Abstract
Background
The prognostic significance of the expression of the miR-221/222 family in cancer remains controversial. We here performed a meta-analysis of published data investigating the effects of miR-221/222 expression on both overall survival (OS) and disease-free survival (DFS) among patients with cancer.
Methods
A systematic search of the PubMed, Embase, Cochrane, and CNKI databases was performed with the last search being updated on March 15, 2013. The hazard ratio (HR) and its 95% confidence interval (95% CI) were used to assess the strength of association.
Results
A total of 17 studies involving 1,204 subjects were included in this meta-analysis. When assessing the prognostic significance of miR-221 expression, the pooled HR was 1.91 (95% CI: 1.28-2.85, p=0.002) for OS and 1.36 (95% CI: 0.88-2.09, p=0.163) for DFS. When assessing the prognostic significance of miR-222 expression, the pooled HR was 2.15 (95% CI: 1.51-3.06, p<0.0001) for OS and 1.37 (95% CI: 0.45-4.13, p=0.581) for DFS. We also found that an elevated miR-221 expression was significantly associated with poor OS when stratifying by ethnicity, cancer type, statistical methodology, sample, and quality assessment. There was no evidence of publication bias.
Conclusion
The meta-analysis demonstrates that the elevated expression of miR-221 and miR-222 is associated with poor OS in patients with cancer. The miR-221/222 cluster might be used as a potential therapeutic strategy in clinical practice. More work is required to fully elucidate the role of the miR-221/222 family in human tumors.
Introduction
In the era of personalized medicine, insights gained from novel molecular tests may improve the ability to make accurate prognostic assessments for patients with cancer (1). The expression of microRNAs (miRNAs) was identified as one of the most promising new biological measurements. MiRNAs represent a class of highly conserved, noncoding RNA molecules of 18-25 nucleotides that regulate gene expression through posttranscriptional interactions with mRNA (2). MiRNAs can be either over or under-expressed, functioning as oncogenes or tumor suppressors, depending on their downstream target genes (3). In human malignancies, miRNA expression profiles can classify different cancer types (4), identify their subtypes (5–6), and distinguish the tissue of origin of the tumors (7). MiRNAs have also shown great promising prognostic associations with major cancer outcomes, including response to therapy, recurrence, and overall survival (OS) (8).
Among the most intensively studied miRNAs, miR-221 and miR-222 are 2 highly homologous miRNAs. They are encoded in tandem on the X chromosome in human, mouse, and rat, and are highly conserved in vertebrates. In addition, they share a seed sequence (9). To date, many studies have reported on the role of miR-221/222 in cancer development, either as oncomiR or as oncosuppressor-miR. Overexpression of miR-221/222 has been observed in various cancer types, including prostate cancer, thyroid papillary carcinoma, breast cancer, hepatocellular carcinoma, and lung cancer (10–14). However, miRNA function depends on the cellular contest and tumor type. A few studies have shown a tumor suppressive role for miR-221/222 in tongue squamous cell carcinoma (OTSCC) and erythrocytes (15–16). Recently, some prognostic studies found that a higher miR-221/222 expression was significantly associated with worse outcome. Nevertheless, some other studies showed insignificant or opposite results. Currently, a formal systematic synthesis of the evidence on the prognostic significance of miR-221/222 for cancer outcomes is lacking.
Every single study may be underpowered to achieve a comprehensive and reliable conclusion. Thus, we carried out a meta-analysis to obtain a more precise estimation of the association between miR-221/222 expression and the prognosis of patients with cancer.
