Abstract
Sputum of cystic fibrosis (CF) patients is a nutrient-rich environment. Higher amino acid content of CF sputum compared to normal sputum plays a major role in the CF-specific phenotype of P. aeruginosa. Presence of amino acids in the sputum-like environment influenced P. aeruginosa quorum-sensing activity and the formation of an unknown exopolysaccharide in the biofilm. Lipopolysaccharides isolated from P. aeruginosa grown in the presence of amino acids enhanced the release of cytokine IL-8 by human kidney and lung epithelial cells. The results of this study provide additional evidence on the role of amino acids towards adaptation of P. aeruginosa to the CF lung environment.
Introduction
The opportunistic pathogen Pseudomonas aeruginosa increases mortality rate of cystic fibrosis (CF) patients by chronically colonizing the lung throughout their lifespan. 1 With its huge genetic repertoire this organism modulates adaptive behaviour in response to local conditions in biotic and abiotic niches. The major macromolecular constituents of normal lung mucus are the mucin glycoproteins, which are large, heavily glycosylated proteins with tandemly repeating sequences of amino acids rich in serine and threonine, the linkage sites for large carbohydrate structures. 2 In contrast to normal individuals, sputum production is enhanced in chronic inflammatory airway diseases such as asthma, chronic bronchitis, and CF. In disease conditions, though sputum has altered macromolecular composition and biophysical properties, the common features are failure of muco-ciliary clearance, resulting in airway obstruction, and failure of innate immune properties. 3 However, in the CF lung airway sputum contains little intact mucin and has increased content of several macromolecules including DNA, filamentous actin, lipids, and proteoglycans. 4 Analysis of CF sputum medium indicated that amino acids and small peptides were major substrates for P. aeruginosa in respiratory secretions.5,6
P. aeruginosa undergoes phenotypic changes from the prototrophic wildtype to an auxotrophic phenotype during the course of chronic colonization in the CF lung. P. aeruginosa frequently exhibits auxotrophy in patients with CF, particularly in patients with severe underlying pulmonary disease. The amino acid content of CF sputum is high during infective exacerbations and correlated with the severity of pulmonary disease.7,8 In a previous study, an artificial sputum medium (ASM) was developed to mimic the CF lung habitat. P. aeruginosa and other bacteria associated with lung infections grew as tight microcolonies attached to the components of ASM. 9 Hitherto, the majority of biofilm models studied cells attached to a solid biotic or abiotic surface in a steady-state culture or under continuous flow.10,11 On the contrary, P. aeruginosa grows in the CF lung under anaerobic conditions as microcolonies, in which bacteria adhere to each other and build a biofilm with the help of self-produced exopolysaccharide (EPS) as well as the sputum components. 12 Amino acids in the environment were responsible for various CF-specific phenotypes of P. aeruginosa, such as variation in colony morphology, alterations in LPS structure and hyper- expression of the outer membrane protein, OprF. 9 In line with those findings, the present study provided additional information, especially, on the role of sputum components and oxygen in biofilm formation by P. aeruginosa. It was evident that amino acids were able to enhance quorum sensing by P. aeruginosa and also the release of cytokine IL8 by human cell lines.
Materials and Methods
Bacterial growth conditions
P. aeruginosa strains (PAO ATCC 15692; strain C, a prototype of clone C, is a CF isolate; C13 and C14, CF isolates; SG17M is a river water isolate from Germany) 13 were grown in tryptic soy broth (TSB) and M9 minimal medium (MM) (6 g/L Na2HPO4, 3 g/L KH2PO4, 1 g/L NH4Cl, 0.5 g/L NaCl, 0.241 g/L MgSO4 and 4 g/L glucose) at 37 °C. The M9 minimal medium was supplemented with casaminoacids (5 g/L) if necessary. Artificial sputum medium (ASM) contained 5 g mucin from pig stomach mucosa (NBS Biologicals), 4 g DNA (Fluka), 5.9 mg diethylene triamine pentaacetic acid (DTPA) (Sigma), 5 g NaCl, 2.2 g KCl, 5 ml egg yolk emulsion (phosphatidyl-choline as source of lecithin) (Oxoid) and 5 g amino acids per 1 l water (pH 7.0) modified after Ghani and Soothill. 14 Different variations of ASM were prepared by excluding one of the components of the medium. For a standard experiment, an inoculum equivalent to OD600 = 0.05 of an overnight culture of P. aeruginosa grown in tryptone soy broth (TSB) was added to 1 ml of ASM in 24-well cell-culture plates and incubated for 16 h at 37 °C with gentle shaking. ASM indicates complete artificial sputum medium throughout the paper.
