Abstract
To isolate specific genomic regions that retain their molecular interactions, allowing direct identification of chromatin-bound molecules, we developed two locus-specific chromatin immunoprecipitation (locus-specific ChIP) technologies, insertional ChIP (iChIP) and engineered DNA-binding molecule-mediated ChIP (enChIP) using the clustered regularly interspaced short palindromic repeats (CRISPR) system or transcription activator-like (TAL) proteins. Essentially, a locus-specific ChIP consists of locus-tagging and affinity purification and can be combined with downstream analyses to identify molecules associated with the target genomic regions. In this review, we discuss the applications of locus-specific ChIP to analyze the genome functions, including transcription and epigenetic regulation.
Keywords
Introduction
Biological activities involving the functions of genomic DNA play vital roles in all aspects of living organisms. These functions include transcription, epigenetic regulation, genomic imprinting, dosage compensation such as X-chromosome inactivation, and others.1–8 To understand the molecular mechanisms underlying the genome functions, it is essential to identify the molecules associated with genomic regions of interest. Recently developed techniques have enabled us to detect molecular interactions on genomic regions. For example, chromosome conformation capture (3C) and its derivatives9,10 have been used to detect interactions between genomic regions. In addition, proteomics of isolated chromatin (PICh) has been used to identify proteins associated with multicopy loci such as telomeres and centromeres. 11
In this review, we discuss two locus-specific chromatin immunoprecipitation (locus-specific ChIP) technologies, insertional ChIP (iChIP) and engineered DNA-binding molecule-mediated ChIP (enChIP), which we developed for locus-specific biochemical analysis of genome functions such as transcription and epigenetic regulation.
iChIP
Scheme of iChIP
Tagging of loci with the insertion of recognition elements of an exogenous DNA-binding protein has been widely used in live imaging.
12
Such locus-tagging was utilized for biochemical purification of specific genomic regions in yeast by Kornberg's group.
13
His group used the
Using the locus-tagging strategy, we developed iChIP technology,14–19 which overcomes many of the drawbacks of previous methods. The original scheme of iChIP is as follows (Fig. 1):
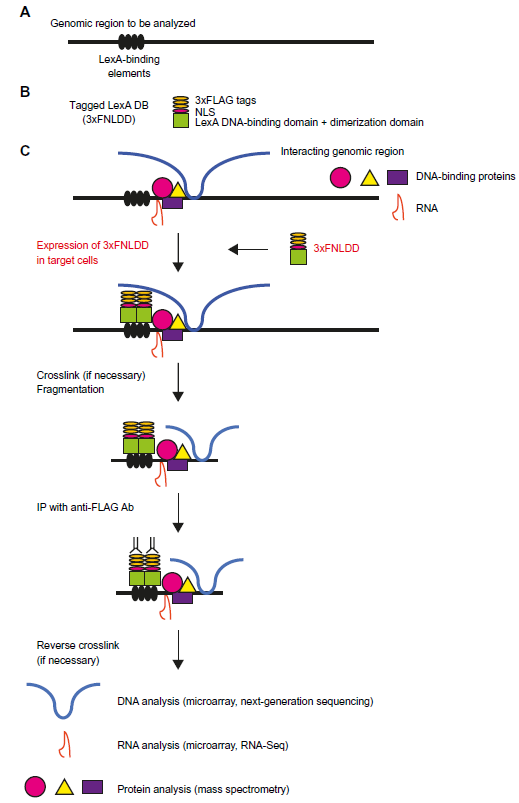
iChIP. (
A repeat of the recognition sequence of an exogenous DNA-binding protein is inserted into the target genomic region of the cell to be analyzed. Examples of these exogenous DNA-binding proteins include LexA, LacI, and Tet repressor.
The DNA-binding domain of the exogenous DNA-binding protein can be fused with a tag(s) and a nuclear localization signal (NLS) and is expressed in the cell to be analyzed.
If necessary, the cells are cross-linked with a compound such as formaldehyde.
Chromatin fraction is prepared, and chromatin DNA is fragmented by sonication or digestion with a endonuclease(s).
Chromatin complexes containing the DNA-binding domain are subjected to affinity purification using an antibody (Ab) or an affinity reagent against the tag(s) or the DNA-binding domain.
Isolated chromatin complexes are reverse cross-linked if a cross-linker was used in step 3.
Proteins contained in the isolated complexes are identified by mass spectrometry (MS), whereas RNAs and DNA are identified by next-generation sequencing (NGS) or microarray analysis.
