Abstract
Background
Our previous work indicated that, first, the embryonic ectoderm development (EED) gene is a candidate gene associated with the pathogenesis of ulcerative colitis (UC) and, second, that the haplotypes of the EED polymorphism are one of the markers for UC susceptibility. The risk of developing colorectal cancer (CRC) increases in patients with inflammatory bowel disease.
Aim
The present study aimed at determining the association between polymorphisms in the EED gene and CRC.
Methods
Genotype analysis of EED single nucleotide polymorphisms (SNPs) was performed with high-resolution melting analysis, and the genotype and allele frequencies of the EED SNPs were compared between CRC patients and healthy controls. The haplotype frequencies of EED for multiple loci were estimated using the expectation maximization (EM) algorithm.
Results
Our study had a power of 76.6% at a 0.05 significance level. Genotype and allele frequencies of the SNPs and haplotype frequencies of the EED gene in CRC patients were not significantly different from those in healthy controls. Only the allele frequency of g.-1850G>C in the rectal cancer (RC) patient group was significantly different from that of the control group (p=0.04). Similarly, the genotype and allelic frequencies of the EED SNPs for either tumor site (left or right) or tumor stage were not significantly different from those in healthy controls. However, our data show an association between the g.-993G>C polymorphism in the EED gene and the presence of lymph node metastasis in CRC.
Conclusions
These results suggest that the SNPs of the EED gene might not be associated with susceptibility to CRC. However, this study shows that the allele frequency of g.-1850G>C in the RC patient group was significantly different from that in the control group (p=0.04) and that g.-993G>C may play a role in the lymph node metastatic process of CRC.
Keywords
Introduction
Colorectal cancer (CRC) is the third most common cancer worldwide (1), and the third leading cause of cancer-related mortality in the United States (2). Its etiology is very complex and is not fully understood. Numerous epidemiological and biological studies have suggested that the risk factors of CRC, including nutrition, physical inactivity, obesity, and diabetes, play a critical role in the etiology of the disease (3, 4). A longer duration and a greater anatomic extent of colitis, and the severity of this inflammation correlate with the risk of CRC in patients with inflammatory bowel diseases (IBD), both ulcerative colitis (UC) and Crohn's disease (CD) (5, 6).
The embryonic ectoderm development protein (EED, also called WAIT-1) is involved in multiple cellular protein complexes and is a member of the superfamily of the Polycomb group (PcG) proteins (7, 8) that mediate silencing of hundreds of genes important for embryonic development, cell proliferation and differentiation (9–11). EED is a member of the Polycomb repressive complexes (PRCs) that are found in Drosophila and mammals (12). Several types of PRCs, such as PRC1, PRC2 and PRC3, have been identified (13). The PRC2/3 complex consists of at least 5 components: EED, EZH2, SUZ12, RbAp38, and AEBP2 (14, 15).
Previously, we conducted a study to find IBD-related genes by cDNA microarray analysis using mRNAs isolated from healthy controls and IBD patients. Several tens of genes were identified as having higher or lower mRNA levels in IBD patients (16). One of them, the EED gene, was identified as a candidate gene because its expression levels were found to decrease in both CD and UC patients but not in healthy controls. We also showed that the genotype and allele frequencies of the g.-1850G>C single nucleotide polymorphism (SNP) of the EED gene in UC patients were significantly different from those in healthy controls (17).
Sporadic and colitis-related CRCs are associated with inflammation and inflammatory diseases, such as IBD (18). The severity of inflammation correlates with the risk of CRC in patients with IBD (6). A recent report suggested that the PRC2/EED-EZH2 complex is overexpressed in breast cancer lymph node metastasis compared with primary tumors (19). However, there was no information on the relation between genetic variations of the EED gene and CRC.
In this study, to determine whether EED SNPs are associated with susceptibility to CRC, we analyzed the allelic and genotypic frequencies of the EED SNPs and compared them between colorectal cancer patients and healthy controls. We also investigated the haplotype frequencies of these SNPs between the colorectal cancer patient group and the healthy control group.