Adherence to Surface
Adherence of P. aeruginosa to polystyrene wells was observed by staining with crystal violet in 24-well plates as described previously. 15
Treatment of Bacterial Aggregates
Bacterial aggregates/biofilm formed in ASM was treated with enzymes or chemicals to analyse their disruptive ability of EPS formed by P. aeruginosa. Alginate lyase in PBS (25 mg/ml), amylase in 10 mM ammonium acetate (pH 6.0) (5 μg/ml), cellulase (Trichoderma viride) in 50 mM ammonium acetate (pH 5.0) (0.1 mg/ml), DNase I in PBS (2 mg/ml), Proteinase K (2 mg/ml in buffer), Beta-mercaptoethanol in PBS (10 μl/ml), SDS in PBS (0.1%), Triton X 100 in PBS (0.1%), Tween 20 in PBS (0.1%) were used. All the chemicals and enzymes were purchased from Sigma-Aldrich (USA) unless otherwise mentioned.
Analysis of Quorum-sensing Molecules
P. aeruginosa strains were cultured in ASM with and without amino acids and the culture supernatants were extracted twice with an equal volume of dichloromethane (J. T. Baker) as described by Middleton et al. 16 The extracts were pooled together, evaporated to dryness in the SpeedVac (Eppendorf) and the dried material was reconstituted in 100 μl acetonitrile. 10 μl of 5-fold serially diluted extracts were added to 90 μl E. coli JM109 reporter strain harbouring plasmid pSB1075 (to detect 3O-C12-HSL), pSB401 (to detect 3O-C-HSL), or plasmid pSB536 (to detect C4-HSL) 17 and incubated overnight at 30 °C with shaking. Bioluminescence was measured as counts per second (CPS) using WALLAC Victor 2 1420 multilabel counter (Perkin Elmer, USA). HSL stands for homoserine lactones with different carbon (C) lengths and an oxo (O) group. Experiments were carried out in triplicates.
Analysis of Lipopolysaccharides
Isolation
Lipopolysaccharides (LPS) from P. aeruginosa were isolated as described by Hitchcock and Brown. 18 Briefly, overnight cultures of P. aeruginosa were adjusted to OD650 = 0.4 and centrifuged to obtain bacterial pellet. The pellet was resuspended in 50 μl lysis buffer containing 2% sodium dodecylsulfate, 4% β-mercaptoethanol, 10% glycerol, 0.002% bromophenol blue in 1 M Tris-Cl (pH 6.8) and heated at 100 °C for 10 min. 25 μg proteinase K per 10 μl of lysis buffer was added and heated at 60 °C for 1 h and stored at -70 °C until further use.
SDS-PAGE
To display LPS, 15% discontinuous SDS-PAGE was used as described. 19 To resolve core-lipid A bands the tricine SDS-PAGE method was used. 20 Briefly, polyacrylamide separating gels with 16.5% T (total acrylamide monomer concentration); 6% C (concentration of cross-linker) and stacking gels with 4% T; 3% C were made. The buffers included anode buffer (0.2 M Tris-HCl, pH 8.9), cathode buffer (0.1 M Tris-HCl, 0.1 M tricine, 0.1% SDS, pH 8.25) and the gel buffer (3.0 M Tris-HCl, 0.3% SDS, pH 8.45). Samples were electrophoresed at 30 V in the stacking gel and at 105 V in the separating gel. LPS was visualized by silver staining.
