Abstract
Background
Recent studies have revealed that microRNAs (miRNAs) play important roles as oncogenes or tumor suppressors in tumorigenesis and tumor development, by negatively regulating protein expression. A previous study of microarrays identified that miR-411 was down-regulated in renal cell carcinoma (RCC), while few studies investigating the role of miR-411 in the pathogenesis of RCC have been performed.
Methods
We assessed the miR-411 expression in RCC and paired adjacent normal tissues, as well as in RCC cell lines and a normal renal cell line, by quantitative reverse transcriptase polymerase chain reaction (qRT-PCR). Furthermore, the effects of miR-411 on RCC and normal renal cell proliferation, apoptosis and migration were determined using MTT assay, CCK-8 assay, flow cytometry and scratch wound assay following restoration of miR-411 with synthetic mimics.
Results
Results of qRT-PCR indicated that the expression of miR-411 was down-regulated in RCC tissues and cell lines when compared with adjacent normal tissues and a normal renal cell line. Further, results of CCK-8, MTT, cell scratch and transwell assay showed that over-expression of miR-411 suppressed RCC cell (786-0 and ACHN) proliferation and migration. Flow cytometry assay revealed that miR-411 could induce RCC cell apoptosis. However, overexpression of miR-411 had no obvious effect on normal renal cell line 293T
Conclusions
To sum up, miR-411 is significantly down-regulated and plays a role as a tumor suppressor in RCC. Further studies are warranted to determine the mechanisms of miR-411 in RCC pathogenesis and define the target genes of miR-411 in RCC.
Introduction
Renal cell carcinoma (RCC) is the third most common uro-logical cancer, it accounts for approximately 3% of all adult malignancies. RCC represents 90% of all kidney cancers (1, 2). In the early stages of the disease, RCC patients have no distinctive clinical features, so RCC is always diagnosed accidentally. However, one third of patients have metastases at diagnosis, and among those with a clinically localized disease, 30% to 40% will develop recurrence and/or metastases after surgery (3, 4). What is more, RCC is resistant to chemotherapy and radiotherapy, and has a poor prognosis (5, 6). So it is essential to understand the mechanism of RCC and search for new biomarkers of RCC based on that mechanism.
MicroRNA (miRNA) is a kind of noncoding RNA with a length of ∼22 nucleotides, which plays important roles in different cellular processes, including cell cycle, proliferation, differentiation, metabolism and apoptosis (7, 8). Recent studies have found that miRNA can regulate gene expression posttranscriptionally by incompletely binding to the 3′ untranslated region (3′ UTR) of mammalian target mRNAs and causing translational inhibition and/or mRNA degradation (9, 10). For the incomplete binding with targeted mRNA, a miRNA can regulate different targeted genes, and a gene can be regulated by different miRNAs. So investigations of miRNAs have provided new directions to understand the mechanism of tumorigenesis in RCC. Previous studies have revealed that miR-411 was down-regulated in clear-cell RCC in a microRNA profiler array (11). In the present study, expression of miR-411 was detected by quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) in RCC tissues and paired normal tissues. And the functions of miR-411 in RCC was also explored.
Materials and methods
Patient samples
Forty-three patients with RCC who received partial or radical nephrectomy were included in this study. RCC tissues and adjacent normal tissues from each patient were snap-frozen in liquid nitrogen immediately after resection. Written informed consent was obtained from all patients. The study was approved by the institutional review board of Shenzhen Second People's Hospital. The tissues collected were reviewed and classified by hematoxylin and eosin (H&E) staining. The clinical and pathological characteristics of the patients are presented in Table I.
