DNA fragments from physiological apoptotic cell death can be found. Tumor cells release DNA into the bloodstream as a consequence of regulated, as well as non-regulated, cell death related to catastrophic events such as necrosis. Apoptotic DNA fragments are usually less than 200 base pair (bp) in length due to a programmed enzymatic cleavage process (8), while larger fragments of variable sizes are generated during necrosis or mitotic catastrophe (6). The established quantitative PCR-based assay (9, 10) uses two amplicons of cfDNA with different length: (i) short (85 bp), indicative of the total load of circulating DNA and derived from a variety of cell-disrupting processes, and (ii) long (244 bp), attributed to non-apoptotic cell death. Elevated levels of long DNA fragments in the blood have been demonstrated as a reliable marker for the presence of a malignant tumor in the patient (11). The integrity of circulating DNA, measured as the ratio of long cfDNA fragments to the total cfDNA load, was higher in patients with cancers of different origin (7, 9, 10, 12–13–14–15).
In this study, we examined the diagnostic value of both total plasma cfDNA levels and integrity ratio as tools for a non-invasive detection of PDAC in patients. Sensitivity and specificity of the potential biomarker were assayed, using plasma from patients with PDAC, pancreatic neuroendocrine tumor (PanNET), and chronic pancreatitis (CP), as well as from healthy volunteers without evidence of pancreatic disease.
Methods
Patients and plasma samples
A series of consecutive plasma samples obtained from 23 PanNET patients (including 5 insulinomas and 18 non-functioning tumors) and 50 PDAC patients were retrieved from the archives of the ARC-NET biobank of the Verona University Hospital (www.arc-net.it). Patient demographic data are summarized in Table I. All patients had been surgically treated at the same institution (The Pancreas Institute of Verona University and Hospital Trust). Tumor characteristics were obtained from both the gross description of the specimen, as recorded at the time of surgery, and from the original histopathological report. PDAC series also included patients who did not undergo radical surgery. In these cases plasma was prepared before palliative surgery or chemotherapy.
Clinicopathological Characteristics Of The Considered Pdac And Pannet *
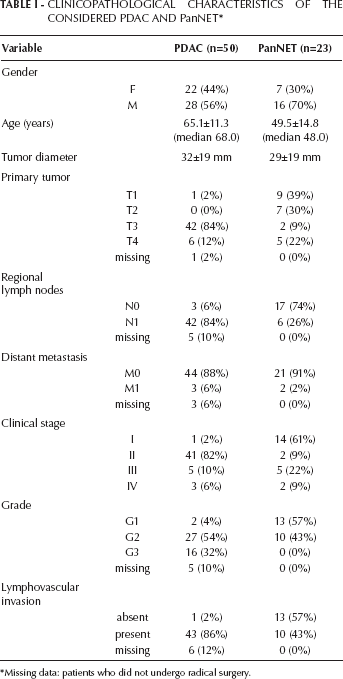
Missing data: patients who did not undergo radical surgery.
To evaluate the specificity of cfDNA as a biomarker for pancreatic tumors versus non-neoplastic disease, plasma preparations from 23 randomly sampled healthy blood donors (of which 13 males; mean age 55.9±8.6 years, median 60 years) and 20 chronic pancreatitis patients (14 males; mean age 52.3±14.2 years, median 51 years) were included in the analysis. Materials used in the experiments have been collected under the Program 853 protocol 298CE 15/02/02 and Program 1885 protocol 52438 on 23/11/2010. The protocols include informed consent of the patient and were approved by the local ethics committee of the Integrated University Hospital Trust of Verona.
Plasma preparation and DNA extraction
All peripheral fasting blood samples were collected in 10-mL sodium heparin tubes (BD Vacutainer; Becton-Dickinson, Milan, Italy) and processed within 2 hours by centrifugation at 2,000g at 4°C for 10 minutes. The plasma obtained was then transferred to new tubes and centrifuged at 3,000g at 4°C for 10 minutes to remove platelet contamination. Final plasma preparations were carefully collected from the upper portion of the supernatant and stored in aliquots at -80°C.
Total DNA was extracted from 500 µL of plasma using the QIAmp UltraSens Virus Kit (Qiagen, Hilden, Germany) following the manufacturer's instructions. Final elution was performed with 50 µL of elution buffer and used directly for Alu sequences quantitation.
PCR quantitation of Alu sequences
Real-time PCR reactions were assembled with Euroclone Alu Reagents Kit (Euroclone, Milan, Italy). Two amplicons were quantified: Alu83 (83 nucleotides), representative of the total load of circulating DNA, and Alu244 (244 nucleotides), representative of the DNA released in necrotic processes.
