Abstract
Killer cell immunoglobulin-like receptor (KIR) genes encode for activating and inhibitory surface receptors, which are correlated with the regulation of Natural Killer (NK) cell cytotoxic activity. Reduced NK cell cytotoxic activity has been consistently reported in Chronic Fatigue Syndrome/Myalgic Encephalomyelitis (CFS/ME) patients, and KIR haplotypes and allelic polymorphism remain to be investigated. The aim of this article was to conduct a pilot study to examine KIR genotypes, haplotypes, and allelic polymorphism in CFS/ME patients and nonfatigued controls (NFCs). Comparison of KIR and allelic polymorphism frequencies revealed no significant differences between 20 CFS/ME patients and 20 NFCs. A lower frequency of the telomeric A/B motif (P < 0.05) was observed in CFS/ME patients compared with NFCs. This pilot study is the first to report the differences in the frequency of KIR on the telomeric A/B motif in CFS/ME patients. Further studies with a larger CFS/ME cohort are required to validate these results.
Keywords
Introduction
Natural Killer (NK) cells are effector cells of the innate immune system, and following recognition of a potential target cell, NK cells mediate a response through cytotoxic activity to remove the target cells or cytokine production to direct an immune response. 1 Activation of NK cell cytotoxic activity is a tightly regulated process governed by the balance of signals received from surface receptors. 2 NK cells constitutively express a myriad of surface receptors, which can be structurally grouped into the immunoglobulin superfamily and the C-type lectin family.2,3 One major family of NK cell receptors includes the Killer cell immunoglobulin-like receptors (KIRs).2,3 Through KIR receptors, NK cells recognize target cells with reduced or absent expression of human leukocyte antigen, which may be the result of infection, malignant transformation, or cellular stress. 4 KIR engagement with specific human leukocyte antigen ligands transduces a cascade of signals to inhibit or activate NK cell cytotoxic activity. 2
KIRs expressed on NK cells are encoded by 17 KIRs, which are located on human chromosome 19q13.4 in the leukocyte receptor cluster. 5 Of the 17 KIRs characterized, nine genes encode inhibitory receptors (KIR3DL3, KIR2DL2, KIR2DL3, KIR2DL5B, KIR2DL1, KIR2DL4, KIR3DL1, KIR2DL5A, and KIR3DL1), six encode activating receptors (KIR2DS2, KIR2DS3/2DS5C, KIR3DS1, KIR2DS3/2DS5T, KIR2DS4, and KIR2DS1), and the remaining two are pseudogenes (KIR2DP1 and KIR3DP1) with unknown functions. 4 Within the leukocyte receptor cluster, the KIR locus is defined by conserved framework genes including KIR3DL3, 3DPl, 2DL4, and 3DL2, which also mark centromeric and telomeric regions. 6 KIRs in the centromeric and telomeric regions are genetically diverse due to variability in gene content and allelic polymorphisms.7,8 The combination of KIR and pseudo genes gives rise to a number of different genotypes, which according to the presence or absence of specific KIRs can be further classified as haplotypes A or B. 6 Haplotype A predominantly consists of inhibitory genes including KIR2DL1, KIR2DL3, KIR3DL1, and KIR3DL2 and the activating KIR2DS4. 5 KIR haplotypes that do not contain the exact copy of haplotype A genes are classified as haplotype B. 5 The predominance of inhibitory genes in haplotype A and activating genes in haplotype B suggests a distinct role of KIR haplotypes in governing effector functions of NK cells. 9 KIR association studies have suggested that haplotype A provides more effective immunity for the clearance of viral infections including hepatitis C and Ebola compared with haplotype B due to the regulation of NK cell activity.10–12 KIR haplotypes present on the centromeric or telomeric motifs are also known to influence NK cell function.5,6 For example, haplotype B on centromeric and telomeric motifs has been identified to provide protection against relapse in hematopoietic stem cell transplantation. 5 In kidney transplant patients, B haplotypes on the telomeric motif has been suggested to protect against cytomegalovirus infection.5,6
Variations in KIR gene content and allelic polymorphism have been identified to influence KIR surface expression and receptor ligation required to initiate NK cell cytotoxic activity and cytokine production.12–16 Reduced NK cell cytotoxic activity has consistently been reported in patients with Chronic Fatigue Syndrome/MyalgicEncephalomyelitis(CFS/ME).17–25 While one study has identified that CFS/ME patients have increased frequencies of KIR3DS1, additional levels of genetic diversity including KIR haplotypes, centromeric and telomeric haplotypes, and allelic polymorphism, which may contribute to reduced NK cell cytotoxic activity, remain to be investigated. 26 The aim of this pilot study was to investigate KIR genotypes, haplotypes, and allelic polymorphism in CFS/ME patients and nonfatigued controls (NFCs).
