Abstract
Background
This study aimed to evaluate the diagnostic performance of a novel nonquantitative methylation-specific reverse hybridization (MSRH) assay to detect secreted frizzled-related protein 2 (SFRP2) promotor methylation in fecal DNA.
Methods
SFRP2 promoter methylation was investigated in stool DNA isolated from 18 colorectal cancer (CRC) patients and 22 healthy controls using the MSRH assay based on methylation-specific DNA amplification followed by reverse hybridization of biotinylated amplicons to sequence-specific methylation detection probes, with MethyLight serving as a reference method.
Results
SFRP2 promotor methylation as determined by MSRH vs. MethyLight showed a sensitivity and specificity of 61.1% and 86.3% vs. 77.7% and 77.3%, respectively. Moderate agreement (κ = 0.54, 95% confidence interval [95% CI], 0.29-0.80, p<0.001) was observed between the 2 methods. However, the differences in SFRP2 promotor methylation observed between CRC patients and healthy individuals by both assays were statistically significant (p<0.001).
Conclusions
Our findings, although limited by the small sample size, do not support the use of the MSRH assay for CRC screening in stool.
Introduction
Colorectal cancer (CRC) is a major cause of cancer death worldwide. DNA methylation changes and mutations in oncogenes and tumor suppressor genes are key pathogenic events in CRC formation, driving the initiation and progression of the adenoma-carcinoma sequence (1, 2). As many CRC patients present with an advanced disease, early detection would seem to be a major factor in reducing rates of death from this disease (3). Established screening modalities – namely, colonoscopy, flexible sigmoidoscopy and barium enemas – are highly specific but limited by the availability of trained examiners and patient compliance (4). In this regard, stool analysis would offer considerable theoretical and practical benefits: it is noninvasive, does not require bowel preparation, might enable screening of the entire colon and rectum, and specimens are transportable. The last feature means that samples for at least some of the tests can be obtained without patients needing to visit their doctors, which is likely to increase patient acceptability (4).
Aberrant DNA methylation patterns have been associated with a large number of human malignancies, and the analysis of stool epigenetic markers, together with other genetic alterations, has now been recognized as an important noninvasive test for the early diagnosis of CRC (4, 5). For example, the genes encoding 4 members of the secreted frizzled-related protein (SFRP) family – namely SFRP1, SFRP2, SFRP4 and SFRP5 – have been found to be hypermethylated and are frequently not expressed in primary CRC (6-8). Moreover, the hypermethylation of SFRP genes is not only detected in CRC but also in adenomas and aberrant crypt foci (7, 8). By contrast, no methylation changes in the SFRP genes have been found in normal colorectal mucosa (8). In a previous study, we were able to show that SFRP2 but not SFRP1, SFRP4 or SFRP5 promotor methylation is a very sensitive DNA-based single marker for the identification of CRC in fecal samples (sensitivity 77%-90%, specificity 77%) (9). Over 10 years later, methylated SFRP2 is still considered a promising marker with great potential for early CRC detection in stool (6).
This study aimed to evaluate the diagnostic performance of a prototype nonquantitative methylation-specific reverse hybridization (MSRH) assay to detect SFRP2 promotor methylation in fecal DNA. For this purpose, SFRP2 promoter methylation was investigated in stool DNA isolated from 18 CRC patients and 22 healthy controls, using the MSRH assay, which is based on methylation-specific DNA amplification followed by reverse hybridization of biotinylated polymerase chain reaction (PCR) products to sequence-specific methylation detection probes, with MethyLight serving as a reference method.
Methods
Ethical standards
This study was approved by the Ethics Committee of the Medical University Innsbruck. It was carried out in accordance with the latest version of the Helsinki Declaration (Fortaleza 2013). All participants gave their written informed consent.
Study population
Fecal DNA samples obtained from 26 CRC patients and 23 endoscopically verified healthy controls were available, for which SFRP2 promotor methylation had been determined previously using MethyLight, a fluorescence-based real-time PCR assay (9). MethyLight enables the quantitative analysis of DNA methylation at a particular locus using DNA oligonucleotides that differentially anneal to deaminated DNA, thereby indicating the methylation patterns present in the native DNA. Of these 49 patients, sufficient amounts of DNA for subsequent analysis by MSRH could be retrieved for 40 individuals only. Of all of the participants (N = 40) investigated here, 25 (62.5%) were men and 15 (37.5%) were women. The median age was 57.4 years (range 18-90). Staging was performed according to the Union for International Cancer Control (UICC) guidelines, with 2 stage I and 16 stage II tumors (Tab. I). Histological subtypes included adenocarcinomas only. Three and 8 CRCs were located in the right and left colon, respectively, and 7 CRCs in the rectum.