Relationship between expression of miR-411 and clinicopathological characteristics of renal cell cancer (RCC) patients
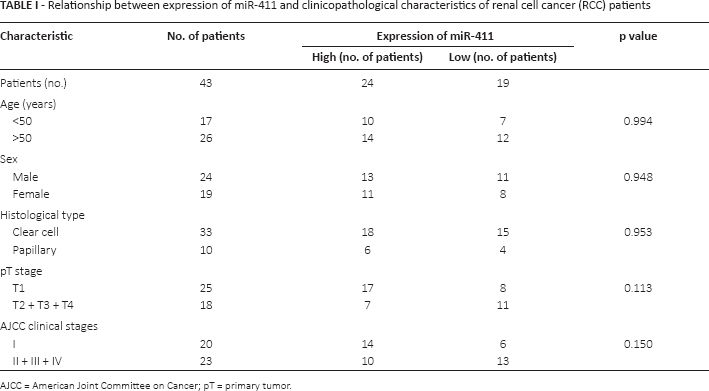
AJCC = American Joint Committee on Cancer; pT = primary tumor.
Cell culture
RCC cells (786-O and ACHN) and human embryonic kidney cell 293T were used in the study (American Type Culture Collection, Rockville, MD, USA). The cells were cultured in Dulbecco's modified Eagle's medium (DMEM; Invitrogen, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, USA) at 37°C in a 5% CO2 atmosphere.
Cell transfection
MiR-411 mimic or mimic Negative control (NC) was synthesized by GenePharma (Suzhou, China) and transfected into cells with Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions.
Real-time quantitative PCR
Total RNA was extracted from the tissue samples or the transfected cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. cDNA was converted from total RNA using RT-PCR with the miScript Reverse Transcription Kit (Qiagen, Hilden, Germany). U6 was used as the internal control. The primer sequences were as follows: miR-411 forward primer: TAGTAGACCGTATAGCGTACG. The reverse primer was provided by Qiagen. U6 forward primer: 5′-CTCGCTTCGGCAGCACA-3′; U6 reverse primer: 5′-ACGCTTCACGAATTTGCGT-3′.
Cell proliferation assay
CCK-8 and methylthiazolyldiphenyl-tetrazolium bromide (MTT) assay were performed to detect cell proliferation. Cells were seeded in a 96-well plate for 24 hours, then transfected with miR-411 mimic or mimic NC and cultured in normal medium. In the CCK-8 assay, 10 μL of Cell Counting Kit-8 (Beyotime Institute of Biotechnology, Shanghai, China) was added into the wells, to be evaluated at 0, 24, 48, and 72 hours after transfection. One hour later, the optical density (OD) of each well was measured with an ELISA microplate reader (Bio-Rad Laboratories, Hercules, CA, USA). For the MTT assay, 20 μL of MTT (5 mg/mL; Sigma-Aldrich, St. Louis, MO, USA) was added into the wells, to be evaluated at 0, 24, 48 and 72 hours after transfection, and 4 hours later, the mixed medium was replaced by 150 μL of dimethyl sulfoxide (DMSO; Sigma-Aldrich). After that, the 96-well plates were agitated for 15 minutes at room temperature. Then the OD of each well was measured. The experiments were performed in sextuplicate and repeated at least 3 times.
Cell scratch assay
Cell scratch assay was performed to assess cells' migratory ability for RCC cell lines (786-0 and ACHN) and a normal renal cell line (293T). Cells were seeded in the 6-well plate. Twenty-four hours later, the cells were transfected by miR-411 mimics or mimic NC. Six hours later, the cell monolayer was scratched with a sterile 1-mL pipette tip to generate a line-shaped wound. Then the cells were cultured in DMEM with 5% FBS. Zero and 24 hours after making the scratches, images of the scratches were acquired with a digital camera. The experiments were performed in triplicate and repeated 3 times.
Transwell assay
Transwell assay was performed to assess the cells' migratory ability. In the assay, transwell chamber inserts (BD, New York, NY, USA) were used following the manufacturer's protocol. Transfected cells (1 × 10 4 ) in 200 μL of DMEM were seeded in the upper channel of the inserts. DMEM with 10% FBS was in the lower channel of the inserts. Cells were allowed to migrate for 36 hours. The cells that had migrated to the bottom of the inserts were stained with crystal violet and counted using a microscope. The experiments were performed in triplicate and repeated at least 3 times.