Materials and Methods
Study participants and inclusion criteria
CFS/ME patients and NFCs were recruited from a database at the National Centre for Neuroimmunology and Emerging Diseases, Menzies Health Institute Queensland, Australia. In the absence of a diagnostic test, the 1994 Fukuda definition was used to identify CFS/ME patients. 27 All participants completed an online questionnaire based on the Fukuda definition for fatigue and symptom presentation to determine suitability for study inclusion. Exclusion criteria included participants presenting with primary mood disorders, thyroid conditions, diabetes, epilepsy, psychosis, cardiac disorders, smoking, pregnant or breastfeeding, and immunological, inflammatory, or autoimmune diseases.
Compliance with ethical standards
Written informed consent was obtained from all participants. This study was conducted with the approval of the Griffith University Human Research Ethics Committee (MSC22/12/HREC) and in accordance with the ethical standards of the 1964 Declaration of Helsinki.
Blood collection
Thirty-five milliliters of peripheral blood was collected into ethylenediaminetetraacetic acid tubes from the antecubital vein of each participant. Blood samples were collected between the hours of 7:30–10:00 am to eliminate circadian variation and analyzed within four hours of collection. 28 Participant blood parameters including full blood counts of white and red blood cells, electrolytes, and erythrocyte sedimentation rate were assessed on all samples by Pathology Queensland.
NK cell isolation and DNA extraction
Peripheral blood mononuclear cells were isolated by density gradient centrifugation with Ficoll-Hypaque (GE Healthcare). From the peripheral blood mononuclear cells, NK cells were isolated by a negative selection kit according to the manufacturer's instructions (Miltenyi Biotec). Isolated NK cells were frozen in liquid nitrogen and stored for deoxyribonucleic acid (DNA) extraction at a later date. DNA from NK cells was extracted using the QIAamp DNA extraction kit (Qiagen) according to manufacturer's instructions, and the concentration and quality of each DNA extraction was assessed using the NanoDrop Spectrophotometer 1000 (NanoDrop Technologies). Prior to genetic typing, NK cell DNA was stored at −20 °C.
KIR gene content
KIR genotyping was performed using reagents and software at Scisco Genetics.6,29,30 Briefly, 14 locusspecific primer pairs were used for the initial polymerase chain reaction (PCR) amplification to detect KIR3DL3, 2DS, 2DL2, 2DL3, 2DL5B, 2DS3/DS5C, 2DP1, 2DLl, 3DP1, 2DL4, 3DL1, 3DS1, 2DL5A, 2DS3/2DS5T, 2DS1, 2DS4, and 3DL2. 29 PCR amplicons generated from each individual were pooled and treated with Exonuclease I and alkaline phosphatase. The amplicon targets were then combined with DNA linkers containing adaptor sequences, which served as primer-binding sites for dual-indexing barcode PCR to ensure unique identification of each sample. Following barcoding, the samples were pooled and multiplex sequencing was performed using the MiSeq platform (Illumina). The generated sequencing data were aligned to sequences obtained from the Immuno Polymorphism-KIR Database to determine KIR and allelic assignments for each participant. 31 Participants were also stratified according to the number of activating (1–6) and inhibitory (6–9) KIRs present.
KIR haplotypes
KIR haplotypes in CFS/ME patients and NFCs were identified according to the presence or absence of specific KIRs. Haplotype A was determined according to the presence of nine KIRs: 3DL3, 2DL3, 2DP1, 2DL1, 3DP1, 2DL4, 3DL1, 2DS4, and 3DL2. 6 Haplotype B was identified according to the absence of all haplotype A genes. 6 Participants presenting with only haplotype A genes were assigned as A/A genotype, homozygous participants for haplotype B were assigned as B/B, and heterozygous individuals containing haplotype A and B genes were assigned as A/B. 32
Centromeric and telomeric motif KIR haplotypes
The position of KIRs within the KIR locus can further define the centromeric and telomeric motifs as genotypes A/A, B/B, or A/B. 6 Haplotype A KIR on the centromeric motif includes KIR3DL3, 2DL3, 2DP1, 2DL1, and 3DP1, while 2DL4, 3DL1, 2DS4, and 3DL2 are found on the telomeric motif. Centromeric and telomeric motifs with only haplotype B genes were assigned B/B, and participants with a combination of haplotype A and B genes on both motifs were classified as A/B.