LPS for infection assay
Crude LPS prepared by tri-reagent method. 21 Briefly, 1–10 mg bacterial cells were suspended in 200 μl of Tri-reagent (Sigma) and incubated at room temperature for 15 min. After incubation 20 μl of chloroform per mg of cells was added to create a phase separation. The mixture was then vortexed vigorously and incubated at room temperature for an additional 10 min. The resulting mixture was centrifuged at 12,000 × g and the upper aqueous phase was collected in a clean centrifuge tube. The remaining organic phase was extracted twice with 100 μl distilled water and aqueous phases were pooled together. The combined aqueous phase was dried in a SpeedVac (Eppendorf) to get the crude LPS. It was dissolved in 500 μl of 0.375 M MgCl2 in 95% ethanol and stored overnight at -20 °C. The mixture was centrifuged at 12,000 × g for 15 min and the supernatant was removed. The pellet was then resuspended in 200 μl water and lyophilised. The lyophilised material was resuspended in a solution containing 0.1 mg/ml DNase I, 0.1 mg/ml RNase, 4 μM CaCl2, 4 μM MgCl2 and incubated overnight at 37 °C. 22 Then the mixture was treated with proteinase K (0.1 mg/ml) for 2 h at 56 °C. Pure LPS was recovered by ultracentrifugation and the resulting pellet was lyophilised, weighed, suspended in water and sonicated in an ultrasound bath until dissolved completely. The concentration and purity of LPS preparations was controlled on a silver-stained SDS-PAGE. Gels were stained with ethidium bromide or coomassie blue to check for contamination with nucleic acids or proteins respectively.
Preparation of lipid A
Crude LPS from the previous procedure was used as the starting material to isolate lipid A by mild acid hydrolysis. Crude LPS was dissolved in 500 μl 1% SDS in 10 mM sodium acetate (pH 4.5 with 4 M HCl) and placed in an ultrasound bath until the sample dissolved completely. It was then heated at 100 °C for 1 h and mixture was dried in a speed vac. SDS was removed by washing the mixture with 100 μl of distilled water and 500 μl acidified ethanol (100 μl 4 M HCl with 20 ml 95% ethanol) followed by centrifugation (2,000 × g for 10 min). The sample was then washed with 500 μl of 95% ethanol and centrifuged (2,000 × g for 10 min). The washing step was repeated twice and the sample was lyophilised to obtain a fluffy white solid lipid A.
IL-8 Measurement
The human epithelial cell lines T24 (ATCC HTB-4, human bladder carcinoma) and A549 (DSMZ ACC 107, human lung carcinoma) cells were grown in 24-well cell-culture plates in RPMI-1640 medium and Dulbecco's MEM respectively supplemented with 10% fetal bovine serum, 25 mM HEPES and 2 mM L-glutamine (Gibco BRL, Life Technologies, Denmark) at 37 °C in 5% CO2 to confluency. LPS was added at appropriate concentrations (0.001, 0.01, 0.1, 1.0, 10.0 and 20.0 μg/ml). Supernatants were collected 6 h post stimulation and the proinflammatory cytokine IL-8 was measured by ELISA (Diaclone). The experiments were carried out in triplicates. Data are presented as mean ± SEM. Statistical analysis was carried out using SPSS software (SPSS Inc.).
Results and Discussion
Pseudomonas aeruginosa colonizing the lung of CF patients is responsible for the decline in health and poor prognosis of these patients. Well-established infections in the form of chronic colonization are often very difficult to eradicate even by an intensive antimicrobial treatment. P. aeruginosa formed tight microcolonies in the artificial sputum medium that grew to form a macromolecular aggregate. This study is an extension of observations published by Sriramulu et al. 9 ASM was modified based on the fact that the average content of amino acids in the CF lung is 5 mg/ml. The CF lung environment is highly heterogenous that leads to patient to patient variation of conditions. To have a homogenous growth condition for the study, a cocktail of amino acids (casamino acids from Sigma) was used in all experiments. All P. aeruginosa strains 13 grown in ASM formed tight microcolonies (Figs. 1b, 2) irrespective of their origin, genotype and phenotype and without attaching to the wall of the well. But the environmental strain, SG17M, attached to the bottom of the well when grown in ASM without either amino acids, DNA, lecithin, or salts (Figs. 1b, 2). This feature might be due to the overexpression of cellular appendages such as flagella, pili, and fimbriae frequently exhibited by environmental isolates, which are not adapted to any unusual niche like the CF lung environment.23–25
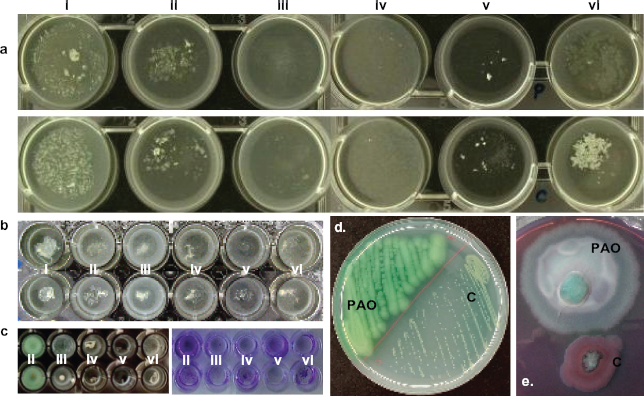
Biofilm formation by P. aeruginosa in (i) ASM and ASM without (ii) amino acids, (iii) DNA, (iv) lecithin source, (v) mucin or (vi) salts. (a) PAO (upper row) and strain C (lower row) in a 6-well plate, (b) PAO (upper row) and strain C (lower row) in a 24-well plate, (c) PAO (upper row) and strain C (lower row) in a 96-well plate, (d) colony morphology of PAO and strain C on TSB agar, (e) ASM-grown PAO and strain C on Congo red agar.