Materials and Methods
Search strategy
We used the PRISMA checklist as protocol of the meta-analysis and followed the appropriate guidelines (Tab. S1) (17). A comprehensive literature search was done for original articles analyzing the prognostic value of miR-221/222 in cancer with PubMed, Excerpta Medica Database (Embase), the Cochrane Central Register of Controlled Trials, and the Chinese National Knowledge Infrastructure (CNKI) database. Studies were selected by using the following search terms variably combined: ‘miR-221’, ‘microRNA-221’, ‘microrna-221’, ‘miRNA-221’, ‘miR-222’, ‘microRNA-222’, ‘microrna-222’, ‘miRNA-222’, ‘cancer’, ‘carcinoma’, ‘tumor’, ‘neoplasm’, ‘prognosis’, ‘survival’, ‘recurrence’, and ‘relapse’. The last search was performed on March 15, 2013. The search was conducted without any restriction on language or publication years. Simultaneously, the reference lists of review papers and original reports were hand-searched for further relevant studies.
Eligibility criteria
The published studies that were eligible for inclusion in this meta-analysis met the following criteria: (i) evaluated the association between miR-221/222 expression and cancer prognosis; (ii) provided hazard ratios (HRs) with their corresponding 95% confidence intervals (CIs) or sufficient data for estimating HR with 95% CI. If the study did not meet the criteria, it was excluded from our research.
Data extraction
Data extraction was performed independently by 2 reviewers. Disagreement was resolved by discussion with our research team. The extracted data elements included the following: first author's name, publication year, ethnicity, number of patients, sample, miR-221/222 assessments, cutoff values, follow-up time, and prognostic information. If the required information were unavailable in relevant articles, a request was sent to the corresponding author for additional data. If a study reported the results on different ethnicities, we treated the data as separate studies.
Quality assessment
We systematically assessed the quality of all the studies included according to the Newcastle-Ottawa quality assessment scale (NOS) for cohort studies (Tab. S2) (18). Each study was judged on 3 broad perspectives: selection, comparability, and outcome. Quality scores ranged from 0 to 9 and a high-quality study was defined as one with a score of ≥6.
Statistical analysis
Outcomes of interest were OS and disease-free survival (DFS). The HR and its 95% CI were used to assess the strength of association. A test of heterogeneity of combined HRs was conducted using Cochran's Q test and Higgins I2 statistic (19). I2 was measured in the interval 0%-100% with increasing I2 values indicating a larger impact of between-study heterogeneity in the meta-analysis. The random-effects model (DerSimonian and Laird method) was used if heterogeneity was observed (p<0.10), while the fixed-effects model (the inverse variance method) was applied in the absence of between-study heterogeneity (p≥0.10) (20–21).
The most accurate method included retrieving the HR estimate and its 95% CI from the original article or from the authors. If not available, the total number of observed deaths/cancer recurrences and the number of subjects in each group were extracted to determine the HR estimate (22). If only Kaplan-Meier curves were available, data were extracted from the graphical survival plots and estimation of the HR was performed using the method described by Parmar and colleagues (23). All data analyses were carried out using the low miR-221/222 expression group as the reference group (HR=1). An observed HR >1 implied a worse prognosis for patients with high miR-221/222 expression.
Publication bias was evaluated using the funnel plot with the Egger's bias indicator test (24). A p<0.05 for the Egger's test was considered to be representative of statistically significant publication bias. All analyses were performed on Stata version 10 (StataCorp LP, College Station, Texas, USA). Kaplan-Meier curves were read by Engauge Digitizer version 4.1 (http://digitizer.sourceforge.net). Statistical tests were two-sided and p values <0.05 were considered significant.
Results
Literature search
Four-hundred and sixteen records for miR-221/222 and cancer were identified from a systematic literature search in PubMed, Embase, Cochrane, and CNKI. Three-hundred and thirty-three records were first screened after duplicates had been removed. After manually screening the records, 310 studies were excluded because they were review articles, conference abstracts, non-human studies, or studies irrelevant to the current analysis. Of the 23 candidate studies, 4 lacked the key HR data (25–28), 1 study investigated the miR-221/222 ratio but not the miR-221 or miR-222 expressions alone (29), and 1 study only investigated the progression-free survival (30). Therefore, a total of 17 studies were eligible for the final meta-analysis (31–47). A flow diagram of the study selection process is presented in Figure 1.