Statistical analysis
Statistical analysis of the data was performed on the Statistical Package for the Social Sciences (IBM Corp, Version 22). For routine blood parameters, Shapiro–Wilk test was used to test for Gaussian distribution. The independent Mann–Whitney test was used to identify any significant differences in blood parameters between CFS/ME patients and NFCs. Frequencies of KIRs, haplotypes, centromeric and telomeric haplotypes, and KIR alleles were compared between CFS/ME patients and NFCs using Fisher's test of association (for frequency counts less than five) and the chi-square test (for frequency counts greater than five). P-values of <0.05 were considered statistically significant.
Results
Participants, blood parameters, and NK cell purity
All participants were Caucasian and a total of 20 CFS/ME patients meeting the 1994 Fukuda definition (mean age [years] α standard error of the mean = 53.2 ± 2.26) and 20 NFCs (mean age [years] α standard error of the mean = 52.85 ± 1.70) were included in this study. No significant differences were observed when the ages, white and red blood cell parameters, electrolytes, C-reactive protein, and erythrocyte sedimentation rate were compared between CFS/ME patients and NFC participants (Supplementary Table 1). The mean purities of isolated CD56+CD3– NK cells for CFS/ME patients and NFCs were 98.0% and 98.9%, respectively.
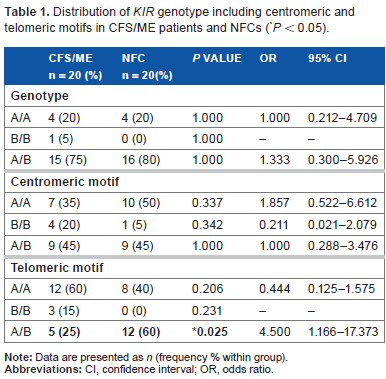
Distribution of KIR genotype including centromeric and telomeric motifs in CFS/ME patients and NFCs (*P < 0.05).
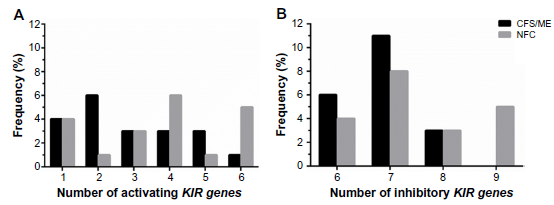
Frequency of activating and inhibitory KIRs present in CFS/ME patients
No significant differences were observed in the frequency of activating and inhibitory genes between CFS/ME patients and NFCs (Fig. 1). The frequency of two and five activating genes (A) and six and seven inhibitory genes (B) was higher in CFS/ME patients compared with NFCs, although this difference was not significant (B).
Frequency of activating (
No significant difference in KIR gene frequencies in CFS/ME patients
The frequency of individual activating and inhibitory KIRs was compared between CFS/ME patients and NFCs, and no significant differences were observed (Fig. 2). Although not significant, frequency of the activating KIR2DS2 was higher in CFS/ME patients when compared with the NFCs (A). For KIR2DS3/2DS5C, KIR3DS1, KIR2DS3/2DS5CT, KIR2DS1, and KIR2DS4, the frequency was lower in CFS/ME patients compared with the NFCs. Inhibitory KIRs, namely, KIR3DL3, KIR2DL1, KIR2DL4, and KIR3DL2, were present in all CFS/ME patients and NFCs (B). In CFS/ME patients, the frequency of KIR2DL2 and KIR2DL3 was higher compared with NFCs. The frequency of KIR2DL5B, KIR3DL1, and KIR2DL5A was lower in CFS/ME patients compared with NFCs, although these differences were not significant.
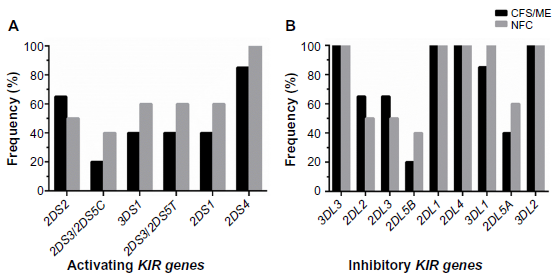
Frequency distribution of activating KIRs (
Telomeric A/B haplotype motif associated with CFS/ME patients
CFS/ME and NFC participants were classified as A/A, B/B, or A/B genotypes according to the presence or absence of specific KIRs, and no significant differences were observed (Table 1). A lower frequency of the A/B telomeric motif was observed in CFS/ME patients (P < 0.05) compared with NFCs.