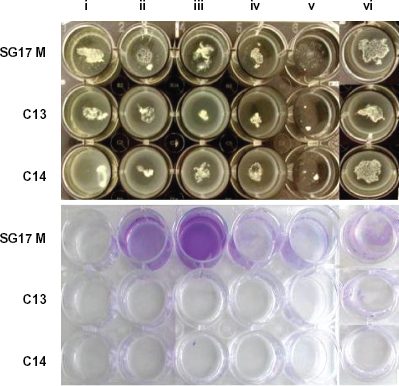
Top panel, P. aeruginosa strains SG17M, C13 and C14 grown in (i) ASM and ASM without (ii) amino acids, (iii) DNA, (iv) lecithin source, (v) mucin or (vi) salts. Bottom panel, wells stained with crystal violet to reveal biofilm attached to the walls.
Though P. aeruginosa formed aggregates in ASM and its variations, the extent of integrity of biofilm depended on the composition of the medium and the growth condition. The presence of oxygen played an important role in determining the integrity of microcolonies in the biofilm structure. 12 In this study, the extent of aeration was varied by using culture vessels of different dimensions. The surface to volume ratio was controlled by using 6-, 24-, and 96-well culture dishes with 9.6, 2.0 and 0.32 cm2 growth area containing 1.0, 1.0, and 0.2 ml of ASM respectively. As the surface area increased, P. aeruginosa formed loose or dispersed microcolonies (Fig. 1a). Microcolonies formed in 6 and 24-well plates did not attach to the bottom of the well as evidenced by crystal violet staining. When grown in 96-well plate P. aeruginosa microcolonies attached to the bottom of the well irrespective of the composition of ASM (Fig. 1c). In addition, the formation of pellicle at the air-liquid interphase was evident when the aeration was restricted by the smaller surface area of the culture vessel. Therefore, presence of microaerophilic to anaerobic condition in the CF lung environment might override the effect of sputum components on the morphology and integrity of biofilms.