Alu83 reactions were assembled with 1x Fluocycle II Master Mix (no ROX), 0.05 µM forward primer, 0.009 µM reverse primer, and 0.25 µM TMR probe in a final volume of 9 µL. Alu244 reactions were assembled as above, but with both primers equimolar at 0.05 µM. In both cases, 4 µL of circulating DNA eluate were used as input. The reactions were performed on a 7900HT Fast Real-Time PCR System (Applied Biosystems; Life Technologies, Monza, Italy) with the following temperatures’ profile: 5 minutes initial denaturation, 40 cycles of 15 seconds denaturation at 95°C, followed by 1 minute annealing/extension at 65°C. Genomic DNA from normal spleen was used to construct the titration curve. Amplification curves were processed using the Sequence Detection Systems software v2.3 (Applied Biosystems). Mean values of at least 5 ng/mL were considered detectable and samples with a standard deviation ≤0.5 from the mean value were kept for further analyses.
Statistical analyses
All statistical analyses were performed using the R 3.0 programming language (CRAN). Lognormal distributions were confirmed for patients’ plasma Alu83 and Alu244 values by qqplot inspection. Subsequent testing was performed on log-transformed values. Variance equality between groups was probed with Brown-Forsyth test (package “HH”).
Either the ANOVA (in case of equal variances) or one-way Welch test (in case of unequal variances between the groups compared) was applied to detect associations between Alu measurements and categorical clinical variables. Associations between continuous clinical variables (subject age, tumor size, and tumor stage) and Alu levels were analyzed by linear models and Spearman's correlation. Multivariate analyses were performed for factors showing significant correlations (p<0.05) in univariate models.
Between-group differences were tested using the Student's t-test (equal variances) or Welch t-test (unequal variances). In case of small non-significant differences in within-group variances (visible on qqplots) and by unequal group sizes, the Welch-modified tests were preferentially performed to maintain stringency. P values are reported accordingly as detailed out in the text.
Performance of predictors was assessed by AUC inspection (package “pROC”) and logistic regression analysis where applicable.
Results
A total of 116 subjects was included in this study. Plasma samples obtained from 73 patients with pancreatic carcinoma were compared to 20 CP patients and 23 randomly sampled healthy blood donors. By using qRT-PCR, we quantified the presence in the plasma of 2 specific amplicons: Alu83, representative of the total load of circulating DNA, and Alu244, representative of the DNA released in necrotic processes.
The total load of cfDNA is increased in PDAC patients
Alu83 quantitation was used to inspect the total load of cfDNA in the subjects’ plasma. The linear amplification range for the Alu83 amplicon calculated on basis of the reference spleen genomic DNA was 5 ng/mL-200 µg/mL. Alu83 values did not differ significantly between males and females. A nonlinear association between Alu83 and subjects’ age was detected (Spearman's rho=0.35; p<0.001) in the overall population. Such association was also evident within the PDAC patients’ group (Spearman's rho=0.47; p<0.001), but not in the other groups.
The total load of cfDNA was inspected across the groups (Fig. 1A, showing the original non-transformed values). Large differences in within-group variances were not observed for Alu83 values (Brown–Forsyth test, p=0.9737). The total cfDNA was associated with disease type (ANOVA, p<0.01). The difference in Alu83 levels was significant between PDAC patients and healthy controls (Welch t-test, p<0.05), as well as between PDAC and CP patients (Welch t-test, p<0.01). The mean Alu83 values were significantly higher in PDAC patients than in all other subjects pooled (Welch t-test, p<0.001), and were an average predictor of PDAC disease (AUC of 0.664; CI, 0.56-0.77; Fig. 1B). Age was a confounding factor for this association. No correlation between Alu83 and tumor size was observed, neither in PDAC nor in PanNET patients (data not shown).
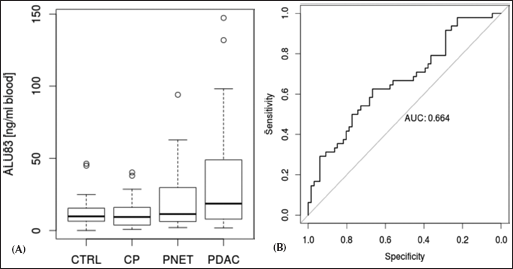
The total cfDNA load, determined by Alu83 quantitation, is increased in PDAC patients. Alu83 values were significantly higher in PDAC patients than in patients of the other groups (A) and were average predictors of PDAC disease (B).