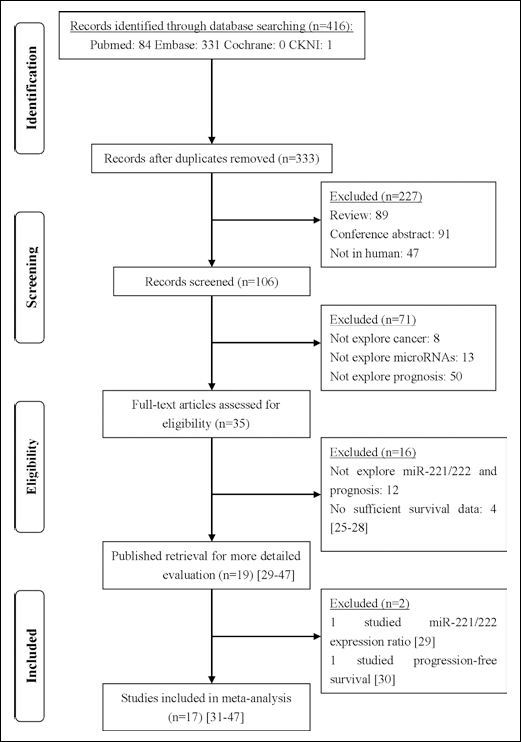
Flow diagram of the studies' selection process.
Study characteristics and quality assessment
The main characteristics of the included studies are shown in Table I. There were 12 studies evaluating the prognostic significance of miR-221 (31, 35–43, 46, 47), 3 studies evaluating the prognostic significance of miR-222 (32, 34, 45), and 2 studies evaluating both miRNAs (33, 44). The main outcomes of interest were OS (10 studies) (32, 35, 37–40, 42–45) and DFS (5 studies) (33, 36, 41, 46, 47), while two studies examined both (31, 34). Among these studies, 11 were performed in Asians (34, 35, 37–41, 43–46), and 6 were performed in Caucasians (31–33, 36, 42, 47). Sample sizes ranged from 32 (35) to 115 patients (41), for a total of 1,204 patients. The patients were affected by a variety of cancers, including hepatocellular carcinoma (HCC), gastric cancer (GC), colorectal cancer, pancreatic cancer, prostate cancer, glioma, leukemia, and lymphoma. Of all the studies, 3 recruited prostate cancer patients (33, 36, 47) and 11 recruited patients with cancers of the digestive system, including HCC (31, 34, 40–42, 46), GC (39, 43, 45), colorectal cancer (37), and pancreatic cancer (32). Frozen or formalin-fixed, paraffin-embedded (FFPE) tumor tissues were used in 14 studies (31–36, 39, 41–47), while 3 studies used serum or plasma samples (37, 38, 40). The quantitative real-time polymerase chain reaction (qRT-PCR) assay was used in 16 studies (31–43, 45–47), whereas the in situ hybridization (ISH) assay was used in only 1 study (44). Notably, miRNA classifiers used the median (9 studies) (31–36, 39, 46, 47), mean (1 study) (43) or other cutoff values (7 studies) (37, 38, 40–42, 44, 45) to classify the expression levels.
