Abstract
Traditionally man has looked to nature to provide cures for diseases. This approach still exists today in the form of ‘bio-prospecting’ for therapeutically-active compounds in venoms. For example, the venoms of many reptiles offer a spectacular laboratory of bioactive molecules, including peptides and proteins. In the last 10–15 years, there have been a number of major proteomic and genomic research breakthroughs on lizard venoms. In this current review, the key findings from these proteomic and genomic studies will be critically discussed and suggestions will be offered for future focused investigations. It is our intention that this article will not only provide a comprehensive picture of the state of current knowledge of the components of lizard venoms, but also engender awareness in readers of the need to protect and conserve such uniquely precious natural resources for several reasons, including the potential benefit of humankind.
Introduction
The study of biologically-active molecules, including peptides and proteins, present in tissues of invertebrates and vertebrates, has been a core research focus in the life and health sciences for many years. Such study has produced many important leads in drug discovery.1,2 It is likely that such peptides and proteins represent an ancient form of intra- and intercellular communication. Scientists have designed peptide/protein phylogenetic trees by translating common amino acid sequences to ancestors of peptides/proteins found to be present in both invertebrates and vertebrates.3,4 For example, some of the mammalian gastroenteropancreatic (GEP) peptides and the vasoactive intestinal peptide (VIP) from humans are also present in the central and peripheral nervous system of lower vertebrates. 5 These findings indicate that the vertebrate hormones/active peptides may have originated in neural tissues before different vertebrate evolutionary lines developed. 3
The last review of lizard venom research was published in 1996 by Raufman. 4 Since then there have been several major research breakthroughs and reports on lizard venom components and evolution in this intervening period. These include 1) the development of a minimally invasive and reliable technique for acquiring lizard venoms, 6 2) the development of robust molecular techniques for constructing cDNA libraries to obtain proteomic and genomic data from lyophilised lizard venom secretions,7,8 3) the discovery of venom toxins in two additional lizard lineages (varanids and iguania), 9 4) the discovery of novel bradykinin-receptor inhibitory peptides (helokinestatins) in helodermatid (Heloderma suspectum and Heloderma horridum) venoms, and 5) the discovery of natriuretic peptide toxins in both helodermatid (H. suspectum and H. horridum) and a varanid (Varanus varius) venom.10,11 In addition, a recent study has also demonstrated that Komodo dragons (Varanus komodoensis) and the extinct giant Megalania (Varanus prisca) possess(ed) venomous bites. 12 In the present review, all the recent proteomic and genomic research on lizard venoms will be discussed and suggestions for future investigations will be offered.
Development of a Technique for Acquiring Lizard Venoms
Kwok and Ivanyi developed a minimally-invasive technique for venom extraction from lizards, 6 and genomic and proteomic studies on these venoms have been published by Chen et al and Kwok et al objectively supporting that this method of obtaining lizard venom worked extremely well.7,10,13 Kwok and Ivanyi's technique offered a rubber-covered object for the lizard to bite. After the lizard was safely restrained, 14 had opened its mouth and the oral cavity had been rinsed with water or saline, they presented it with a plastic-reinforced rubber squeeze bulb. As soon as the bulb contacted the lizard's mouth, it would voluntarily bite down on the bulb. Care was taken to ensure that the posterior teeth (toward the angle of the jaw) engaged the bulb. Then the lizard was held with its head at a 30°–45° angle (below horizontal) and its mouth was suspended above a glass or plastic vessel. 6 After a lizard bit the rubber bulb and compressed and released its jaws several times, drops of venom would be produced and the venom would drop into the vessel from along the gums of the lower jaw.
The advantages of this modified technique are that it is generally non-injurious to the lizard, reduces animal stress and the shedding of blood from breaking teeth that might be inadvertently collected using other methods. 15 It also maximizes the quantity of venom that can be safely collected. Overall, the technique appears safe and reliable for use in venom extraction from lizards, which is of great benefit when research requires investigations into the venoms of rare/protected species, and provides a robust source of valuable lizard venoms for biomedical researchers performing intensive proteomic and genomic research for potential drug discovery.
