Abstract
This research intended to analyze the expression pattern of proteins in roots of the Brazilian soybean cultivar Conquista when inoculated with Bradyrhizobium japonicum CPAC 15, a strain broadly used in commercial inoculants in Brazil. At ten days after bacterial inoculation, whole-cell proteins were extracted from roots and separated by 2-D gel electrophoresis. Comparative analysis revealed significant changes in the intensity of 37 spots due to the inoculation (17 up-regulated and 20 down-regulated proteins), identified by MALDI-TOF/TOF-TOF. Identified proteins were associated with COG functional categories of information storage and processing, cellular processes and signaling, metabolism, and also in the “poorly characterized” and “not in COG” categories. Among the up-regulated proteins, we identified sucrose synthase (nodulin-100), β-tubulin, rubisco activase, glutathione-S-transferase, a putative heat-shock 70-kDa protein, pyridine nucleotide-disulphideoxidoreductase and a putative transposase. Proteomic analysis allowed for the identification of some putative symbiotic functions and confirmed the main biological processes triggered in the nitrogen-fixing symbiosis with soybean.
Introduction
The symbiotic associations of soybean [Glycine max (L.) Merrill] with bacteria belonging to the species Bradyrhizobium japonicum, B. diazoefficiens and B. elkanii have global economic and social importance. Brazil is an outstanding example, as biological nitrogen fixation (BNF) by soybean crops represents a key process in agricultural production systems, resulting in estimated savings of about US$15 billion in N-fertilizers per season. 1 The establishment of the symbiosis begins with the exchange of molecular signals between the bacterium and the host plant, involving a succession of complex processes which trigger profound changes in both symbionts. Once the plant recognizes the specific Nod signal produced by the rhizobia, the plant follows a pre-determined developmental pathway to form the nodule. Nodule formation is a highly ordered process that involves the de-differentiation of root cortical cells and their subsequent division to form the mature organ.2–4
Proteomic analysis, which focuses on investigating the cumulative changes and modifications of proteins, helps to acquire a more comprehensive understanding of the responses occurring in host plants under symbiotic conditions. 5 Proteomic studies of soybean bradyrhizobia have been reported, including studies with B. japonicum CPAC 15 (= SEMIA 5079),6,7 a strain belonging to the same serogroup as USDA 123 and broadly used in Brazilian commercial inoculants since 1992, due to symbiotic efficiency and competitiveness characteristics. 1 , 8 , 9 However, few genomic and proteomic studies have been reported with nodulated soybean, especially with Brazilian genotypes. Recently, Barros de Carvalho and collaborators 4 reported a transcriptional analysis of genes involved in nodulation of soybean cv. Conquista, when inoculated with strain CPAC 15, revealed a variety of transcripts related to primary metabolism, cell-wall modifications, and an antioxidant defense system. Complementing this study, we are now analyzing differential protein expression patterns in the same symbiosis of cultivar Conquista with strain CPAC 15.
Material and Methods
Plant Material
Soybean seeds (cultivar Conquista = MG/BR46) were surface-sterilized 10 and germinated on absorbent paper moistened with sterile distilled water at 22 ± 2°C (in the dark) for three days. Seedlings were transferred to sterile plastic bags containing 200 mL of N-free nutrient solution. 11
Inoculum Preparation and Plant Inoculation
B. japonicum strain CPAC 15 (= SEMIA 5079) was grown until the exponential phase of growth in yeast mannitol broth (YMB). 10 The cells were centrifuged and washed with saline solution (NaCl 0.85%). Aliquots of washed cell suspension were counted in YMB medium, indicating a concentration of 2.27 X 107 cells mL-1. For the inoculated treatment, 1 mL of the inoculum was applied at the base of each radicle of 3-day-old seedlings. The experiment had a fully randomized design with three replicates, each consisting of 20 plants per treatment. Treatments consisted of: soybean roots inoculated with strain CPAC 15 and non-inoculated soybean.
Plants were grown under greenhouse conditions with a 12-h day/night period and mean temperature of 25–28°C/15–18°C (day/night) for ten days. Roots of 13-day-old soybean were then separated from shoots, immediately frozen in liquid nitrogen and stored at–-80°C. Effectiveness of inoculation was proven by the inspection of abundant nodule primordia at the harvest. As at this stage only nodule primordia were present, the analyses were performed on whole roots. To confirm that inoculation was successful, we left several plants to grow until flowering stage, when we confirmed abundant nodulation and high production of biomass and N content in the shoots.
Proteomics Analysis
Whole-cell proteins of soybean roots were extracted from both the inoculated and control treatments following the simplified method described by Rodrigues et al. 12 Total-protein extract was solubilized in isoelectric focusing (IEF) buffer, quantified by the method described by Bradford 13 and mixed with DeStreak buffer (GE Healthcare) at a final concentration of 350 μg, which was employed to rehydrate IPG strips (linear pH 4–7, 13 cm, GEBiosciences). 2-D assays were performed, as described by Rodrigues et al, 12 in triplicate and the gels were analyzed by Image Master 2D Platinum v. 5.0 (GE Healthcare), after being stained with Coomassie Blue PhastGel™ R-350 (GE Healthcare). Well defined spots, present in all three gels, were selected, excised, and processed as described before. 14 Digestion was performed with trypsin (Gold Mass Spectrometry Grade, Promega) at 37°C overnight.