Main Characteristics and Results of the Eligible Studies
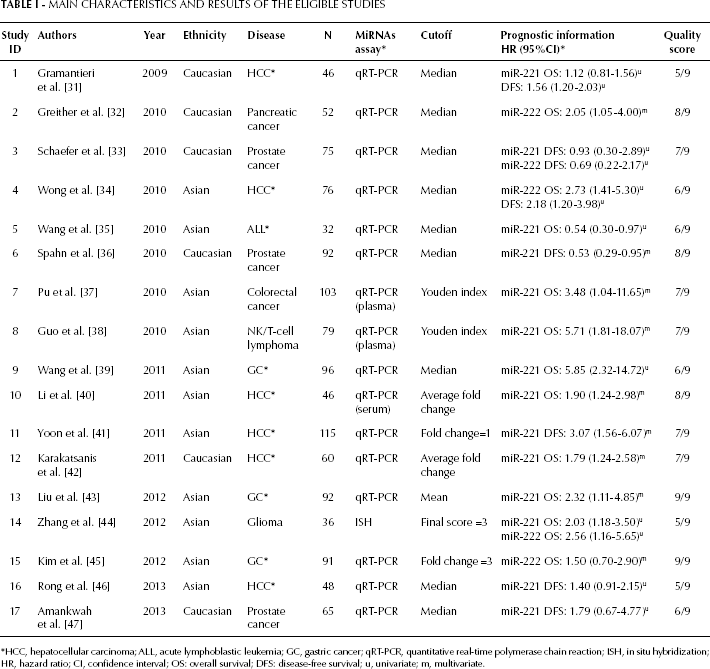
HCC, hepatocellular carcinoma; ALL, acute lymphoblastic leukemia; GC, gastric cancer; qRT-PCR, quantitative real-time polymerase chain reaction; ISH, in situ hybridization; HR, hazard ratio; CI, confidence interval; OS: overall survival; DFS: disease-free survival; u, univariate; m, multivariate.
Quality scores of the included studies ranged from 5 to 9 stars. Fourteen studies that had ≥6 stars were categorized as high-quality studies (32–43, 45, 47), while 3 studies that had <6 stars were categorized as low-quality studies (31, 44, 46).
Meta-analysis
The summary of our meta-analysis results for the prognostic significance of miR-221/222 expression in cancer is presented in Table II. Among the 17 studies eligible for the meta-analysis, 11 provided the estimated HR and 95% CI (33, 35–38, 40–43, 45, 47). In the remaining studies, these values were calculated from the presented data (32, 39, 46) or extracted from survival curves (31, 34, 44).
Meta-Analysis Results for Prognostic Significance of miR-221/222 Expression with Cancers
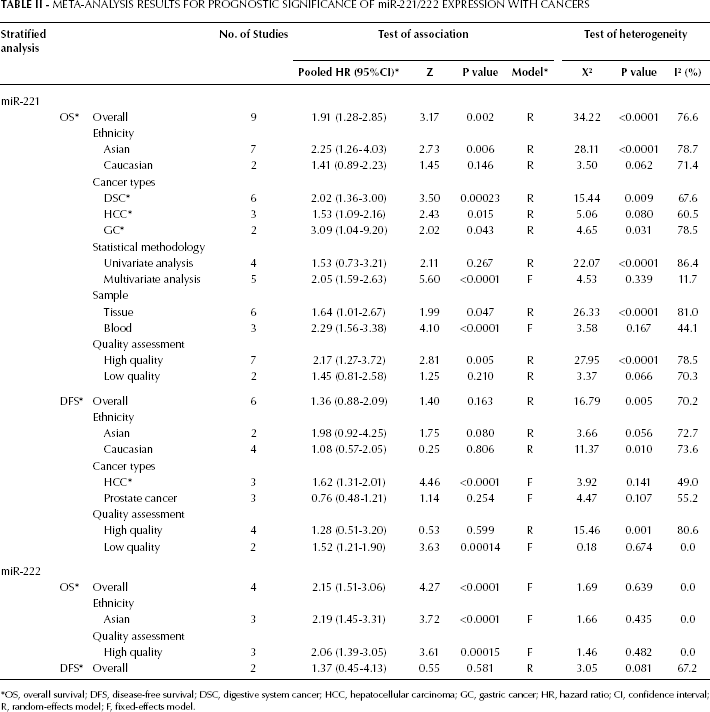
OS, overall survival; DFS, disease-free survival; DSC, digestive system cancer; HCC, hepatocellular carcinoma; GC, gastric cancer; HR, hazard ratio; CI, confidence interval; R, random-effects model; F, fixed-effects model.