Necrosis-derived cfDNA is barely detectable in patients with pancreatic diseases
In most plasma samples of our cohort the Alu244 levels, indicative of necrotic DNA fragmentation, were below or at the beginning of the qPCR linear amplification range established with the reference spleen genomic DNA (5 ng/mL-200 µg/mL). As expected for measurements close to the detection threshold, the inter-replicate standard deviation was often more than 0.5 times the mean value; this forced us to remove these samples from the analysis. Consequently, the Alu244/Alu83 ratios were obtained only for a few samples, and their high variability made them unsuitable for statistical predictions.
As in the case of total Alu cfDNA load (Alu83), necrotic DNA fragments (Alu244) were not related to gender, although they were fairly correlated to subjects’ age (Spearman's rho=0.25, p<0.05).
We did not find any association of the Alu244 levels with PDAC or PanNET tumor size. In contrast to Alu83, the low levels of necrotic Alu fragments were comparable between patient groups (Fig. 2). Subjects with the highest measured Alu244 values (>5 ng/mL) included the oldest patients (>65 years).
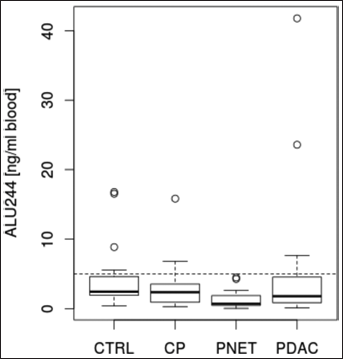
Necrosis-derived cfDNA levels, determined by Alu244 quantitation, are barely detectable in patients with pancreatic diseases and are comparable among the different patient groups. Horizontal line: detection threshold at 5 ng/mL.
Discussion
Multiple hypotheses concerning the origin of circulating DNA have been proposed, including cell necrosis, apoptosis, mitotic catastrophe, autophagy, and circulation of micrometastases (16).
Most of the previous studies focused on cfDNA derived from cell apoptosis and/or necrosis (17, 18), as these may be distinguished by the length of the circulating DNA fragments detected. In fact, in case of apoptotic death, cells release DNA fragments mostly of 186 bp in length, while tumor necrosis is characterized by the presence of fragments that vary in size and are generally greater than 200 bp. As a result, in cancer patients with prominent necrosis, DNA fragments of >200 bp are readily detectable, resulting in a DNA integrity (ratio between long fragments and total DNA) increasing towards 1; by contrast, in healthy individuals only shorter fragments are present, indicating the event of apoptosis, and DNA integrity is close to 0 (11).
In this study, we sought to test the potential capacity of cfDNA circulating in plasma to discriminate pancreatic cancer patients from subjects with non-malignant pancreatic disease and healthy controls. We used a robust method of analysis that has previously been successfully applied in the early detection of solid tumors by others (7, 9, 10, 12–13–14–15, 19, 20).
In contrast to what has been observed in colon and breast cancers, the load of circulating tumor DNA in our pancreatic tumor patients seems to derive from non-necrotic cell death. In fact, necrosis-related Alu244 sequences were feebly detectable in most of our samples, and the overall levels were comparable among the different patient groups. Of interest, the few plasma samples that contained Alu244 above the ~5 ng/mL detection threshold were from patients that were >65 years old, had a clinical history of necrotic acute pancreatitis or had PDAC residual/inoperable disease (i.e. patients who did not undergo radical surgery; Tab. I). This could be partially explained by the limited presence of necrotic areas in PDAC and PanNET compared with colorectal tumors.
The limited levels of Alu244 hampered the evaluation of cfDNA integrity, which could therefore not be used for prediction of PDAC disease at an early stage.
On the other hand, the total load of cfDNA (i.e. Alu83 levels) was significantly higher in PDAC subjects than in the subjects of the other groups, suggesting an increased rate of apoptosis in this malignancy. Similarly, Giacona and colleagues observed that most of the cfDNA in PDAC patients is associated to apoptosis of neoplastic cells (8).
No association was found for Alu83 or Alu244 and PDAC or PanNET tumor size or other pathological variables. Major limitations of the present study are related to the relatively small number of patients evaluated, the clinical heterogeneity of the considered neoplasm, and the skewed disease stage distribution in our cohort. For these reasons, the current work should be considered as a pilot study, and the association between Alu values and clinicopathological tumor features should be validated on a larger series of patients.
Conclusions
A significant load of cfDNA in plasma characterizes PDAC patients. However, necrosis-derived cfDNA fragments, quantified with the Alu244 amplicon, were barely detectable in any of the considered samples and, in that respect, were comparable between the different subject groups. Whether other methods of cfDNA quantitation could be used as early biomarkers of PDAC detection will require further investigation.