Flow cytometry analysis of cell apoptosis
The apoptotic rates of cells were measured by flow cytometry assay with an Alexa Fluor® 488 Annexin V/Dead Cell Apoptosis Kit (Invitrogen). Cells were seeded in the 6-well plate, and 24 hours later the cells were transfected with miR-411 mimics or mimic NC. Then 48 hours later, the cells were harvested and stained with an Alexa Fluor® 488 Annexin V/Dead Cell Apoptosis Kit according to the manufacturer's instructions. Then flow cytometry (EPICS, XI-4; Beckman Coulter, Inc.250 S.Kraemer Boulevard Brea, CA 92821, USA) was used to analyze the apoptosis rate. The experiments were performed in triplicate and repeated at least 3 times.
Statistical analysis
A paired t-test was used to compare the expression levels of miR-411 in matched tissues. Student's test was used to analyze assays for characterizing phenotypes of cells. All statistical analysis was carried out with SPSS 19.0 statistical software package (SPSS Inc., Chicago, IL, USA). Results in all figures are shown as means ± standard error. A p value of <0.05 was deemed statistically significant.
Results
MiR-411 was significantly down-regulated in RCC tissues and cell lines
qRT-PCR was performed to detect the expression levels of miR-411 in 43 pairs of tissues. The expression of miR-411 in adjacent normal tissue was 5.00 ± 0.32 times that in RCC tissues (p<0.001) (Supplementary Figure 1A, available online at www.biological-markers.com – Expression of miR-411 in tissues). The Log2 ratios (T/N) in all tissues are shown in Supplementary Figure 1B (available online at www.biological-markers.com – Expression of miR-411 in tissues), and miR-411 was down-regulated in 30 RCC tissues compared with adjacent normal tissues. In cell lines, the results revealed that miR-411 was significantly down-regulated in RCC cell lines (ACHN and 786-O) compared with the 293T cell line (Supplementary Figure 1C, available online at www.biological-markers.com – Expression of miR-411 in tissues). Subsequently, the relationship between the expression levels of miR-411 and clinicopathological characteristics of RCC patients was explored. Patients were separated into 2 groups (high-expression group and low-expression group) according to the mean of the-ΔΔCT values for all of the samples. Then the chi-square test was performed to explore the relationship between the expression and clinicopathological characteristics of patients. However, no relationship was found between the expression of miR-411 and the features of age, sex, T stage, histological type, Fuhrman grade and American Joint Committee on Cancer (AJCC) clinical stage (p>0.05; Tab. I). However, bias cannot be ruled out, due to the limited number of cases, and these results should be verified in more cases.
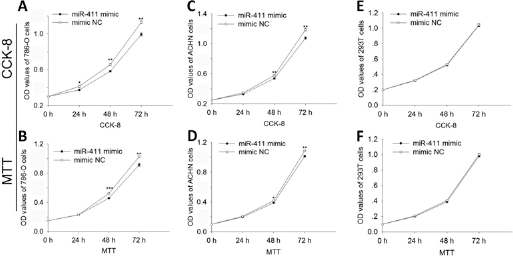
Cell proliferation assay for 786-O (
Transfection of miR-411 mimic overexpressed miR-411
RCC cell lines (786-O and ACHN) and normal renal cell line (293T) were cultured and transfected with miR-411 mimic or mimic NC. At 48 hours after transfection, the expression levels of miR-411 in 293T, 786-O and ACHN cells were detected with qRT-PCR. Results are shown in Supplementary Figure 2 (available online at www.biological-markers.com – Relative expression of miR-411 in 786-O, ACHN and 293T cells after transfection). The expression level of miR-411 was 648.31 in 786-O cells (p<0.01), which was 358.64 times higher than that in cells transfected with mimic NC.