Analysis for the prognostic significance of miR-221
Analysis for OS
When assessing the association of miR-221 expression with OS, 9 studies presented data valuable for analysis (31, 35, 37–40, 42–44). We found statistical heterogeneity among the studies evaluating miR-122 and OS, and thus we applied the random-effects model to calculate a pooled HR and its 95% CI. We found that an elevated miR-221 expression significantly predicted poor OS, with the pooled HR being 1.91 (95% CI: 1.28-2.85, p=0.002). The forest plot of studies comparing the HR of high vs low miR-221 expressions for OS is shown in Figure 2A. There was no evidence of publication bias either from the funnel plot or from the Egger's test. The funnel plot was symmetric and the p value of the Egger's regression intercepts was 0.122 (data not shown).
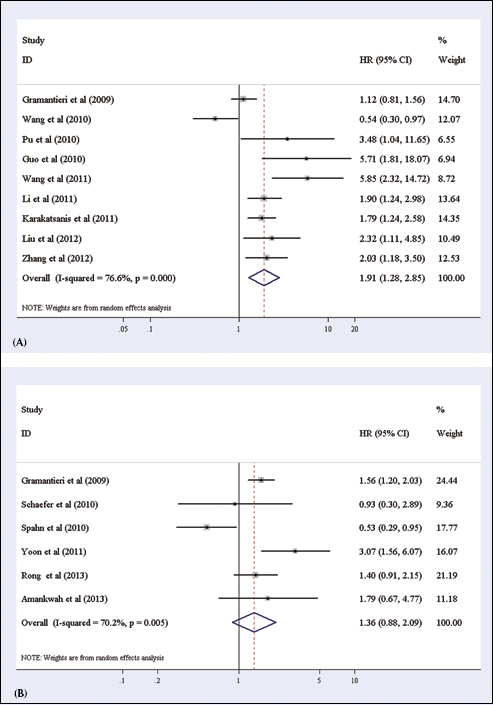
Forest plots of studies evaluating hazard ratios of high miR-221 expression as compared to low miR-122 expression:
Subgroup analyses were further performed to evaluate the effect of miR-221 expression on OS. Statistically significant heterogeneity was observed in all the subgroup analyses except for that of the multivariate results and blood sample. When stratified by ethnicity, miR-221 expression was significantly correlated with OS in Asians, with the combined HR being 2.25 (95% CI: 1.26-4.03, p=0.006). When stratified by cancer types, elevated miR-221 expression was associated with poor OS, with the combined HR being 2.02 (95% CI: 1.36-3.00, p=0.00023) in patients with digestive system cancers, 1.53 (95% CI: 1.09-2.16, p=0.015) in HCC patients, and 3.09 (95% CI: 1.04-9.20, p=0.043) in GC patients. When stratified by statistical methodology, miR-221 expression was significantly correlated with OS in subgroup analysis of multivariate results, with the combined HR being 2.05 (95% CI: 1.59-2.63, p<0.0001). When stratified by sample, elevated miR-221 expression was associated with poor OS, with the combined HR being 1.64 (95% CI: 1.01-2.67, p=0.047) in tissue, and 2.29 (95% CI: 1.56-3.38, p<0.0001) in blood. When stratified by quality assessment, elevated miR-221 expression was associated with poor OS in subgroup analysis of high-quality studies, with the combined HR being 2.17 (95% CI: 1.27-3.72, p=0.005).
Analysis for DFS
When assessing the association of miR-221 expression with DFS, 6 studies presented data valuable for analysis (31, 33, 36, 41, 46, 47). For studies evaluating RFS, a random-effects model was applied because of the heterogeneity between studies. We found no significant association between miR-221 expression and DFS, with the pooled HR being 1.36 (95% CI: 0.88-2.09, p=0.163). The forest plot of studies evaluating the HR of high vs low miR-221 expression for DFS is shown in Figure 2B.