Development of Molecular Techniques for Constructing Lizard Venom cDNA Libraries
The emerging science of functional genomics integrates data derived from proteomics, genomics, and physiomics to provide a holistic view of the life process–-a necessary prerequisite for a global understanding. In the case of lizard venom proteomics, where many toxic proteins are of relatively high molecular mass, direct sequencing to obtain full primary structure is the method of choice. Many researchers resort to molecular cloning to this end using either degenerate primers designed for small sequence tags derived from the protein of interest or those designed for conserved nucleic acid domains of related molecules from other species. In 2002, Chen et al first reported the simple and rapid technique that unmasks venom protein mRNAs present in lyophilized lizard venom samples. 7 The traditional approach to isolation of mRNA from lyophilized venom samples fails to recover appropriate RT-PCR-compatible mRNA and, hence, its absence was assumed.5,7 However, Chen and his co-workers developed a novel methodology wherein they used magnetized oligo-dT DynaBeads for polyadenylated mRNA capture and circumvention of precipitation, recovering venom protein poly(A) mRNA, which is RT-PCR compatible and representative of all the studied classes of proteins from reptile venoms. 7 These findings imply that the apparent absence and/or lability of mRNA in complex biological matrices is not always real and pave the way for accelerated acquisition of molecular genetic data on venom toxins for scientific and potential therapeutic purposes without sacrificing animals. By using the principle of this robust technique, Kwok et al have further demonstrated that parallel proteome and venom gland transcriptome analyses can be performed on such samples. 8 They reported the presence of DNA that facilitates the more widely used applications of gene sequencing, such as molecular phylogenetics, in a non-invasive manner that does not require specimen sacrifice. From this “surrogate” tissue, they successfully acquired partial 12S and 16S rRNA gene sequences that were presented for illustration purposes. Thus, from a single sample of lizard venom, robust and complementary proteome, transcriptome and genome data can be generated for applications in diverse scientific disciplines.
Discovery of Additional Venomous Lizards
Until recently, it was believed that only one lineage of extant lizards, the helodermatids (represented by two species H. suspectum and H. horridum), had evolved a venom delivery system.9,16 In 2005, Fry and his international coworkers made a major scientific breakthrough in lizard venom history by investigating the evolution of venomous function in squamates. They obtained sequence data from five nuclear protein-coding genes representing major squamate lineages. Phylogenetic analyses showed that the closest relatives of venomous snakes are anguimorph (which includes helodermatids) and iguanian lizards (Fig. 1). 9 In a further investigation, they reported the presence of venom toxins (Table 1) in two additional lizard lineages (varanids and iguania), and through Magnetic Resonance Imaging (MRI), showed that all lineages possessing toxin-secreting oral glands form a clade.9,11,17
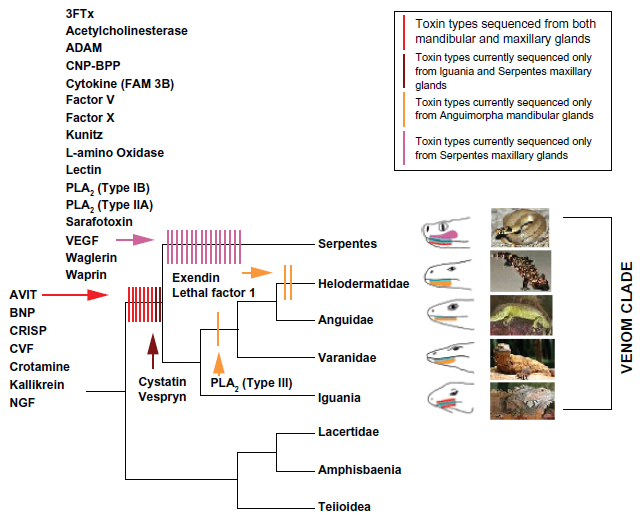
Source from Fry et al. 9
Lizard venom proteins and characterised activities.