SFRP2 methylation results obtained for 22 controls and 18 CRC patients
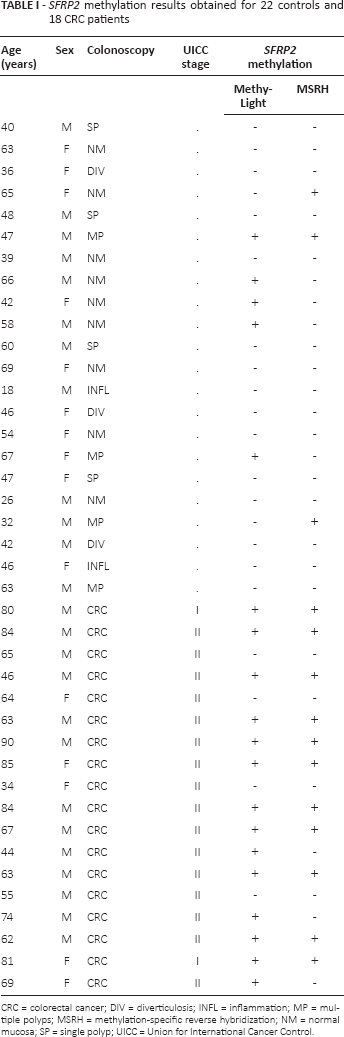
CRC = colorectal cancer; DIV = diverticulosis; INFL = inflammation; MP = multiple polyps; MSRH = methylation-specific reverse hybridization; NM = normal mucosa; SP = single polyp; UICC = Union for International Cancer Control.
MSRH
A series of candidate 15- to 20-mer oligonucleotides, encoding 2 CpG dinucleotides (CpG 1 and 2) each, were designed using the NCBI Reference Sequence Databank (NCBI RefSeq ID: NC_000004.12). Capture probes were immobilized as an array of parallel lines on a nitrocellulose membrane, which was then sliced into 3-mm test strips (10). Chemically synthesized DNA (Eurofins Genomics, Ebersberg, Germany) covering the genomic sequence 100 bp upstream and 100 bp downstream of each SFRP2-specific capture oligonucleotide was used to generate recombinant plasmid clones (TOPO TA Cloning Kit; Live Technologies, Carlsbad, USA) for each possible permutation of CpG methylation status (i.e., 4 plasmid clones for each capture probe with no methylation, CpG 1 or CpG 2 methylated, and both CpG 1 and 2 methylated). The plasmid clones served as PCR templates to control for hybridization specificity and to optimize oligonucleotide design. Bisulfite genomic DNA modification, which converts all unmethylated cytosines to uracils, followed a protocol published earlier (9). For reverse hybridization, methylation-specific PCR was performed on deaminated DNA using 5′-biotinylated primers.
To verify the DNA quality, each PCR also contained primers specific for the human β-actin gene (ACTB) and consisted of a total volume of 25 μL containing 10× PCR buffer (Qiagen, Hilden, Germany), 0.2 μmol/L of the forward and reverse primers, 1 unit of HotStar Taq DNA polymerase (Qiagen) and up to 50 ng of fecal DNA. Amplification conditions included a heat activation step at 95°C for 15 minutes, followed by 45 cycles of denaturation at 95°C for 45 seconds, annealing at 66°C for 45 seconds, and extension at 72°C for 45 seconds.
Biotinylated amplification products were hybridized to nitrocellulose test strips using fully automated instrumentation (profiBlot T48; TECAN, Grödig, Austria), strictly controlling temperature (45°C ± 0.5°C). Specifically bound PCR products were visualized at room temperature using streptavidin-alkaline phosphatase conjugate and enzymatic color reaction (10). The final array design was based on 2 capture probes both located within the same CpG island, that discriminated best between methylated (i.e., CpG 1 and 2 methylated) and unmethylated DNA (i.e., CpG 1 and/or 2 unmethylated). A sample was considered positive if a hybridization signal was obtained from either one or both capture probes. Primer and oligonucleotide capture probes specific for methylated DNA are listed in Table II.