Frequency distribution of KIR alleles in CFS/ME patients
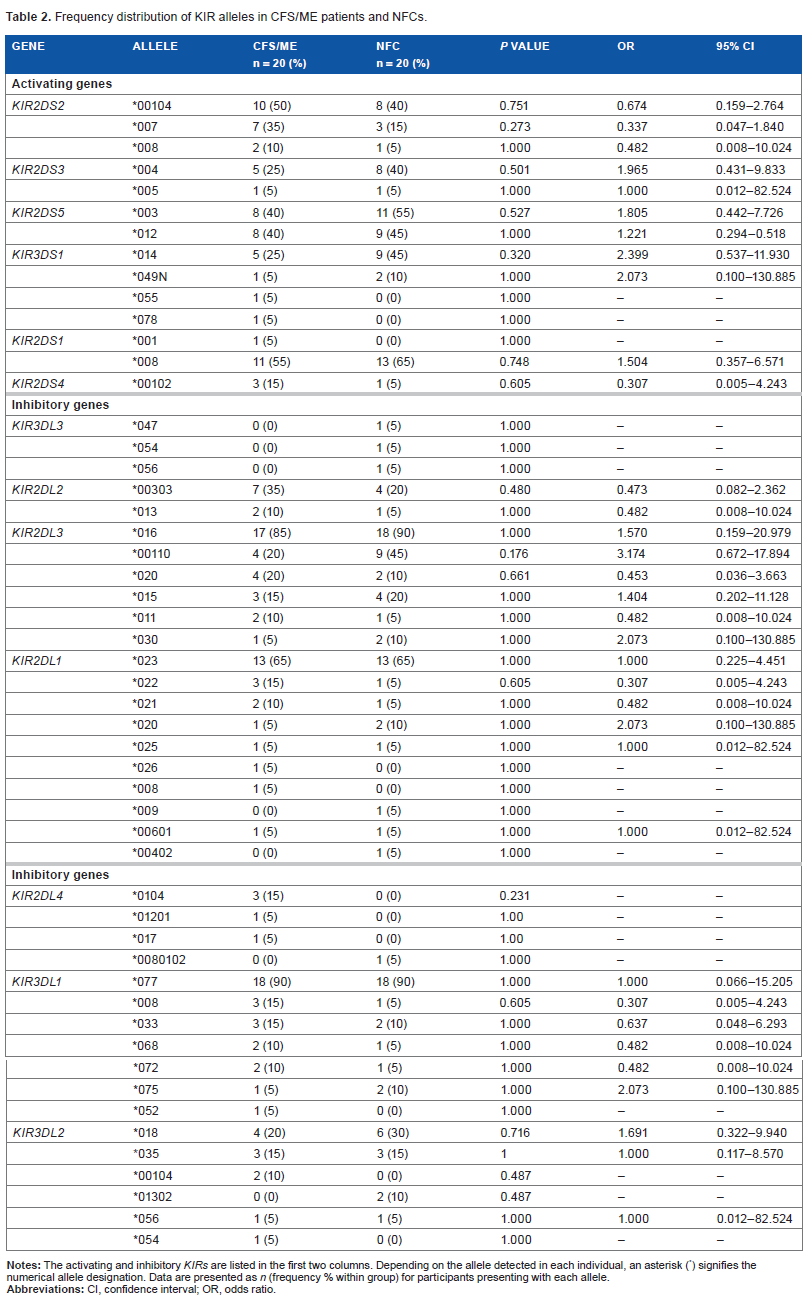
The presence of alleles associated with activating and inhibitory KIRs was compared between CFS/ME and NFC participants. While CFS/ME patients presented with increased frequencies of KIR2DS2*007 and lower frequencies of KIR3DS1*014 and KIR2DL3*00110 compared with NFCs, no significant differences were observed (Table 2).
Frequency distribution of KIR alleles in CFS/ME patients and NFCs.
Discussion
This pilot study is the first to genotype NK cell KIRs in an Australian population of CFS/ME and also the first to investigate KIR haplotype frequencies in CFS/ME patients. KIRs encode for activating and inhibitory surface receptors, which have previously been correlated with the regulation of NK cell cytotoxic activity.2,13,33–35 Reduced NK cell cytotoxic activity has been consistently reported in CFS/ME patients, and investigation of KIRs in the present study has revealed a significantly lower frequency of the telomeric A/B motif in CFS/ME.