From previous studies, it was evident that alginate is not the only EPS building material that contributed to the development of mature P. aeruginosa biofilms.26,27 To understand the nature of the EPS material responsible for tight microcolony formation, P. aeruginosa grown in ASM was plated on agar plate containing congo red indicator, which was widely used to study cellulose- and curli fimbriae-based multicellular morphology of Salmonella enterica serovar Typhimurium. 28 P. aeruginosa PAO formed big colony with subsequent spreading (Fig. 1e). Alginate production was induced extensively by this strain, which usually formed non-mucoid colonies on TSB agar. Whereas, P. aeruginosa strain C formed a brown, non-mucoid colony with a limited swarming behaviour on the Congo red agar surface. This observation indicated that there was a complete repression of alginate production in the CF lung-adapted strain C even under stimulating conditions. However, it was difficult to purify the EPS material from other components. Therefore, an attempt was made to disintegrate the P. aeruginosa PAO aggregate formed in complete ASM using enzymes and chemicals (Fig. 3) to understand the tentative composition of EPS. Enzymes like alginate lyase, amylase, cellulase, DNAse I, Proteinase K and denaturing agents like tween 20, triton × 100, SDS, beta-mercaptoethanol were used to disintegrate the biofilm. P. aeruginosa biofilm was stable under the experimental control conditions like, in water, saline or PBS even after 48 h incubation at 37 °C with shaking at 200 rpm. Amylase, a glycoside hydrolase that cleaves α-1, 4-glycosidic bonds, could not disintegrate the biofilm even after 48 h of treatment. Alginate lyase, which cleaves β-(1–4)-D-mannuronic bonds, had a meager effect on the aggregate by disintegrating the periphery after 24 h incubation and the rest of the large clump could not be disrupted by aspiration. The extent of disruption by this enzyme remained the same even after prolonged incubation of up to 48 h. Cellulase (0.1 mg/ml) had a profound effect on the bacterial aggregate and it was dissolved completely after 2 h of incubation. At lower concentrations (0.01, 0.002 and 0.001 mg/ml with less than overnight incubation) the aggregate was disrupted completely to form a homogenous solution containing planktonic cells. Treatment with DNase I resulted in a moderate disruption of the aggregate after 24 h incubation leaving a loose backbone structure probably containing EPS material and other ASM components. The backbone structure was stable even after 48 h incubation and could be disrupted only to some extent by aspiration. Tiny aggregates remained after aspiration (the solution was not homogenous as in the case of digestion with cellulase). Proteinase K had similar effect on the aggregate as DNase I, leaving only the backbone structure of the biofilm. SDS had a minimal effect on the biofilm by dissolving the periphery of the biofilm and leaving a translucent mass. The remaining structure could not be disrupted by aspiration. Tween, Triton-X and β-mercaptoethanol had the lowest disruptive effect on the aggregate. As a control, the aggregates could not be disrupted by any of the buffers used in the treatment. Taken together, complete disintegration of tight biofilm structure was achieved by DNase I and cellulase when compared to other agents used for the treatment.
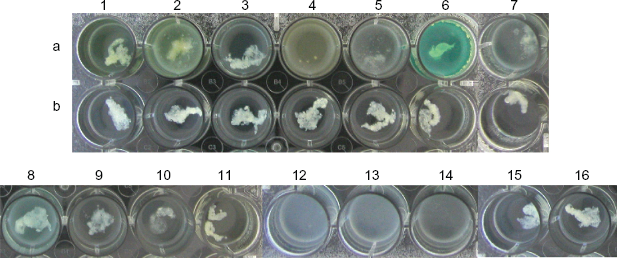

Treatment of ASM-grown P. aeruginosa PAO biofilm. Row (a) treated set; row (b) control set treated with PBS or other buffers. 1. control, 2. alginate lyase, 3. amylase, 4. cellulase, 5. DNase I, 6. Proteinase K (12.5 mg/ml), 7. Proteinase K (2.0 mg/ml), 8. Tween 20, 9. Triton X-100, 10. SDS, 11. β-mercaptoethanol, 12. cellulase (0.002 mg/ml), 13. cellulase (0.001 mg/ml), 14. cellulase (0.01 mg/ml), 15. water, 16. saline.
The DNA, released through the necrosis of lung epithelial cells, is one of the major components of CF sputum and it had been shown to contribute to the biofilm architecture. 29 In this study, the disruptive effect of DNase I on ASM-grown P. aeruginosa biofilm indicated that the DNA included in ASM was indeed incorporated in to the biofilm. As an additional source, DNA released from dead bacteria could not be excluded. Cellulase had the maximum disruptive capacity of loosening the P. aeruginosa biofilm in to planktonic cells. Cellulase hydrolyses 1, 4-beta-D-glycosidic linkages in cellulose, lichenin and cereal beta-D-glucans. 30 Surprisingly, the genome of P. aeruginosa PAO (www.pseudomonas.com) did not harbour genes involved in cellulose biosynthesis. Characterization of the EPS from the P. aeruginosa biofilm was unsuccessful due to background contamination with other components of the biofilm architecture. This study, however, is in conformation with previous findings that P. aeruginosa secretes a yet unknown/unnamed EPS to reinforce the biofilm.
