Abstract
Background
Inflammatory bowel disease (IBD) results from an inappropriate inflammatory response in which genetic, immune, and environmental factors all play important roles. Recently, single nucleotide polymorphisms (SNPs) in the CD24 gene have been associated with the development of several autoimmune diseases.
Aim
To evaluate whether CD24 SNPs, are associated with risk of ulcerative colitis (UC) and Crohn's disease (CD).
Methods
The CD24 polymorphisms C170T (rs8734), TG1527del (rs3838646), A1626G (rs1058881), and A1056G (rs1058818) were assessed in a case-control study of an Israeli cohort comprising 117 IBD patients and 105 age and gender-matched healthy controls. Restriction fragment length polymorphism (RFLP) analysis was performed using BstX1, Bsr1, Mfe1, and BstU1 restriction enzymes. Odds ratios (OR) and 95% confidence intervals (CI) were estimated by logistic regression models.
Results
Carriers of the C170T SNP were at increased risk of IBD (OR=3.022, 95% CI: 1.748–5.223, p=0.001), UC (OR=3.002, 95% CI: 1.661–5.427, p=0.001) and CD (OR=3.077, 95% CI: 1.334–7.095, p=0.008). Carrying the A1626G and A1056G SNPs was found to be a risk factor for IBD (OR=2.460, 95% CI: 1.420–4.259, p=0.001 and OR=1.856, 95% CI: 1.011–3.405, p=0.01), UC (OR=2.218, 95% CI: 1.207–4.075, p=0.01 and OR=1.944, 95% CI: 0.995–3.798, p=0.01) but not for CD (p=0.086 and p=0.299). The A1626G and TG1527del were found to be associated with younger age of IBD onset (p=0.022 and p=0.027, respectively).
Conclusions
The CD24 C170T polymorphism is associated with IBD risk. The A1626G and A1056G SNPs might be associated only with UC risk. These findings suggest CD24 as a new genetic susceptibility factor, with clinical implications in the prediction of IBD prognosis and therapy.
Keywords
Introduction
Described more than a century ago, inflammatory bowel disease (IBD) is an inflammatory disorder of the gastrointestinal mucosa, and is composed of two types of disease: Crohn's disease (CD) and ulcerative colitis (UC). The presenting symptoms are diarrhea, abdominal pain, vomiting, fever, weight loss, and extra-intestinal manifestations such as arthritis, uveitis, and skin rashes (1). The age of onset of UC is usually higher that that of CD. UC affects only the colon and the rectum, while CD can affect the entire gastrointestinal (GI) tract (1). The etiology of IBD is uncertain. Numerous studies suggest that it may result from a combination of certain genetic variations, changes in the immune system, as well as environmental and infectious factors. Recent studies have identified variations in specific genes, including ATG16L1, IL23R, IRGM, and NOD2, that influence the risk of developing IBD (2, 3). The application of cutting-edge genomics technologies and genome-wide association (GWA) scanning has recently identified several IBD susceptibility genes, such as NOD2 and IBD5, which account for 30% of the genetic contribution to CD (4). Additional genes and genetic variants remain to be identified.
CD24 is a glycophosphatidylinositol (GPI)-anchored mucin-like cell surface glycoprotein (5), encoded by a gene located on chromosome 6. The CD24 gene has several pseudogenes on 3 different locations in the human genome (chromosomes 1, 15, and Y) (6). CD24 is expressed in a variety of cell types, including activated T cells (7, 8), B cells (9), macrophages (10), and dendritic cells (11). CD24 was also shown to be expressed in a variety of solid tumors, including colorectal cancer (CRC) (5). The CD24 gene contains 4 single nucleotide polymorphisms (SNPs); among them, the 170C→T and 1527TG SNPs were shown to influence the risk of several chronic inflammatory and autoimmune diseases, such as chronic hepatitis B virus (HBV) infection (12), experimental autoimmune encephalomyelitis (EAE) (13), multiple sclerosis (MS) (14), and systemic lupus erythomatosus (SLE) (15). The other SNPs confer protection to MS and SLE (16). Mohamed et al (13) recently demonstrated that CD24 expression is upregulated in regenerating tissue in IBD, thus supporting its role in tissue healing in the colon.