Materials and Methods
Patients and DNA Samples
DNA samples used in this study were provided by the Biobank of Wonkwang University Hospital, a member of the National Biobank of Korea; this Biobank is supported by the Ministry of Health and Welfare Affairs. We obtained genomic DNA samples from 292 CRC patients (107 males and 185 females) and 535 healthy controls (334 males and 201 females). The mean age of the CRC patients was approximately 62.9 years while it was 40.7 years for the healthy controls. CRC patients included 150 cases of rectal cancer (RC) and 142 cases of colon cancer (CC). For some of the analysis the CC patients were divided into 2 groups according to the site of lesion: right or left CC. Cancer stages of CRC were classified using the American Joint Committee on Cancer (AJCC) staging manual, 7th edition (20). Genomic DNA was extracted from peripheral blood leukocytes using a standard phenol-chloroform method or using a Genomic DNA Extraction kit (iNtRON Biotechnology, Korea) according to the manufacturer's instructions. CRC patients were recruited from our outpatient clinic at the Wonkwang University Hospital. Patients were classified according to their clinical features, endoscopic findings and histopathological examinations. Healthy controls were recruited from the general population, and they underwent comprehensive medical screening at the Wonkwang University Hospital. All subjects in this study were of Korean ethnicity and lived in the same area.
SNP selection
On the basis of the Genbank dbSNP database (http://www.ncbi.nlm.nih.gov), we selected 3 SNPs (rs7952481, rs1391221, and rs974144) on the EED gene. Information on rs7952481 (g.-1850G>C), rs1391221 (g.-993G>C), and rs974144 (g.12351C>T) SNPs was derived from the NCBI SNP database. SNPs with unknown heterozygosity and minor allele frequency (below 5%) in the Asian population were excluded. The reference sequence for the EED gene was based on the sequence of the human chromosome 11, NC_000011.9.
Genotype analysis
Genotype analysis of the SNPs on the EED gene was performed by high-resolution melting analysis, as previously described (17).
Statistical analysis
The χ2 test was used to estimate the Hardy-Weinberg equilibrium (HWE). Pairwise comparison of the biallelic loci was employed for the analysis of linkage disequilibrium (LD). Haplotype frequencies of the EED gene for multiple loci were estimated using the expectation maximization (EM) algorithm with SNPAlyze software (DYNACOM, Japan). Logistic regression analysis (SPSS 11.5) was used to calculate the odd ratios (with 95% confidence intervals). A p value of less than 0.05 was considered statistically significant.
Results
To determine whether the EED SNPs are associated with susceptibility to CRC, genotypes of the EED SNPs were determined using the high-resolution melting analysis. We compared genotypes and allele frequencies in CRC patients (n=292) and healthy controls (n=535). This study had a power of 76.6% at a significance level of 0.05. The genotype and allele frequencies of g.-1850G>C, g.-993G>C and g.12351C>T of EED in CRC patients were not significantly different from those in healthy controls (Tab. I).
Genotype and Allele Frequency Analyses of the EED Gene Polymorphisms in CRC Patients and Healthy Controls
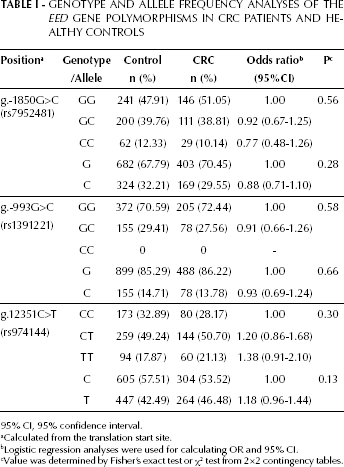
95% CI, 95% confidence interval.
Calculated from the translation start site.
Logistic regression analyses were used for calculating OR and 95% CI.
Value was determined by Fisher's exact test or χ2 test from 2×2 contingency tables.
We also analyzed the EED genotype and allele frequencies of the CC patients, RC patients, and healthy controls. The genotype frequency of g.-1850G>C, and the genotype and allele frequencies of g.-993G>C and g.12351C>T for EED were also not significantly different between healthy controls and CC or RC patients. However, the allele frequency of g.-1850G>C in the RC patient group was significantly different from that in the control group (p=0.04; Tab. II). We further compared EED genotype and allele frequencies between the 2 subgroups of CC (left CC and right CC) and healthy controls. The genotype and allele frequencies of g.-1850G>C, g.-993G>C, and g.12351C>T of EED were not significantly different between healthy controls and both right and left CC (Tab. III).