Mass spectra were acquired in a MALDI-TOF-TOF Autoflex Spectrometer (Bruker Daltonics), which was operated in the reflector mode for MALDI-TOF peptide mass fingerprint (PMF) and in the “LIFT” mode for MALDI-TOF-TOF in the fully manual mode, using Flex Control software v. 2.2 and processed using Flex Analysis v. 3.0 (Bruker Daltonics). The PMFs and MS/MS ion spectra generated were searched against the public database NCBInr (National Center for Biotechnology Information non-redundant)/ Viridiplantae, using the Mascot software v. 2.3 (Matrix Science) as previously described. 6 Identifications, available at PRIDE (http://ebi.ac.uk/pride) with the experiment accession number 14817, were validated only when the MOWSE (Molecular Weight Search) score was significant, and both decoy score and false discovery rates were considered.
Protein Characterization
A set of bioinformatics tools was used for enhanced characterization of identified proteins. The proteins were fit into COG (Clusters of Orthologous Groups) categories according to their functional inference, using the COGnitor program (http://www.ncbi.nih.gov/COG). 15 Software packages PSORT-B 16 and PSLpred 17 were used for prediction of subcellular localization.
Results and Discussion
Thirty-seven differentially expressed spots were analyzed by MALDI-TOF/TOF-TOF, resulting in the identification of 37 proteins that are highlighted by their numbers in Figure 1 and listed in Supplementary Table S1.

2-D profile of soybean root extract indicating the differentially expressed proteins after inoculation with Bradyrhizobium japonicum strain CPAC 15. The molecular weight of protein standards is indicated on the left.
Seventeen root proteins were up-regulated (increased expression) and twenty were down-regulated (decreased expression), when compared to the control, after inoculation of soybean Conquista with strain CPAC 15 (Fig. 2).
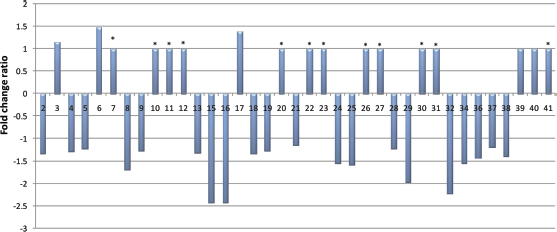
Fold change ratio of differentially expressed proteins of soybean roots, after inoculation with Bradyrhizobium japonicum strain CPAC 15. See Supplementary Table S1 for more details.
Proteins were distributed into 13 categories according to the functional classification in COG, belonging to four functional groups: information storage and processing (A, L), cellular processes and signaling (M, O, U, Z), metabolism function (C, E, F, G, H), and poorly characterized proteins. Ten proteins were classified as hypothetical/conserved hypothetical and another ten did not fit into any of the categories, being assigned as “not in COG” (Fig. 3).
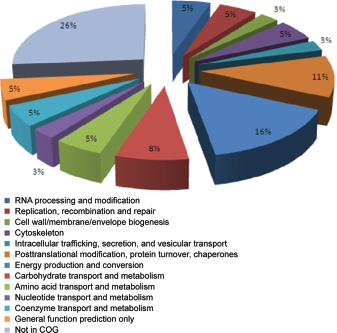
Distribution in COG categories of proteins of soybean roots after inoculation with Bradyrhizobium japonicum strain CPAC 15.
The most representative category was the “metabolic function,” with 37% of the identified proteins, 43% of which were related to energy production and conversion. Proteins related to cellular processes and signaling comprised 21%, followed by the information storage and processing category with 11%.