Subgroup analyses were further performed to evaluate the effect of miR-221 expression on DFS. Statistically significant heterogeneity was observed in the subgroup analysis of ethnicity and high-quality studies. Subgroup analyses were stratified by ethnicity, cancer types, and quality assessment. The miR-221 expression was significantly correlated with DFS in subgroup analysis of HCC patients with the combined HR being 1.62 (95% CI: 1.31-2.01, p<0.0001), and in subgroup analysis of low-quality studies with the combined HR being 1.52 (95% CI: 1.21-1.90, p=0.00014).
Analysis for prognostic significance of miR-222
Analysis for OS
When assessing the association of miR-222 expression with OS, 4 studies presented data valuable for analysis (32, 34, 44, 45). The Q test of heterogeneity was not significant and we conducted the analysis using the fixed-effects model. We found that an elevated expression of miR-222 significantly predicted poor OS, with the pooled HR being 2.15 (95% CI: 1.51-3.06, p<0.0001). The forest plot of studies evaluating the HR of high vs low miR-222 expression for OS is shown in Figure 3A. There was no evidence of publication bias either from the funnel plot or from the Egger's test. The funnel plot was symmetric and the p value of the Egger's regression intercepts was 0.994.
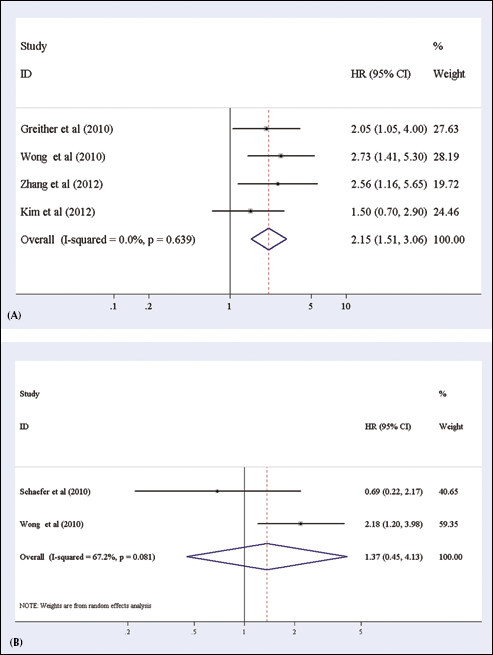
Forest plots of studies evaluating hazard ratios of high miR-222 expression as compared to low miR-122 expression:
Subgroup analyses were further performed to evaluate the effect of miR-222 expression on OS. The Q test of heterogeneity was not significant and we conducted the analysis using the fixed-effects model. MiR-222 expression was significantly correlated with OS in subgroup analysis of Asians with the combined HR being 2.19 (95% CI: 1.45-3.31, p<0.0001), and in subgroup analysis of high-quality studies with the combined HR being 2.06 (95% CI: 1.39-3.05, p=0.00015).
Analysis for DFS
When assessing the association of miR-222 expression with DFS, 2 studies presented data valuable for analysis (33–34). For studies evaluating RFS, a random-effects model was applied because of the heterogeneity between studies. We found no significant association between miR-222 expression and DFS, with the pooled HR being 1.37 (95% CI: 0.45-4.13, p=0.581). The forest plot of studies evaluating the HR of high vs low miR-222 expression for DFS is shown in Figure 3B.
Discussion
To the best of our knowledge this is the first meta-analysis to investigate the association between miR-221/222 expression and the survival of patients with cancer. Findings from the current meta-analysis demonstrated that the elevated expression of miR-221 and miR-222 both significantly predicted poor OS. Meanwhile, we found no significant association between miR-221/222 expression and DFS.