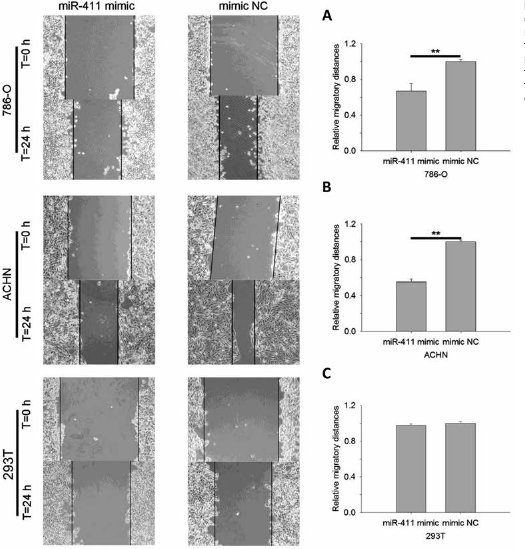
Wound scratch assay for 786-O, ACHN and 293T cells. (
Overexpression of miR-411 inhibited cell proliferation
CCK-8 and MTT assay were performed to determine whether miR-411 could affect cell proliferation. In both 786-O (Fig. 1A, B) and ACHN (Fig. 1C, D) cells transfected with miR-411 mimic, cell growth arrest was observed. However, there was no obvious growth arrest in 293T cells that were transfected with miR-411 mimic (Fig. 1E, F). The results showed that miR-411 could regulate RCC cell proliferation.
Overexpression of miR-411 inhibited cell migration
Cell scratch assay and transwell migration assay were performed to assess the function of miR-411 in RCC cells. Results of cell scratch assay are shown in Figure 2. MiR-411 suppressed cell migratory distance by 32.85% (786-O, p<0.01) and 44.69% (ACHN, p<0.01). There was no significant difference in the distances between 293T cells transfected with miR-411 mimic and those transfected with NC. Results of transwell assay showed that migratory cells transfected with miR-411 mimic were suppressed by 46.68% (786-O; p<0.05) and 54.03% (ACHN; p<0.01) (Fig. 3). For 293T cells, only a few cells migrated to the bottom surface of the transwell (results not shown). The results showed that miR-411 could suppress RCC cell migration.
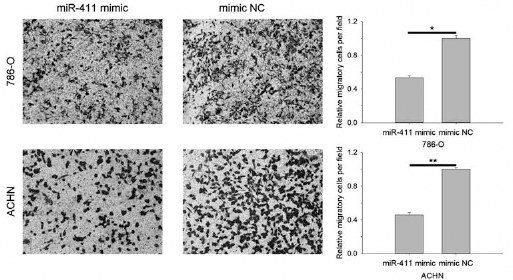
Results of transwell migration assay. MiR-411 suppressed 786-O and ACHN cells' ability to migrate; NC = Negative control (*p<0.05, **p<0.01).
MiR-411 induced cell apoptosis
Whether miR-411 could regulate RCC cell apoptosis was assed by flow cytometry. Forty-eight hours after transfection, cell apoptosis was detected, as shown in Figure 4. In 786-O cells, the apoptosis rate was 25.21% (miR411 mimic) vs. 13.00% (mimic NC) (p<0.05). In ACHN cells, the apoptosis rate was 17.70% (miR411 mimic) vs. 7.51% (mimic NC) (p<0.05). In 293T cells, the apoptosis rate was 5.86% (miR411 mimic) vs. 5.38% (mimic NC) (p>0.05). The results revealed that miR-411 could induce cell apoptosis in RCC.
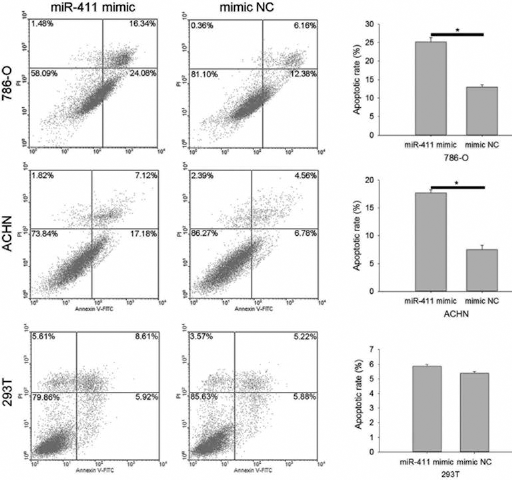
Up-regulation of miR-411 induced cell apoptosis of 786-O and ACHN; NC = Negative control (*p<0.05).