Three (1056A→G, 1527TG/del, and 1626A→G) of the 4 CD24 gene SNPs are located on the 3′-untranslated region (UTR), while the fourth SNP, a C>T transition (rs52812045) at position 170, a putative GPI-anchor cleavage site (-1 position), results in a non-conservative substitution of alanine (A) with valine (V) (6). The association of these SNPs with chronic autoimmune diseases and a recent publication regarding the association of 1527TG/del with CD in a Spanish population (14) prompted us to explore all the 4 known SNPs in patients with UC and CD.
Materials and Methods
Study population
The whole sample collection and experimental procedure were approved by the institutional review board of the Tel-Aviv Sourasky Medical Center and the Israeli Ministry of Health. Informed consent from all participants was obtained prior to sample collection and entry into the study.
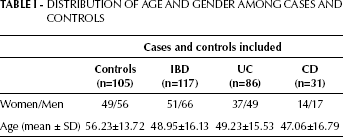
The study population comprised 117 IBD patients (86 UC and 31 CD) and 105 age and gender-matched healthy controls that were prospectively enrolled through the Integrated Cancer Prevention Center at Tel-Aviv Sourasky Medical Center from October 2000 to May 2010. IBD diagnosis was assessed according to established clinical guidelines and endoscopic, radiologic, and histopathologic criteria. Demographic and routine clinical data, including behavior and location of IBD and disease-related complications, were recorded by retrospective analysis of the patients’ clinical charts. The baseline characteristics of the study population are shown in Table I.
Distribution Of Age And Gender Among Cases And Controls
DNA extraction and genotyping
DNA was extracted from peripheral blood leukocytes using standard procedures as described by Miller et al (17). DNA was amplified by a standard polymerase chain reaction (PCR) using PCR Master Mix Taq polymerase (Thermo Scientific, www.thermoscientific.com/onebio). The CD24 170C→T, 1056A→G, 1527TG/del, and 1626A→G variants were identified using restriction fragment length polymorphism (RFLP) analysis as described below.
The subjects were examined for the 4 SNPs in the CD24 gene using restriction RFLP analysis: the C→T substitution at position 170 (rs8734), at the open reading frame, results in an A to V substitution; the other 3 SNPs, located in the 3′ UTR, are the A→G substitution at position 1056 (rs1058818), the TG dinucleotide deletion at position 1527 (rs3838646), and the A→G substitution at position 1626 (rs1058881).
Determination of the 170C→T polymorphism in the CD24 gene
DNA fragments bearing the SNP site were amplified by a PCR using a forward primer (5′ - TTG CCA CTT GGC ATT TTT GAG GC - 3′) and a reverse primer (5′ - GGA TTG GGT TTA GAA GAT GGG GAA A - 3′).
The PCR was carried out in a total reaction volume of 25 μL using the following amplification protocol: denaturation at 95°C for 5 minutes, followed by 30 cycles of denaturation at 90°C for 10 seconds, and annealing and extension at 55°C for 30 seconds. The Taq polymerase was used for the amplification. After PCR, the genotype of each sample was determined by BstXI restriction digestion. The T→C change creates a BstXI restriction site at nucleotide 170, allowing for the differentiation between these 2 CD24 alleles by RFLP analysis. A total of 10 μL of CD24 PCR products were digested with BstXI for 16 hours at 55°C. The digested products were then separated in a 2% agarose gel. The predicted digestion pattern was the following: PCR products of the T170 allele were cut into 2 small fragments (325 bp and 129 bp), whereas those of the C170 were not cut, resulting in a fragment of 454 bp.
Determination of the polymorphisms at the 3′UTR of the CD24 gene
Nested PCR was used to extract only the active gene sequence and to distinguish it from the other products. The first PCR forward primer sequence included both the intron and the second exon, allowing obtaining only the coding gene fragment and not the pseudogenes.
Determination of the 1056A→G, 1527TG/del, and 1626A→G polymorphisms in the CD24 gene by nested PCR
DNA fragments bearing the other 3 SNP sites were amplified by PCR using a forward primer (5′-CTA AAG AGA ATG ACC TTG GTG GGT TGA-3′) and a reverse primer (5′-CAC AGT AGC TTC AAA ACT GTT CGA-3′). The PCR was carried out in a total reaction volume of 25 μL using the following amplification protocol: denaturation at 95°C for 5 minutes followed by 94° for 30 seconds, 55°C for 30 seconds, and 68°C for 2 minutes for 20 cycles. The length of the expected PCR product was 2,017 bp.