In our initial attempts, using the first-generation tagged LexA, the yields of the target loci were about 2.5% of input. 14 After optimization of tags and LexA domains, we could achieve yields greater than 10% of input using 3xFNLDD, our second-generation tagged LexA, which consists of a 3xFLAG-tag, an NLS, and the DNA-binding domain and dimerization domain of LexA. 16 The yields of iChIP are important to decide how many cells are necessary for downstream analysis. In our experience, we were able to identify proteins associated with a single-copy locus from 5 × 10 7 cells when the yield (% input) was around 10%. Therefore, we suggest iChIP users to change the cell numbers to be used according to the yield. In this regard, if the yield is significantly lower than 1%, it might be difficult, if not impossible, to harvest sufficient number of cells. In that case, we would suggest choosing other genomic regions for locus-tagging. In this regard, enChIP is much more flexible in testing multiple target genomic regions (see below). In contrast, for the identification of nucleic acids associated with the target locus, smaller number of cells can be used.
in vitro iChIP
In addition to the original iChIP procedure, in which the tagged exogenous DNA-binding protein is expressed in the cells to be analyzed, we developed
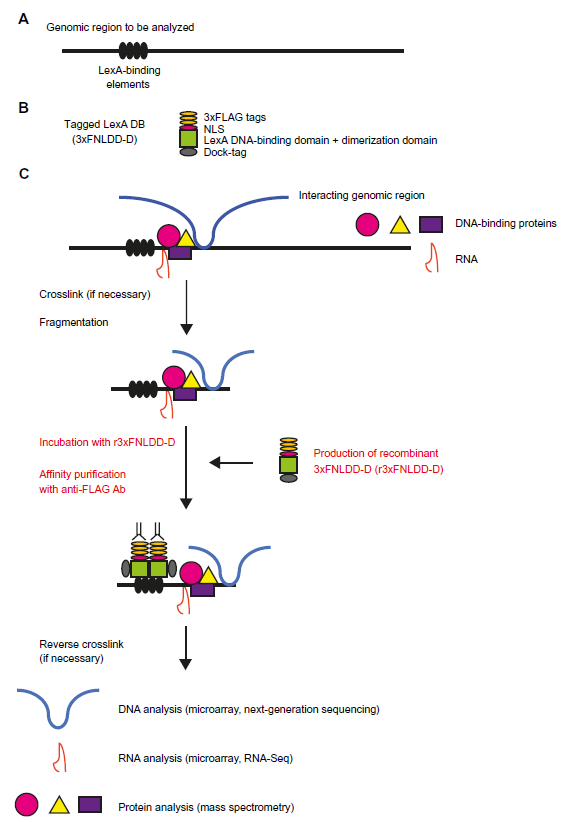
Applications of iChIP
iChIP can be performed using cells in which recognition sequences of an exogenous DNA-binding molecule have been knocked in at the endogenous locus as well as using cells harboring transgenes that retain such recognition sequences (Fig. 3). If a transgene contains functional sequences such as gene regulatory elements, random integration of transgenes could be beneficial because the cells might acquire multiple copies of the transgene, allowing more target genomic regions to be obtained by iChIP. On the other hand, tagging the endogenous locus would be physiologically more relevant. It has not been easy to perform knock-ins in cell lines, but the advent of genome engineering technologies makes the knock-in approach much more straightforward.
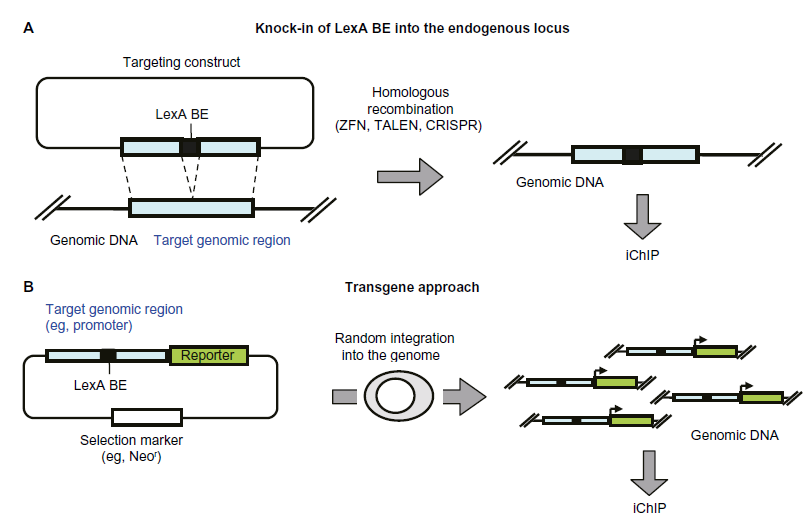
Knock-in and transgene approaches of iChIP. (
We hold patents on iChIP (“Method for isolating specific genomic regions,” US patent 8,415,098; Japan patent 5,413,924). Since we developed iChIP in 2009,
14
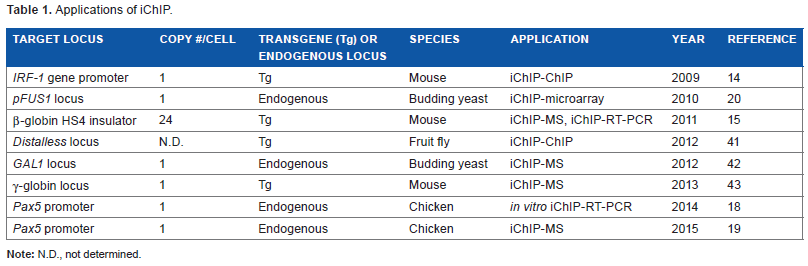
it has been used to identify molecules associated with specific genomic regions (Table 1). For example, iChIP combined with MS (iChIP-MS) enabled us to identify proteins associated with specific genomic regions,
15
including a single-copy locus in cells of multicellular eukaryotes.