Genotype and Allele Frequency Analyses of the EED Gene Polymorphisms in RC Patients, CC Patients and Healthy Controls
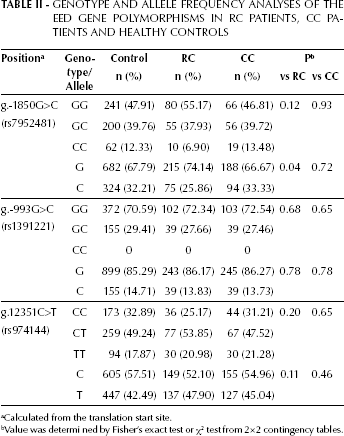
Calculated from the translation start site.
Value was determi ned by Fisher's exact test or χ2 test from 2×2 contingency tables.
Genotype and Allele Frequency Analyses of the EED Gene Polymorphisms in Healthy Controls and the two Subtypes of CC Patients (Right and Left Colon Cancer)
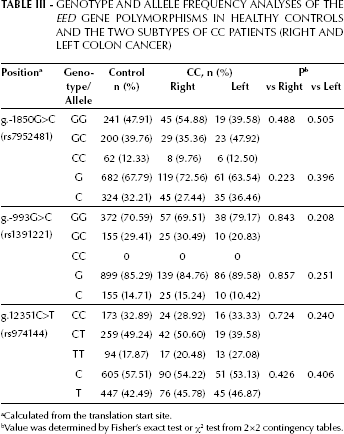
Calculated from the translation start site.
Value was determined by Fisher's exact test or χ2 test from 2×2 contingency tables.
In addition, to determine whether the EED SNPs were associated with cancer stage, CRC patients were divided on basis of the primary tumor and regional lymph node staging. Considering the stage of the primary tumor, patients were included in the TA group if tumor stage was T1 or T2, and in the TB group if tumor stage was T3 or T4. Considering the regional lymph node involvement, patients were included in the N(-) group if staged N0, and in the N(+) group if staged N1 or N2 The genotype and allele frequencies of g.-1850G>C, g.-993G>C, and g.12351C>T in different primary tumor or lymph node stages in CRC patients were not significantly different from those in healthy controls (Tab. IV). Thus, these results suggest that the EED SNPs were not associated with susceptibility to CRC. Interestingly, however, we found that the genotype and allelic frequencies of the g.-993G>C SNP were significant different between patients in the N(+) and N(-) groups (p=0.02 and p=0.03, respectively).
Genotype and Allele Frequencies of the EED Gene Plymorphisms in Healthy Controls and Tumor and Lymph Node Stages of Colorectal Cancer Patients
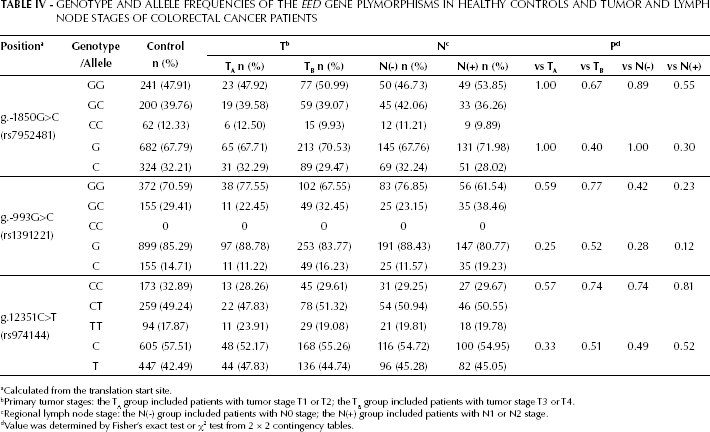
Calculated from the translation start site.
Primary tumor stages: the TA group included patients with tumor stage T1 or T2; the TB group included patients with tumor stage T3 or T4.
Regional lymph node stage: the N(-) group included patients with N0 stage; the N(+) group included patients with N1 or N2 stage.
Value was determined by Fisher's exact test or χ2 test from 2 × 2 contingency tables.