The pooled risks of miR-221 and miR-222 for OS were both statistically significant and quite high with regard to global HRs (1.91 and 2.15, respectively). Subgroup analyses were further stratified by ethnicity, cancer types, statistical methodology, sample, and quality of studies when assessing the association between miR-221 expression and OS. When stratified by ethnicity, the results have shown that an elevated expression of miR-221 was a predictor of worse outcome in Asians, but not in Caucasians. Ethnic differences may partly be due to various genetic backgrounds, dietary behaviors, environmental exposures, and other factors. When stratified by cancer types, a high expression of miR-221 was significantly correlated with OS in digestive system cancers, especially in HCC and GC. When stratified by the type of sample used (tissue or blood), the results were consistent with the overall analysis. When stratified by statistical methodology, the association between miR-221 expression and OS remained significant in multivariate analysis, but not in univariate analysis. In the multivariate analysis, the HR for high expression of miR-221 is adjusted for confounders such as some clinicopathological factors. The result is more convincing after adjustment in multivariate analysis than in univariate analysis. When excluding the low-quality studies, the association can still be observed in high-quality studies, which made the results even more convincing. When further assessing the association between miR-222 expression and OS, we found that the elevated expression of miR-222 was a predictor of poor OS in Asians, as well as in high-quality studies.
Recently, a few meta-analyses were performed to summarize the global predicting role of specific miRNAs in the survival of patients with a variety of cancers (48–49). Results from these studies provided supporting evidence for the prognostic value of miR-21 and miR-155 in cancer patients. Even though our data indicate that miR-221/222 is a promising prognostic factor of cancers, the current conclusion should be tempered for several reasons. Firstly, different sample sources were used among the studies. While tissue samples are widely used for miRNA studies, measuring circulating miRNAs might be more useful as a non-invasive technique (2). In order to minimize the effect of the different types of samples used, we conducted a subgroup analysis according to the sample source. The present meta-analysis examined the miR-221 expression levels in tissue and showed similar HR results for OS as those measuring miR-221 levels in serum/plasma. Secondly, there is no clear cutoff value for the expression of miR-221/222. Most of the researchers used median or mean values as the cutoffs. The lacking of abundant data on miR-221/222 expression in the global population remains an issue to be solved.
The association between miR-221/222 expression and cancer prognosis may be partly explained by the biological function of the miRNA cluster. MiR-221/222 can act as oncomiRs by targeting important tumor suppressor genes, such as phosphatase and tensin homolog deleted from chromosome 10 (PTEN), tissue inhibitor of metalloproteinase 3 (TIMP3), p27Kip1, p57, and Bim. Their overexpression can have several effects: inducing cell proliferation through the activation of the cell cycle, blocking TRAIL-induced apoptosis, and enhancing tumorigenicity of lung and liver cancer cells (9, 50). Moreover, recent studies have also demonstrated that miR-221 and miR-222 could regulate radiosensitivity, cell growth, and invasion of gastric cells (51). Accordingly, miR-221/222 could be used as a therapeutic tool to modulate the resistance or sensitivity to anti-cancer agents. Increasing miR-221/222 levels often affected various characteristics associated with epithelial to mesenchymal transition (EMT), including the invasive capacity (52, 53). This may lead to the overexpression of miR-221/222 in poorly differentiated, aggressive cancers, which is linked to poor outcome for patients. However, some controversy exists. MiR-221/222 can play a tumor-suppressive role in erythroleukemic cells (16). Consequently, much remains to be explored to fully determine the effective influence of the miRNA families on cancer development.
Our meta-analysis has some limitations. Firstly, a marked heterogeneity of subjects was detected in the OS and DFS analysis, a fact that may distort the meta-analysis. Differences in the baseline characteristics of the patients (age, stage, and ethnicity), study design, patient selection, cancer type, various cutoff values for miR-221/222 expression, and the duration of follow-up may be possible explanations for the heterogeneities we found across the studies. In this case, we attempted to minimize the effect by using a random-effects model. Secondly, the lack of individual HR data makes it harder to exclude the influences of confounding factors in this meta-analysis. Thirdly, we should be cautious when interpreting the results of DFS analysis and subgroup analysis in Caucasians due to the relatively small sample sizes of the studies.
In conclusion, our meta-analysis, representing a quantified synthesis of all eligible studies, has shown that elevated expressions of miR-221 and miR-222 are associated with poor survival in patients with cancers. The miR-221/222 cluster might be used as a potential therapeutic strategy in clinical practice. More work is required to fully elucidate the role of the miR-221/222 family in human tumors.