19
In our analysis of chicken β-globin HS4 insulator (cHS4), we detected p68 as a known cHS4-interacting protein.
15
In addition, we found that the cHS4 complex binds to a nuclear matrix protein, Matrin-3, mediated by CCCTC-binding factor (CTCF),
15
suggesting that the tethering of cHS4 insulator complexes to the nuclear matrix by Matrin-3 might be important for the function of the cHS4 insulator. In our analysis of chicken
Applications of iChIP.
enChIP
Scheme of enChIP
Although iChIP is a powerful technique, it requires the insertion of recognition sequences of an exogenous DNA-binding protein such as LexA into target loci. The advent of engineered DNA-binding molecules such as zinc-finger proteins, 21 transcription activator-like (TAL) proteins, 22 and the clustered regularly interspaced short palindromic repeat (CRISPR) system23,24 prompted us to utilize such engineered DNA-binding molecules for the isolation of specific genomic regions, leading to the development of enChIP.25–31 The scheme of enChIP is as follows (Fig. 4):
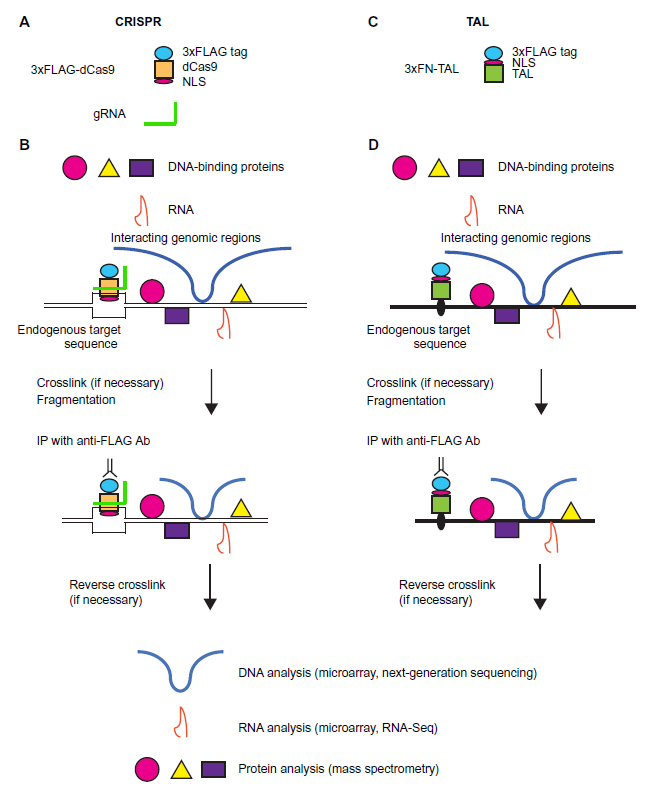
enChIP. A locus to be analyzed is tagged with engineered DNA-binding molecules such as the CRISPR complex, consisting of a catalytically inactive form of Cas9 (dCas9) and gRNA (
An engineered DNA-binding molecule recognizing the target genomic sequence is generated.
The engineered DNA-binding molecule can be fused with a tag(s) and an NLS, and the fusion protein is expressed in the cell to be analyzed.
If necessary, the cells are cross-linked with a compound such as formaldehyde.