Symbiotic nitrogen fixation is an energy-demanding process, and the supply of sucrose to the nodule may limit fixation. In addition, nodule organogenesis requires imported sucrose for cellulose biosynthesis. 18 The sucrose synthase (nodulin-100), up-regulated in our study, contributes significantly to the development of the cell wall, among other known functions in nodulation. 19 Its activity increases rapidly during nodule development and declines during senescence.18,20 In addition, it is well known that several proteins are involved in plant-cell-wall penetration and cytoskeletal reorganization, given the need for structural modification of the root during the infection process by rhizobia.4,21
Rubisco activase was another up-regulated protein (Supplementary Table S1). This finding is interesting since this enzyme normally accumulates in greening or photosynthetic tissues expressing rubisco. 22 Nevertheless, rubisco activase is a member of a sequence superset of the AAA+ family, which are ATPases associated with diverse cellular activities 23 and include ATP-dependent proteases, membrane fusion, DNA processing, and microtubule severing and trafficking. 24 Some of the AAA+ proteins can also exhibit a classic chaperone activity in preventing the aggregation of denatured proteins and, in some cases, refolding them.24,25
We also identified proteins with potential antioxidant properties. Glutathione-S-transferase and pyridine nucleotide-disulphideoxidoreductase were up-regulated in inoculated plants; in contrast, glutathione reductase and mono-dehydroascorbatereductase were down-regulated. Reactive oxygen species (ROS) are produced at high rates when plants are exposed to abiotic, biotic, or xenobiotic stress, and the main sources of ROS in plants, under physiological conditions, are respiration, photosynthesis, and N2 fixation. 26 In this context, high levels of antioxidants can increase the rates of nitrogen fixation by up to 4-fold in planta as well as in vitro reconstitution systems containing leghemoglobin and bacteroids.27,2 8
A putative heat-shock 70 kDa protein, identified in our study, was up-regulated in inoculated plants. The 70 kDa heat-shock proteins are a group of ubiquitous, highly conserved molecular chaperones that have been implicated in a variety of processes ranging from DNA replication to protein folding and transport. 29 Although 70 kDa heat-shock proteins were originally identified on the basis of their stress inducibility, it is now understood that constitutively expressed 70 kDa heat-shock proteins also perform critical roles in normal metabolism. 30
Another down-regulated enzyme was the peroxisomalbetaine-aldehyde dehydrogenase (BADH). The most commonly studied aspect of BADH in plants is its role in tolerance of abiotic stresses, including salinity, drought, and high temperatures. Moreover, recent research has identified the potential for BADH as an antibiotic-free marker for selection of transgenic plants. 31
A probable cysteine synthase–-a primary enzyme of sulphur metabolism with significance to stress response–-was down-regulated. Plant thiols are apparently involved in responses to almost all stress factors, and their accumulation, redox status, and regulation are crucial to plant-stress tolerance. Minor changes in gene expression or redox state of thiols may define the difference between tolerant and susceptible genotypes or species. 32
We also identified an up-regulated putative transposase. By the criterion of inheritance instability, transposable elements have been described in at least 35mono- and dicotyledonous plant species. 33
A transcriptomic study performed in conditions similar to those of our study–-including the same cultivar and strain–-resulted in 3,210 differentially expressed transcripts. 4 Two proteomic spots were used to validate the transcriptional data, represented by a putative glutathione-S-transferase and a sucrose synthase. 4 In accordance with this transcriptional study, our study demonstrated intense metabolic activity during the nodulation process. Amongst the major processes, we highlighted the metabolic pathways of primary metabolism, cell-wall modification, and antioxidant-defense systems. Barros de Carvalho et al 4 found various nodulins (nodulin-21, nodulin-22, nodulin-26 and nodulin-36), and we now report nodulin-100.
It is important to note that data arising from genome and transcriptome studies are not always fully exploitable, since some sequences do not correspond to an assigned function, and direct information is not available about translation or co-and post-translational events of deduced gene products. 34
In our study, proteomics allowed the identification of some putative symbiotic functions and also confirmed the main biological processes triggered in the Brazilian soybean cultivar Conquista inoculated with B. japonicum strain CPAC 15, contributing to an understanding of critical events at the cellular level.
Concluding Remarks
By using proteomic tools, we evaluated the protein-expression pattern in roots of Brazilian soybean cultivar Conquista inoculated with B. japonicum CPAC 15, a strain broadly used in commercial inoculants in Brazil. The analyses allowed the identification of some putative symbiotic functions and also confirmed main biological processes triggered in the development of the nitrogen-fixing symbiosis in soybean.
Author Contributions
Conceived of and designed the experiments: EPR, JSSB and MH. Analyzed the data: ART, EPR, JSSB and DFG. Wrote the first draft of the manuscript: ART. Contributed to the writing of the manuscript: ART, EPR, JSSB, DFG, MH. Agreed with manuscript results and conclusions: ART, EPR, JSSB, DFG, MH. Jointly developed the structure and arguments for the paper: ART, EPR, JSSB, DFG, MH. Made critical revisions and approved final version: MH. All authors reviewed and approved of the final manuscript. ART and EPR contributed equally to the manuscript.
Disclosures and Ethics
As a requirement of publication the authors have provided signed confirmation of their compliance with ethical and legal obligations including but not limited to compliance with ICMJE authorship and competing interests guidelines, that the article is neither under consideration for publication nor published elsewhere, of their compliance with legal and ethical guidelines concerning human and animal research participants (if applicable), and that permission has been obtained for reproduction of any copyrighted material. This article was subject to blind, independent, expert peer review. The reviewers reported no competing interests.
Supplementary Data
Footnotes
Acknowledgements
The authors would like to thank Dr. L. Huergo, Dr. E.M. Souzaand Dr. F.O. Pedrosa for their help in the protein analyses, and Dr. Allan R. J. Eaglesham for suggestions on the manuscript. Approved for publication by the Editorial Board of Embrapa Soja as manuscript 347/2013.