Discussion
Many studies have identified the associations between miRNAs and tumorigenesis. MiRNAs are a kind of noncoding RNA which can negatively regulate the expression of targeted gene posttranscriptionally by binding with the 3′UTR of targeted mRNA (12-15). Previous studies have indicated that in different tumors, miRNAs have unique expression profiles, suggesting their potential role as cancer biomarkers (16). In RCC, a lot of miRNAs have been shown to be dysregulated, and they play important roles during RCC initiation and progression, such as miR-362-3p (17), miR-16 (18), miR-508-3p (19) and so on. To the best of our knowledge, the present study is the first to demonstrate that miR-411 is down-regulated in RCC tissues, which is similar to miR-411 in breast cancer (20, 21) as reported previously. However, in the present study, bias cannot be excluded, due to the limited number samples. Thus, in future studies, expression of miR-411 in RCC tissues and the relationship between its expression and clinicopathological characteristics remain to be verified.
In the present study, results of qRT-PCR showed that miR-411 was down-regulated in RCC tissues compared with adjacent normal tissues, which suggested that miR-411 was a potential biomarker for RCC. Results of cellular function assay indicated that overexpression of miR-411 by transfection can suppress RCC cell (786-O and ACHN) proliferation and migration. It was also observed that overexpression of miR-411 can induce RCC cell apoptosis. However, in the normal renal cell line 293T, overexpression of miR-411 has no obvious effect on 293T cell proliferation, migration and apoptosis. Thus, we could imagine that the downstream gene or protein of miR-411 plays an important role in RCC cells, while not in a normal renal cell line.
MiR-411 has been shown to be abnormally expressed in other cancers, including hepatocellular carcinoma (HCC) (22), lung adenocarcinoma (23) and breast cancer (20, 21). In HCC tissues, miR-411 is overexpressed (22), which is different from the expression of miR-411 in RCC. In this study, it was also described that miR-411 could down-regulate ITCH, which resulted in the up-regulation of cyclin D1 and c-Myc, and thus promote HCC cell proliferation. Nadal et al (23) reported that knockdown of miR-411 significantly reduced cell migration in a lung AC cell line, and miR-411 was overexpressed in tumors from patients who relapsed systemically compared with others. Another study, conducted by Zhao et al, investigated the role of miR-411 in lung cancer (24) and revealed that miR-411 could promote proliferation of lung cancer cells by inhibiting the expression of tumor suppressor FOXOl. Thus, in lung cancer and HCC, miR-411 plays a role as an oncogene. However, miR-411 plays a role as a tumor suppressor in breast cancer by inhibiting breast cancer cell growth and metastasis via targeting GRB2 (20). In different cancer types, miR-411 could have different roles, and finally, our study suggested that miR-411 functions as a tumor suppressor in RCC.
MiR-411 was also involved in some diseases other than tumors. Wang et al (25) reported that miR-411 was down-regulated in osteoarthritis and could regulate matrix metalloproteinase 13 (MMP-13) in chondrocytes. Another study, conducted by Harafuji et al (26), found that miR-411 was over-expressed in facioscapulohumeral muscular dystrophy myoblasts, and YAF2 was a potential regulatory target of miR-411. In addition, miR-411 was reported to be associated with drug resistance in malignant pleural mesothelioma (MPM) (27). In that study, the researchers demonstrated that miR-379/411 cluster could regulate IL-18 and contribute to drug resistance in MPM, which provided a potential therapeutic opportunity for MPM patients.
In conclusion, the results of the present study demonstrate that miR-411 is significantly down-regulated in RCC tissues and plays a role as tumor suppressor in RCC cell lines. And in a 293T cell line, miR-411 has no obvious effect. Further study will focus on the pathway of miR-411 in RCC and the possibility of miR-411 being a biomarker for RCC patients.
Footnotes
Financial support: This work was supported by the Shenzhen Basic Research Project (JCYJ20140414170821327).
Conflict of interest: None declared.