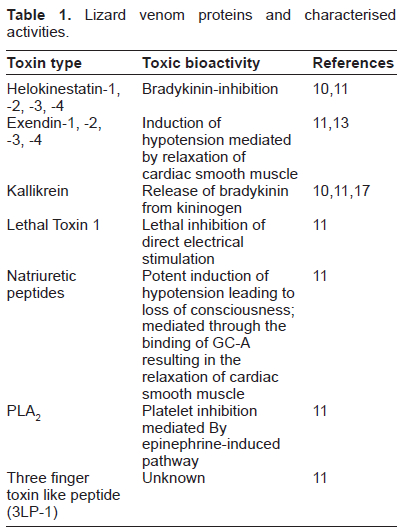
The venom toxins in varanids and iguania exhibit considerable sequence diversity, which is consistent with the birth-and-death mode of protein evolution that has given rise to a wide diversity of bioactivities in reptile venoms. These molecules represent a tremendous relatively unexplored resource, which is valuable to drug design and development, as well as aiding our understanding of reptile evolution.9,18
Following these discoveries Fry et al described the immediate effects of bites from these lizards as mild envenomations. Bites on human digits by a lace monitor (Varanus varius), a Komodo dragon (V. komodoensis), and a spotted tree monitor (V. scalaris), all produced similar effects: rapid swelling, localized disruption of blood clotting, and shooting pains up to the elbow, with some symptoms lasting for several hours. 9 Recently, Fry and his co-workers published further evidence demonstrating that the Komodo dragon possesses a venomous bite. 12 MRI scans of a preserved skull showed the presence of two venom glands in the lower jaw, and found that it secreted venom containing several different toxic proteins (Table 1). The known functions of these proteins include inhibition of blood clotting, lowering of blood pressure, muscle paralysis, and the induction of hypothermia, leading to shock and loss of consciousness in envenomated prey. 12 Moreover, Fry et al further analysed the structure and mechanics of the Komodo dragon's skull and jaw muscles. It was discovered that the bite force of the Komodo dragon is not particularly strong. 12 Rather than inflicting devastating bites, the jaws are best suited for clamping on to prey and resisting it from pulling away, giving time for the venom to penetrate the wound. 12 This means that less effort is required to subdue its victim, and allows larger animals to be preyed upon. As a result of these findings, the previous theory that bacteria were responsible for the deaths of Komodo dragon prey could be disputed. 12 In the same study, Fry et al also hypothesised that the Komodo dragon's extinct and even larger relative, Megalania (V. prisca), may have used the same method of prey capture, since this lizard also had a strong skull but relatively slender jawbones. 12 It was very closely related to the Komodo dragon and the lace monitor, both of which have now been proven to be venomous. Fry et al had previously shown that the capacity to produce venom evolved once in the common ancestor of snakes and lizard groups like the iguanas and monitors. 9 There is every reason to believe that Megalania used venom as well, which would make it the largest venomous animal to have ever lived.
Discovery of Helokinestatin and Natriuretic Peptides
The venom of helodermatid lizards has been studied as far back as 1967,19,20 but only recently has much of the biochemistry been elucidated. Over the last decade, the level of research into helodermatid venom components has intensified; particularly that directed towards the discovery of novel bioactive agents. The therapeutically useful exendin peptide toxin (marketed as Byetta™), isolated from helodermatid venoms for the treatment of diabetes, is the most remarkable example of lizard venom-drug discovery. 21
According to Russell's report in 1983, approximately 90%–95% of freeze-dried helodermatid venoms consist of proteinacious material made up of non-toxic proteins, toxins, enzymes and peptides. 22 Kwok and his co-workers believed that a large number of novel proteins/bioactive peptides within the venoms remained to be discovered. But, due to a lack of contemporary focus on the study of such venoms, this had not been determined. In 2007, Kwok et al reported a novel bradykinin-receptor inhibitory peptide from helodermatid venom named helokinestatin. 10 Helokinestatin represents an entirely novel structural class of helodermatid venom peptide displaying a unique bradykinin-receptor inhibitory activity. The discovery of helokinestatin in venoms of helodermatids represents an additional novel factor acting on the kinin/kininogen system–-at the level of selective bradykinin inhibition. 10 Recently, Fry et al investigated novel venom proteins produced by differential domain-expression strategies in helodermatid lizards. Multidisciplinary techniques were used to investigate the structure, function and evolutionary diversification of the unique venom system in these enigmatic lizards. 11 In their study, for the first time more protein types (e.g. natriuretic peptide) were reported from Heloderma venoms. 11 They also isolated additional isoforms of previously characterised Heloderma toxins belonging to the CRISP, exendin, helokinestatin, kallikrein and phospholipase A2 (PLA2) groups (Table 1). 11
The findings of Kwok's and Fry's studies, combined with the information available from the literature, 23 reveal how little we still know about the toxinology of the enigmatic Heloderma lizards. With respect to their venoms, we do not know enough of their composition, role of individual venom components in prey capture, or in vitro activities relative to subsequent physiological sequelae following envenomation.