DNA fragments bearing the SNP sites from the first PCR were amplified by PCR using the following primers: frw 5′-GGC ATT TCC TAT CAC CTG TTT-3′, and rev 5′-AAT CTA CCC CCA GAT CCA AGC A-3′ for 1056A→G; frw 5′-GCA ATT TTG CCT TCA AAA CAG-3′, and rev 5′-TTT AGG CTT AGG ACC AGG TTC-3′ for 1527TG/del; frw 5′-CAA CTA TGG ATC AGA ATA GCA ACA AT-3′, and rev 5′-GGAACATCTAAGCATCAGTGTGTG-3′ for the 1626A→G polymorphism. The PCR conditions were as follows: 94°C for 30 seconds, 55°C for 30 seconds, and 72°C for 30 seconds, for 35 cycles.
Following PCR, the genotype of each sample was digested by BstUI (at 60°C) for P1056, BsrI (at 65°C) for P1527, and MfeI (at 37°C) for P1626 (New England Biolabs, http://www.neb.com) for 16 hours; then, 10 μL of the digestion product were electrophoresed on 2.0% agarose gel (16). The expected fragments’ length of the second PCR were: for 1056A→G the A/A fragment is of 550 bp, and the G1056 allele is cut into 2 smaller fragments of 400 bp and 150 bp; for 1527TG/del the TG allele is cut into 318-bp and 158-bp fragments, whereas the del is the uncut fragment of 474 bp; for 1626A→G the A1626 allele is of 209 bp, and the G1626 allele is cut into 2 fragments of 181 bp and 28 bp. For small fragments, as in the case of the 1626A→G variant, a 4% agarose gel was used.
Statistical analysis
The association between each polymorphism and UC or CD, alone and in combination, was evaluated by logistic regression and presented as odds ratio (OR) with confidence intervals (CI); the calculated probability (p) values are two-sided (p=0.05) at the 95% level of significance.
The association between the risk factors (polymorphisms, age, physical activity, education, gender, dietary supplementation, ethnicity) and outcomes (findings of UC or CD) was examined by the Student's t-test and presented as p values, all two-sided and reported at the 95% level of significance (p=0.05). Database organization and management was carried out in Microsoft Excel. All statistical analyses were carried out using SPSS software, version 14 (Chicago, IL).
Results
A total of 222 consecutive subjects were included in this study. The IBD patients included were 117: 86 with UC and 31 with CD. Each subject underwent regular screening tests and additional evaluation according to his personal risk profile. The demographic characteristics of the subjects included are presented in Table I. The mean age of the study population was 52.36 years (range 32–83 years) and 54.05% of them were males.
The Ala57Val and A1626G variants and the risk for IBD
The overall prevalence of the genetic polymorphisms is showed in Table II and Figure 1, and a representative example of all 4 polymorphisms is shown in Figure 2. The CD24 170C→T SNP was present in 117 (52.7%) cases, 75 (63%) of which were IBD cases (68% UC and 62% CD) and 42 (40.5%) were controls. The heterozygote genotype was present in 35.5% of the controls and 55.5% of the IBD patients (57% UC and 59% CD), while the val/val genotype was present in 5% of the controls and 7.5% of the IBD patients (9% UC and 3% CD). The CD24 1056A→G SNP was present in 166 cases (74.8%), 92 (78%) of which were IBD (79% UC and 75% CD) and 74 (71%) were controls (p>0.05). The CD24 1527TG/del SNP was present in 25 cases (11.3%), 14 (12%) of which were IBD cases (10.5% UC and 15% CD) and 11 (11%) were controls (p>0.05). The CD24 1626A→G SNP was present in 95 cases (42.8%), 62 (52%) of which were IBD cases (54% UC and 45% CD) and 33 (32%) were controls (p<0.05). The distributions of the homozygous and heterozygous cases are shown in Table II.