The numbers of activating and inhibitory KIRs present were compared between CFS/ME patients and NFCs as gene quantity has been associated with NK cell activation.36,37 While differences were reported in the number of activating and inhibitory genes between CFS/ME patients and NFCs, statistical significance was not observed. Specific KIRs were also examined and no significant differences were reported between CFS/ME patients and NFCs. These findings contrast a previous association report of increased activating KIR3DS1 in CFS/ME patients. 26 An increased frequency of KIR3DS1 has been correlated with increased NK cell degranulation and production of interferon-gamma. 38 Previously, we have also reported increased degranulation and interferon-gamma production in NK cells from CFS/ME patients, which suggests that frequencies of KIR3DS1 may contribute to NK cell dysfunction in CFS/ME.18,26
Inherited diversity of KIR genotypes through the combination of maternal and paternal haplotypes on the centromeric and telomeric motifs has been associated with susceptibility or resistance to pathogen infection due to the regulation of NK cell activity. 39 Within the telomeric motif of the KIR locus, CFS/ME patients presented with a lower frequency of the A/B genotype compared with the NFCs. More than half of the CFS/ME cohort presented with homogenous A/A telomeric motif, which only contains one activating receptor, KIR2DS4. Due to the predominance of inhibitory KIRs in homogenous A/A genotypes, ligation of KIR2DL3, KIR2DL1, and KIR3DL1 confers strong inhibition through the immunoreceptor tyrosine-based inhibitory motifs.2,40 In contrast, inhibition of NK cells in haplotype B individuals are mediated by fewer ligands due to the absence of these inhibitory genes. 40 Differences in the presence of activating and inhibitory KIRs between A, B, and AB suggests that each haplotype may have different activation thresholds for NK cells, which maybe dysfunctional in CFS/ME patients.
In addition to the KIR content variation between the haplotypes, allelic polymorphism caused by insertions, deletions, substitutions, or single-nucleotide polymorphisms also contributes to the regulation of NK cytotoxic activity.14,16,41–43 The inhibitory function of NK cells is affected by substitutions of KIR3DL1 producing KIR3DL1*004, KIR3DL1*002, and KIR3DL1*007.14,32 KIR3DL1*004 results in the production of a misfolded protein, which is retained in the endoplasmic reticulum, while KIR3DL1*002 transduces a stronger inhibitory response than KIR3DL1*007 due to conformational changes in the extracellular region of the receptor. 16 As allelic polymorphisms have been associated with changes in the levels of KIR surface expression and strength of signals integrated due to ligand affinity, KIR alleles were investigated in CFS/ME patients and no significant differences were observed.
Conclusions
This pilot study is the first to report differences in the frequency of KIR on the telomeric A/B motif in CFS/ME patients. As the activity of NK cells is governed by the balance between activating and inhibitory signals, differences in the gene content profile of KIR haplotypes may create different activation thresholds for NK cells.39,40 In CFS/ME patients, further investigations are required to determine if lower frequencies of A/B on the telomeric motif may contribute to dysfunctional regulation of NK cell cytotoxic activity. It is also paramount for future studies to include a larger sample size to ensure that there is enough statistical power to identify the differences between CFS/ME patients and NFCs. Future studies into NK cell KIRs have the potential to identify if genetic predispositions may contribute to reduced NK cell cytotoxic activity in CFS/ME patients.
Author Contributions
Performed all the experimental protocols for NK cell isolation and DNA extraction from NK cells, analyzed the data, and wrote the article: TKH. Helped design the study, analyzed the data, and drafted the article: EWB. Conceived the study, sought ethics approval, provided the CFS/ME, NFC cohorts from the National Centre for Neuroimmunology and Emerging Diseases database, critically revised the intellectual content and interpretation of data analysis, and drafted the article: DRS, SMM-G. All the authors read and approved the final manuscript.
Footnotes
Acknowledgments
The authors thank Scisco Genetics for performing the KIR content haplotyping and Stephen Rudd, Michael Thang, and Anne Bernard at the Queensland Facility for Advanced Bioinformatics for completing the statistical analysis of the KIR data. We also acknowledge the National Centre for Neuroimmunology and Emerging Diseases, Stafford Fox Medical Research Foundation Grant, Mason Foundation, Queensland Smart State Co-investment Futures Fund, and Edward P. Evans Foundation.