Summary, Conclusions and Directions for Further Investigations
In summary, there have been a number of major scientific breakthroughs in lizard venom research over the last decade. To begin with, the development of a minimally invasive technique for acquiring samples of lizard venoms, along with robust molecular techniques, help provide a consistent and reliable source of valuable lyophilized lizard venoms for intensive proteomic and genomic research studies.6,7,8,10,13 Secondly, two additional lizard lineages, monitors–-including the Komodo dragon–-and the iguania, were confirmed as possessing toxin-secreting oral glands and venom toxins.9,12 As a result of these discoveries and reports, new insights into the evolution of the venom system in squamate reptiles have been provided and new avenues have been opened for biomedical investigations into the bioactive profiles of novel components establishing a path of new leads for drug design and development.
Thirdly, Raufman's 1996 review article reported five bioactive peptides that had been isolated from the venom of Heloderma lizards. 5 Additionally, four novel helokinstatin toxin variants have been discovered in helodermatid venoms.10,11 These peptides possessed a potent bradykinin receptor inhibitory activity and represent a prototype of a novel structural class of helodermatid venom peptide. 10 Thus, they represent additional and novel factors acting on the kinin/kininogen system–-at the level of selective bradykinin inhibition. In addition, more protein types (e.g. natriuretic peptide), previously reported only from snake venoms, were for the first time identified in Heloderma venoms.11,22,24 The natriuretic peptide transcript from the helodermatids also encoded tandem repeats of functional helokinestatin peptides, revealing that a single gene expresses both toxin types, with post-translational processing liberating the different peptides. 11 These newly identified venom toxins from lizards may herald a new era of research focus in proteomics and genomics of lizard venoms and their use for drug discovery.
This review shows how little we know about the fundamental evolution and function of these unique natural bioweapon systems and molecules, or the molecular evolution and function of the associated peptides/toxins. Therefore, it would be beneficial for future investigations in lizard venom research to focus on these proteomic and genomic techniques, which have cast a new light on our understanding of the history of the venom system among lizards. It is obvious that lizard venoms possess a high potential for drug discovery, as evidenced by the commercial success of the drug Byetta™, which is used to treat human diabetes and is derived from a helodermatid exendin-4 toxin. Results from venom research over the last decade suggest that further studies and sampling from well-known or newly-identified venomous lizards are likely to recover novel isoforms of types already identified, as well as discovering entirely new classes of bioactive peptides/toxins. These peptides/ toxins have great potential for use against a wide variety of pharmacological targets in humans, such as the treatment of pain, diabetes, multiple sclerosis and cardiovascular diseases, making them an invaluable source of ligands for studying the properties of these targets in different experimental paradigms for drug development.
The foremost clinical disadvantage to venom peptide/protein drugs such as Byetta™ is that they cannot ordinarily be administered via the oral route because they are likely to be either degraded by peptidase enzymes in the gastro-intestinal tract, or be too large to be absorbed. As a consequence, these drugs are typically delivered via the parenteral route, which makes the therapy intrusive for the patient and increases the costs for the healthcare provider. The intravenously administered peptidomimetic glycoprotein IIb/IIIa inhibitor, tirofiban (Aggrastat™), which was based upon compounds derived from Malayan pit viper (Calloselasma rhodostoma) venom is a fine case in point. 25 The transdermal, nasal, buccal/sublingual and pulmonary routes have all shown potential as means of peptide/protein drug delivery. Although Exubera™, Pfizer's insulin inhalation device, was launched and subsequently withdrawn from the market, there are several promising transdermal peptide/protein delivery systems which have recently been launched or are currently undergoing clinical trials. 26 As such, it seems that the search for potential lead compounds from lizard venom will need to proceed in tandem with research into formulation development to ensure that pharmacological discoveries are translated into commercially successful therapies.
We hope this review stimulates further interest into this dynamic field and we would also like to emphasize that, in the last 10 years, many reports suggest that many reptiles are experiencing dramatic range reductions and are threatened with extinction.27–30 Loss of biodiversity will result in the loss of important proteomic and genomic information that these animals contain, along with potentially unknown and untapped sources of new pharmacologically active agents. Therefore, in addition to stimulating further research in reptile venoms, we hope that this article serves to raise people's awareness regarding the value of these animals, and fosters a sense of responsibility to protect and conserve these precious natural resources.
Disclosures
This manuscript has been read and approved by all authors. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The authors and peer reviewers of this paper report no conflicts of interest. The authors confirm that they have permission to reproduce any copyrighted material.
Footnotes
Acknowledgments
We thank Dr Bryan G. Fry for helpful suggestions and critical reading of the manuscript.