Next, we compared the haplotype frequencies of the g.-1850G>C, g.-993G>C, and g.12351C>T SNPs of EED between CRC patients and healthy controls. Out of 8 possible haplotypes only the frequencies of 4 major haplotypes (GGT, CGC, GCC, and GGC), which have been previously identified in healthy controls and UC patients (17), and of 1 minor haplotpye (CGT) were not significantly different between CRC patients and healthy controls (Tab. V). In addition, there was no significant difference in the haplotype frequencies of the EED SNPs between healthy controls and CC or RC patients, or between healthy controls and right or left CC (data not shown). These results suggest that the haplotypes of the EED polymorphisms are not associated with CRC susceptibility.
Haplotype Frequencies of the EED Gene Polymorphisms in Colorectal Cancer Patients and Healthy Controls
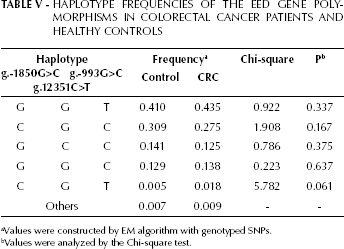
Values were constructed by EM algorithm with genotyped SNPs.
Values were analyzed by the Chi-square test.
Discussion
Multifactorial diseases such as CRC occur as a result of the interplay between a number of genetic and environmental factors. Unlike single-gene disorders, the genetic factors in CRC include many loci with relatively small effects, and only few, if any, are absolutely required for CRC occurrence. SNPs are the most common known form of human genetic variation, and may contribute to the increased susceptibility to cancer, including colorectal cancer (21). Recently, several genome-wide association studies reported an association between the variants of chromosome 8q24 and CRC (22–24). However, most of the underlying susceptibility loci for CRC remain unknown.
Sporadic and colitis-related CRCs are associated with inflammation and inflammatory diseases such as IBD (18). The severity of inflammation correlates with the risk of CRC in patients with IBD (6). We previously suggested that the g.-1850G>C polymorphism in the EED gene might be associated with the susceptibility to UC (17). Additionally, a recent report suggested that the PRC2/EED-EZH2 complex is overexpressed in breast cancer lymph node metastasis compared to primary tumors (19). However, there was no information on the relation between genetic variations of the EED gene and CRC. Therefore, the present study aimed at investigating whether polymorphisms in the EED gene are associated with susceptibility to CRC.
In our study, the genotype and allele frequencies of the EED SNPs in CRC patients were not significantly different from those in healthy controls (Tab. I). Since CC and RC have each been associated with different genetic features (25), CRC patients were divided into CC and RC patients, and their genotypes and allele frequencies for the EED SNPs were analyzed. The genotype and allelic frequencies of the EED SNPs in both CC and RC patients were also not significantly different from those in healthy controls. Only the allele frequency of g.-1850G>C in the RC patient group was significantly different from that in the control group (p=0.04).
Some studies suggest that right-sided cancers may have unique molecular features, which may be associated with rapid cancer development (26). However, our data do not show an association between the EED SNPs and the side of the CC lesions (Tab. III).
Identification of the factors regulating carcinogenesis and progression of CRC would contribute to reduce the occurrence of CRC, as well as to improve the clinical outcome for treatment of the disease. However, the genotype and allele frequencies of the EED SNPs in different primary tumor and lymph node stages of CRC were also not significantly different from those in healthy controls (Tab. IV). These results suggest that the genotype and allelic frequencies of the EED SNPs cannot be associated with susceptibility to CRC. However, we found that the genotype and allelic frequencies of the g.-993G>C SNP were significantly different between patients in the N(+) and N(-) group (p=0.02 and p=0.03, respectively). This result indicates a possible role of the EED SNP in the tumor aggressiveness of CRC. This is consistent with the finding by Yu et al (19) that PRC2/EED-EZH2 may play a pivotal role in the lymph-node metastatic process of breast cancer.
Similarly, the haplotype frequencies of the EED SNPs in healthy controls were not significantly different from those in CRC patients (Tab. V). Furthermore, there was no significant difference in the haplotype frequencies of the EED SNPs between healthy controls and CC or RC patients, or between healthy controls and right CC or left CC (data not shown). These results suggest that the haplotype of the EED SNPs may not be associated with CRC susceptibility.
In conclusion, the results of our study, with a power of 76.6%, suggest that EED may not be associated with the pathogenesis of colorectal cancer. However, our data show an association between the g.-993G>C polymorphism in the EED gene and lymph node metastasis of CRC.