Distribution Of The Different Genotypes For The Polymorphisms
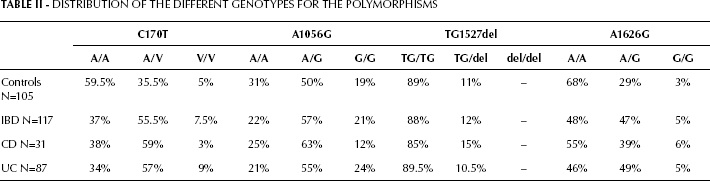
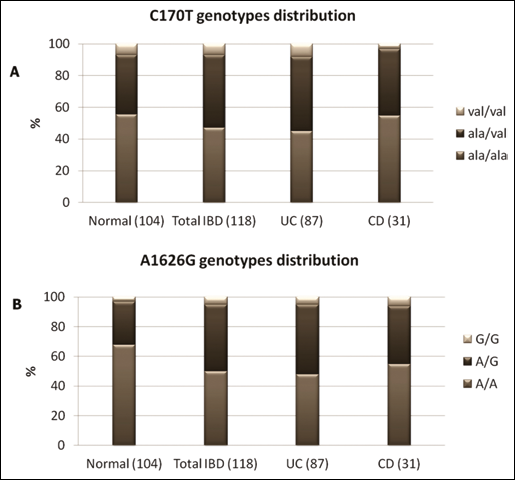
C170T (A) and A1626G (B) genotype distribution. Distribution of CD24 C170T and A1626G genotypes among IBD (Total), UC, and CD patients compared to the group of healthy controls (Normal). Genotyping was performed by PCR followed by restriction enzyme digestion using BstXI and MfeI. The data are based on the analysis of 104 normal controls and 117 IBD patients.
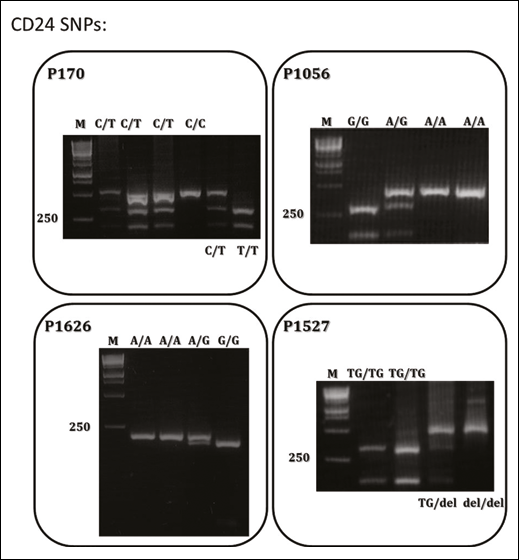
Example of genotyping by PCR-RFLP analysis using BstXI, BstUI, BsrI, and MfeI restriction enzymes for, respectively, the polymorphisms P170, P1056, P1527, and P1626. The genotypes of the individuals are indicated in the lanes. M: molecular size marker. The number on the left side (250) is the size of a standard DNA marker (bp).
The means of 2 or more groups were compared using the analysis of variance (ANOVA) between groups. By comparing carriers versus non-carriers in IBD patients versus healthy controls we obtained: CD24 170C→T (p=0.01), 1056A→G (p=0.224), 1527TG/del (p=0.730), and 1626A→G (p=0.007). This indicated that 1056A→G and 1527TG/del were not significantly different.
Using univariate logistic regression, we found that carriers of the CD24 170C→T, 1626A→G, and 1056A→G SNPs were more common among IBD patients (respectively OR=3.022, 95% CI 1.748–5.233, p=0.001; OR=2.460, 95% CI 1.420–4.259, p=0.001; and OR=1.856, 95% CI 1.011–3.405, p=0.046). These findings prompted us to perform the same comparisons between the UC and CD patients versus healthy controls, separately, in spite of the small size of the groups. A statistically significant difference was observed among almost all the different groups. An increased risk was observed in UC patients carrying the 170C→T (OR=3.002, 95% CI 1.661–5.427, p=0.001), the 1626A→G (OR=2.629, 95% CI 1.455–4.751, p=0.01), and the 1056A→G SNPs (OR=1.944, 95% CI 0.995–3.798, p=0.052). Similarly, an increased risk was observed also, among CD patients (OR=3.077, 95% CI 1.334–7.095, p=0.008 and OR=2.045, 95% CI 0.904–4.626, p=0.086, respectively). The 1527TG/del SNP showed no significant association with IBD (p=0.73, p=0.224, respectively for UC and CD patients).
The next step of the study was to find whether there is an association between the 3 significant CD24 SNPs, namely 170C→T, 1056A→G, and 1626A→G. Therefore, we combined the carriers of 2 of each polymorphism and examined whether there is a higher risk for IBD in these individuals. Carriers of the 170C→T and 1626A→G SNPs were at significantly increased risk of harboring IBD (OR=6.875, 95% CI 3.275–14.433, p=0.001). An increased risk was also observed for UC subjects as compared to healthy controls (OR=5.000, 95% CI 2.340–10.682, p=0.001), but not for CD patients (OR=1.875, 95% CI 0.795–4.422, p=0.151). The other two combinations showed non-significant results.