ACTB and SFRP2 gene primer and probe sequences for the MSRH
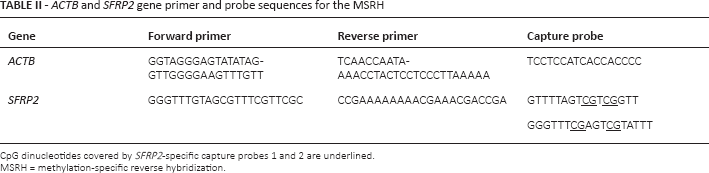

CpG dinucleotides covered by SFRP2-specific capture probes 1 and 2 are underlined.
MSRH = methylation-specific reverse hybridization.
With respect to our reference method MethyLight, the MSRH assay covers the same CpG island but addresses different CpG dinucleotides contained in the 2 capture probes. The MSRH operators were blinded to the disease status of patients as well as to the MethyLight results.
Statistical analysis
Pearson's chi-square test was used for comparison of SFRP2 promotor methylation and KRAS mutation status between 18 CRC patients and 22 healthy controls. Cohen's kappa (κ) with 95% confidence intervals (95% CI) was performed to calculate the agreement between MSRH and MethyLight. A p value <0.05 was considered statistically significant. SPSS software version 11.0 for Windows (IBM Corporation, Armonk, NY, USA) was used for statistical analysis.
Results
The MSRH assay detected SFRP2 promotor methylation in 11 of 18 CRC patients, and in 3 of 22 healthy controls, demonstrating a sensitivity of 61.1% and a specificity of 86.3%. Figure 1 shows some typical reactivity patterns obtained after reverse hybridization and color development. By comparison, MethyLight identified SFRP2 promotor methylation in 14 of 18 individuals with CRC (sensitivity: 77.7%) and in 5 of 22 healthy controls (specificity: 77.3%) (Tab. I). Moderate agreement (κ = 0.54, 95% CI, 0.29-0.80, p<0.001) was observed between the 2 methods. However, the differences in SFRP2 promotor methylation observed between CRC patients and healthy individuals by both assays were statistically significant (p<0.001).
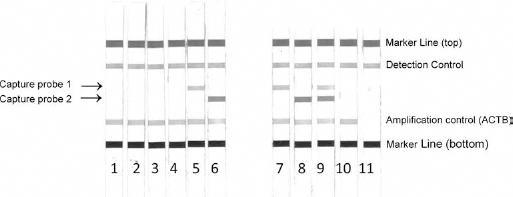
Detection of SFRP2 promotor methylation by methylation-specific polymerase chain reaction (PCR) and reverse hybridization. Typical reactivity patterns after enzymatic color development are shown. Strips on the left show combinations of recombinant plasmid clones incorporating unmethylated (strips 1-2), hemimethylated (strips 3-4) and fully methylated (strips 5-6) DNA sequences. Strips on the right show fecal DNA samples, which were found to be methylated (strips 7-9) or unmethylated (strip 10). None template control (strip 11). To control for quality of extracted DNA and functionality of reagents, an ACTB-specific (amplification control) and a detection control signal must be present, respectively.
Conclusions
Recently, several biomarkers have been proposed for CRC screening in serum and stool. In particular, DNA methylation patterns of genes such as SEPT9, SFRP2 and ALX4 are among the most interesting markers (11, 12). In this respect, SFRP2 promotor methylation has been identified as a promising noninvasive biomarker for the detection of CRC in stool (4-6, 9, 13). Therefore it was of interest to evaluate the diagnostic performance of a prototype MSRH assay for the detection of SFRP2 promoter methylation in stool DNA isolated from 18 CRC patients and 22 healthy controls, with MethyLight serving as a reference method.
The MSRH assay under investigation here detected SFRP2 promotor methylation in 11 of 18 CRC patients and in 3 of 22 healthy controls, demonstrating a sensitivity of 61.1% and a specificity of 86.3%, respectively. Hence, MSRH analysis resulted in a distinct decrease in sensitivity (61.1% vs. 77.7%) and a slight increase in specificity (86.3% vs. 77.3%) compared with our reference method MethyLight. However, the MethyLight results were in agreement with 2 recent meta-analyses which found the pooled diagnostic performance of SFRP2 methylation for CRC detection in stool to range from 71% to 79% (5, 13).
Therefore, our findings, although limited by the small sample size, do not support the use of the MSRH assay for CRC screening in stool.
Footnotes
Acknowledgments
The authors are grateful to Bettina Rauscher (ViennaLab) for excellent technical assistance.
Financial support: No grants or funding have been received for this study.
Conflict of interest: None of the authors has any financial interest related to this study to disclose.