Quorum-sensing
Establishment of chronic infection by P. aeruginosa requires an array of cell-associated and extracellular virulence factors. Several of these virulence factors have been demonstrated to be regulated by quorum sensing (QS).31–34 QS is a microbial cross-talk mechanism by which an individual bacterium produces small diffusible molecules that has a behavioural impact on surrounding cells or organisms.35–37 P. aeruginosa produces two such signal molecules called acyl homoserine lactones (AHLs), 3O-C12-HSL and C4-HSL, and a third molecule called Pseudomonas quinolone signal (PQS). These signals are encoded by the las, rhl, and pqs systems respectively. 33 The regulatory effect of these QS systems follows a hierarchal pattern from las to pqs to rhl in a top-down manner. In this study, the quantity of QS molecules was measured from the culture supernatants of P. aeruginosa PAO and a set of quorum-sensing mutants (δlasR, δrhlR, δpqsL and δpqsD) grown in ASM with and without amino acids (Fig. 4).
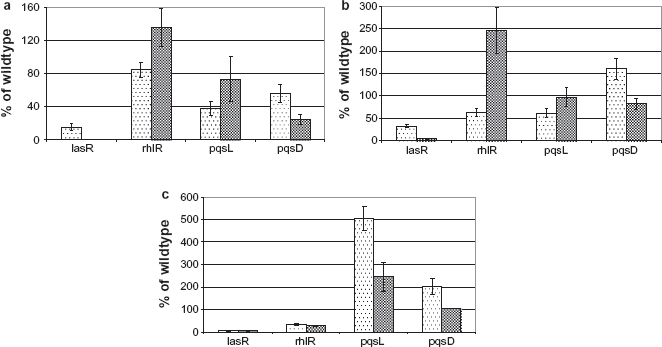
Relative quantification of quorum-sensing molecules produced by P. aeruginosa lasR, rhlR, pqsL and pqsD mutants grown in ASM (dotted bars) and ASM without amino acids (shaded bars), in comparison with the wildtype. (a) 3O-C12-HSL, (b) 3O-C6-HSL, (c) C4-HSL.
The las system, being the master regulator, completely abolished the production of C4-HSL by the lasR mutant. However, the presence of amino acids in the ASM resulted in the production of 3O-C12-HSL and 3O-C6-HSL up to 40% of the wildtype level. The rhlR and pqsL mutants produced lower levels of 3O-C12-HSL and 3O-C6-HSL in the presence of amino acids, the difference was wider in the production of 3O-C6-HSL by the rhlR mutant. Interestingly, in the pqsL mutant the production of C4-HSL was enhanced by 5- and 2.5-fold in the presence and absence of amino acids respectively. Mutation in pqsL was known to increase the levels of PQS by interrupting its degradation. 38 An increase in the production of C4-HSL by pqsL mutant indicates that the rhl system is also controlled by the pqs system with the latter being at an intermediate level in the QS hierarchy. 39 Mutation in pqsD gene involved in PQS biosynthesis showed reduced levels of 3O-C12-HSL compared to the wildtype. Presence of amino acids increased the level of 3O-C6-HSL and C4-HSL. Taken together, this study indicated that one or more amino acids might serve as a precursor for the biosynthesis of QS molecules by the corresponding QS system genes or by yet unknown genes functional through alternative pathway. The presence of amino acids in the CF lung environment enhances the release of QS molecules, which modulate virulence and adaptive behaviour of P. aeruginosa. Production and secretion of EPS is a QS-controlled phenotype.
IL-8 Response
Cytokines and polymorphonuclear leukocytes play a key role in immune mediated inflammation in progressive pulmonary damage due to CF disease. CF is a complex multisystem disorder caused by mutations in a membrane glycoprotein, the CF transmembrane conductance regulator, which serves as a Cl– channel. 40 One of the complications in CF is characterized by persistent infection with clonal populations of P. aeruginosa.41,42 The inflammatory response to such chronic infection is mediated primarily by cytokines such as tumor necrosis factor, interleukin-1 (IL-1), IL-6, and IL-8, which resembles the immune response elicited by LPS. 43 P. aeruginosa showed a modified LPS, particularly in lipid A anchor component, during growth in CF sputum-like environment. 9 In this study, an attempt was made to investigate the biological impact of such changes in the antigenicity of LPS by measuring the level of IL-8 released by human epithelial cells. Two cell lines were chosen for the purpose, the T24—a urinary bladder carcinoma cell line, and the A549—a carcinomic alveolar basal epithelial cell line. In CF research, while A549 cell line is widely used to study the host IL-8 response to different LPSs, the T24 cell line was included for the purpose of comparison. LPS and lipid A were successfully isolated from planktonic cells of bacteria. Isolation of LPS from bacterial biofilms has always been a challenge. In this study, LPS was isolated for cellular assays using established procedures (see Materials and methods). Compositional and structural analysis of LPS is a labour-intensive task and requires special expertise and enough material. Modification of lipid A component of LPS has been a hallmark of CF lung isolates (see above). Though well-established procedures were followed to isolate lipid A, it was not possible to obtain pure preparation suitable for structural analysis.