Several life-style and disease-related parameters were investigated, such as smoking, age at diagnosis, IBD-related surgeries, median follow-up time, and location of the disease. For this purpose, a multivariate logistic regression analysis was performed. Among all the parameters that were examined, only the age of onset was associated with the CD24 gene SNPs. The 1527TG/del polymorphism exhibited a protective effect since the age of diagnosis was older among patients than among healthy controls (p=0.027). The 1626A→G SNP increased the IBD risk since the age of disease onset was significantly younger (p=0.022). The CD24 170C→T and 1056A→G SNPs had no influence on IBD risk (p=0.506 and p=0.431, respectively).
Discussion
The results of the current study demonstrate that CD24, a genetic risk factor previously implicated in malignant, autoimmune, and other chronic inflammatory disorders, is also a risk factor for IBD susceptibility. The frequency of 3 out of the 4 CD24 genetic SNPs was significantly different between the patients’ groups and the controls’ group. Among the 4 SNPs, 2 (170C→T and 1626A→G) were found to be significantly associated with an increased risk for both UC and CD. Specifically, for 170C→T we found an OR=3.022 for IBD, OR=3.022 for UC, and OR=3.077 CD, and for the 1626A→G SNP we found an OR=2.460 for IBD, OR=2.629 for UC, and OR=2.045 for CD. By contrast, the presence of the 1056A→G SNP increased only the risk of total IBD and UC (OR=1.856 for IBD, and OR=1.944 for UC). This could be due, at least in part, to the relatively low number of CD patients in our study. In the past 2 decades, multiple genes that predispose to IBD susceptibility have been identified and confirmed. The great advance in understanding the genetic background of IBD is related to recent case-control and genome-wide association studies (GWAS) (5), which have identified 85 distinct loci associated with the disease. However, the genes identified thus far only explain around a quarter of the genetic contribution to CD (16).
CD24 polymorphisms have primarily been studied in the context of malignancy (11). Recent studies investigating their role in autoimmune disorders have found an association with several diseases, such as MS and SLE (13–15). Two recent reports provide insights into the role of CD24 in the genetic background of IBD. Ahmed et al (18) demonstrated that CD24 is upregulated in regenerating epithelium in IBD and plays a role in mucosal healing. The association of the CD24 gene and IBD has been supported by a recent study of Diaz-Gallo et al. (19), involving 369 CD and 323 UC Spanish patients, which demonstrated that a TG deletion in the 3′-UTR (rs3838646) was associated with an increased risk of CD (OR=1.61, 95% CI=1.17–2.21), but not of UC. In addition, the authors found that this allele was significantly associated with early onset of disease, ileocolonic location, and an inflammatory type of CD, and not with perforating or structuring disease. This association is supported by a recent meta-analysis of 6 CD GWAS that showed the presence of a signal of association in the CD24 region (6q21, GeneID 100133941) (3). The findings of the current study confirm those of Diaz-Gallo et al. (19) concerning the association between the TG1527del SNP and CD, showing that the deletion allele is associated with an increased risk of CD, but not UC, in a Spanish population. However, the association of the 2 other SNPs in the 3′-UTR that was observed in our study is novel and has not been described in IBD patients before. Despite the low number of subjects, a statistically significant association was found between these SNPs and the IBD risk in an Israeli population. We also found that the A1626G and TG1527del SNPs were associated with a younger age of disease onset (respectively p=0.022 and p=0.027). These findings further indicate that not only the occurrence of the disease, but also its different sub-phenotypes, such as age of onset and disease behavior, might be related to CD24, of which biological functions might be involved in generating different IBD phenotypes.
The association of CD24 with MS and SLE, and now with IBD, is in line with the concept of shared genetic determinants for clinically distinct disorders (14, 15). It seems that there may be a general set of susceptibility genes for autoimmunity, which are modulated by disease-specific genes, as well as by the host's human leukocyte antigen status. A specific combination of polymorphisms, combined with environmental factors, could determine the type of disease developed by a subject.
A weakness of this study derives from the relatively low number of subjects involved, in particular those with CD. A main limitation, thus, relies on the lack of sufficient statistical power and low resolution when compared with the current gold standard of GWA scans (3). However, the good quality of the clinical data, the long follow-up, and the high number of age and gender-matched controls suggest that these observations may be true and significant.
Clearly, much remains to be investigated about the influence and interactions between specific genetic polymorphisms and IBD. Identifying and understanding these interactions can improve differential diagnosis and optimize treatment efficacy.