Footnotes
Acknowledgements
We thank all the people who gave help for this study.
Supplementary Material
Methodologic Quality of Studies Included in the Meta-Analysis
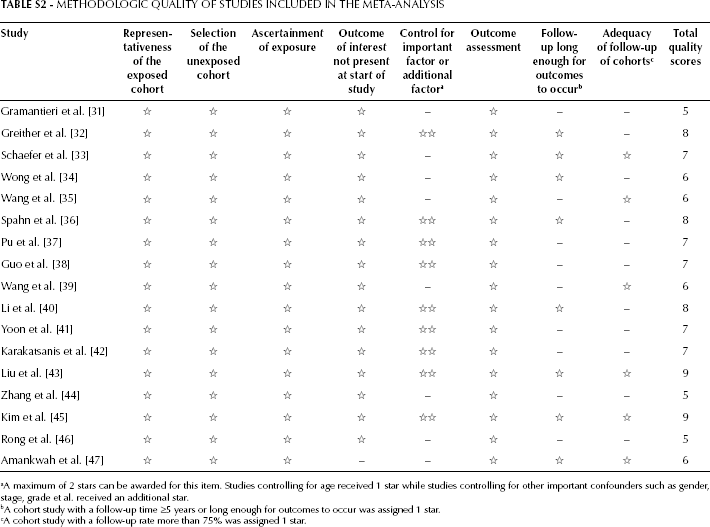
| Study | Representativeness of the exposed cohort | Selection of the unexposed cohort | Ascertainment of exposure | Outcome of interest not present at start of study | Control for important factor or additional factor a | Outcome assessment | Follow-up long enough for outcomes to occur b | Adequacy of follow-up of cohorts c | Total quality scores |
|---|---|---|---|---|---|---|---|---|---|
| Gramantieri et al. [31] | ⋆ | ⋆ | ⋆ | ⋆ | – | ⋆ | – | – | 5 |
| Greither et al. [32] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | ⋆ | – | 8 |
| Schaefer et al. [33] | ⋆ | ⋆ | ⋆ | ⋆ | – | ⋆ | ⋆ | ⋆ | 7 |
| Wong et al. [34] | ⋆ | ⋆ | ⋆ | ⋆ | – | ⋆ | ⋆ | – | 6 |
| Wang et al. [35] | ⋆ | ⋆ | ⋆ | ⋆ | – | ⋆ | – | ⋆ | 6 |
| Spahn et al. [36] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | ⋆ | – | 8 |
| Pu et al. [37] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | – | – | 7 |
| Guo et al. [38] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | – | – | 7 |
| Wang et al. [39] | ⋆ | ⋆ | ⋆ | ⋆ | – | ⋆ | – | ⋆ | 6 |
| Li et al. [40] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | ⋆ | – | 8 |
| Yoon et al. [41] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | – | – | 7 |
| Karakatsanis et al. [42] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | – | – | 7 |
| Liu et al. [43] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | ⋆ | ⋆ | 9 |
| Zhang et al. [44] | ⋆ | ⋆ | ⋆ | ⋆ | – | ⋆ | – | – | 5 |
| Kim et al. [45] | ⋆ | ⋆ | ⋆ | ⋆ | ⋆⋆ | ⋆ | ⋆ | ⋆ | 9 |
| Rong et al. [46] | ⋆ | ⋆ | ⋆ | ⋆ | – | ⋆ | – | – | 5 |
| Amankwah et al. [47] | ⋆ | ⋆ | ⋆ | – | – | ⋆ | ⋆ | ⋆ | 6 |
A maximum of 2 stars can be awarded for this item. Studies controlling for age received 1 star while studies controlling for other important confounders such as gender, stage, grade et al. received an additional star.
A cohort study with a follow-up time ≥5 years or long enough for outcomes to occur was assigned 1 star.
A cohort study with a follow-up rate more than 75% was assigned 1 star.