When challenged with LPS isolated from P. aeruginosa PAO grown in ASM with and without amino acids, TSB or MM, different levels of IL-8 release were measured from the cell culture supernatants (Fig. 5). LPS from P. aeruignosa PAO grown in TSB and MM showed no change in the level IL-8 response in both cell lines. LPS from ASM-grown P. aeruginosa induced the release of an elevated amount of IL-8 by both cell lines. T24 cells released at least 18-fold higher amount of IL-8 compared to A549 cells in response to LPS showing an overall difference in the immune response between cell lines (Figs. 5a and b). Interestingly, LPS from P. aeruginosa grown in ASM without amino acids induced lower levels of IL-8 in T24 cells compared to that from TSB or MM-grown cells. There was no significant difference (P < 0.05) in the level of IL-8 released by A549 cells in response to LPS from P. aeruginosa grown in ASM without amino acids, TSB and MM. These changes in proinflammatory immune response must be due to the result of specific structural changes in the LPS molecule isolated from the same strain grown under different conditions. Existence of a variety of penta- and hexa-acylated lipid A structures in the LPS of P. aeruginosa grown under different environmental conditions had been reported. 44 CF isolates of P. aeruginosa synthesized LPS with specific lipid A that contained palmitate and amino-arabinose. 45 Modification of lipid A was not only CF-specific but was also associated with resistance to cationic antimicrobial peptides and an increased inflammatory response.44–47 An attempt to isolate lipid A from ASM-grown P. aeruginosa PAO resulted in preparations that were heavily contaminated with the components of the medium. However, in this study, the observed increase in IL-8 levels in response to LPS from P. aeruginosa grown in ASM was indeed due to modification of lipid A molecule. Further study is necessary to find the specific modification in this molecule.
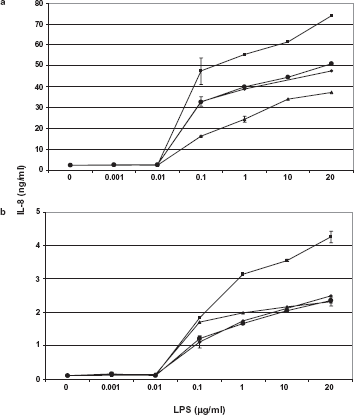
IL-8 response towards LPS isolated from P. aeruginosa PAO grown in TSB (♦), MM (•), ASM (▪) and ASM without amino acids (▴) by (a) T24 kidney epithelial cells and in (b) A549 lung epithelial cells.
Taken together, the above findings indicate that the availability of free amino acids in the CF lung environment might act as a driving force towards adaptive evolution of P. aeruginosa during the course of chronic infection.
Publish with Libertas Academica and every scientist working in your field can read your article
“I would like to say that this is the most author-friendly editing process I have experienced in over 150 publications. Thank you most sincerely.”
“The communication between your staff and me has been terrific. Whenever progress is made with the manuscript, I receive notice. Quite honestly, I've never had such complete communication with a journal.”
“LA is different, and hopefully represents a kind of scientific publication machinery that removes the hurdles from free flow of scientific thought.”
Available to your entire community free of charge
Fairly and quickly peer reviewed
Yours! You retain copyright
Footnotes
Acknowledgements
I would like to thank David D'Argenio and Steve P. Diggle for providing QS mutants. I thank G. S.
Stewart for providing reporter plasmid constructs to measure QS molecules.
This manuscript has been read and approved by the author. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The author and peer reviewers of this paper report no conflicts of interest. The author confirms that they have permission to reproduce any copyrighted material.