The chromatin fraction is prepared, and chromatin DNA is fragmented by sonication or digestion with a endonuclease(s).
Chromatin complexes containing the engineered DNA-binding molecule are subjected to affinity purification using an Ab or an affinity reagent against the tag(s) or the engineered DNA-binding molecule.
Isolated chromatin complexes are reverse cross-linked, if a cross-linker was used in step 3.
Proteins contained in the isolated complexes are identified by MS, whereas RNAs and DNA are identified by NGS or microarray analysis.
Because enChIP does not require the insertion of recognition sequences of an exogenous DNA-binding molecule, which is necessary for iChIP, the time and cost of the analysis can be significantly reduced. This is one of the greatest advantages of enChIP over iChIP.
enChIP using CRISPR
In our first report of the use of an engineered DNA-binding molecule for the purification of specific genomic regions, we utilized the CRISPR system, which consists of a catalytically inactive form of Cas9 (dCas9) and guide RNA (gRNA). 25 dCas9 contains two mutations that abolish endonuclease activities but preserves the DNA-binding activity. 32 Because the recognition of target genomic sequences is solely determined by gRNA, we can easily perform enChIP in a short period of time. The yields of enChIP using CRISPR were comparable with those of iChIP.25,27
enChIP using TAL
enChIP can be performed using other engineered DNA-binding molecules. For example, when we performed enChIP using a tagged TAL protein that recognized the telomeric repeat to isolate telomere regions, we were able to successfully isolate telomere regions.
26
The yields of enChIP using TAL proteins were about 1%,
26
lower than those of enChIP using CRISPR. At this stage, it is not clear whether the difference in yields is intrinsic to the system employed or because of the difference in the target loci.
Identification of proteins associated with a specific genomic region using enChIP
In our initial attempt, using transient expression of dCas9 and gRNA targeting the promoter region of interferon (IFN)-inducible
Identification of RNAS associated with a specific genomic region using enChIP
In addition to proteins associated with specific genomic regions, we could detect RNAs associated with specific loci using enChIP combined with RT-PCR (enChIP-RT-PCR).
26
For example, we detected the RNA component of telomerase
Comparison of iChIP and enChIP
As described earlier, both iChIP and enChIP enable the isolation of specific genomic regions that retain molecular interactions, allowing the identification of associated molecules. One of the advantages of enChIP over iChIP is that it does not require knock-in of the recognition sequence of an exogenous DNA-binding molecule, such as LexA, which is necessary for iChIP. This is important because the insertion of such sequences might abrogate normal functions of the targeted genomic regions; if this was the case, it would be necessary to insert the sequences into different genomic regions. In this regard, even if enChIP is used, the binding of an engineered DNA-binding molecule might affect the function of the genomic region. However, in this case, engineered DNA-binding molecules that recognize different DNA sequences could be tested more easily. In this regard, the CRISPR system is most flexible because many different gRNAs can be generated and tested easily and economically. By contrast, generating multiple TAL proteins recognizing different target sequences can be expensive and time consuming.
On the other hand, iChIP has some advantages over enChIP. First, because only one allele can be tagged with an exogenous DNA-binding molecule and its recognition sequences, allele-specific analysis can be performed by iChIP. To perform allele-specific analysis in enChIP, sequence differences such as single nucleotide polymorphisms must exist between alleles of the target genomic region, and therefore, careful evaluation is necessary to confirm allele-specific isolation. Second, the
Management of Detection of Nonspecific and off-Target Interactions in Performing Locus-Specific ChIP
One of the major problems of detecting specific interactions of genomic regions with proteins, RNAs, and other genomic regions is nonspecific binding. In addition, it has been shown that dCas9 exhibits significant binding to off-target sites.36–39 Therefore, it should be necessary to cancel out the effects of nonspecific and/or off-target interactions. We suggest putting the following comparison sets in performing locus-specific ChIP:
Comparison of different conditions such as different stimulations (eg, the absence or presence of IFNγ stimulation in reference 33) or different cell types (eg, B cells and macrophages in reference 19).
Comparison of different engineered DNA-binding molecules for a target region in enChIP. Molecules detected commonly using different engineered DNA-binding molecules would represent true positives. In this regard, enChIP using CRISPR would be advantageous over that using zinc finger and TAL proteins because different gRNAs can be easily tested for each genomic region. For enChIP using CRISPR, it would be better to include cells expressing only the tagged dCas9 but not gRNA as a negative control to exclude molecules associated with dCas9.
Conclusions
Locus-specific ChIP technologies, including the complementary approaches iChIP and enChIP, are powerful tools for the isolation of specific genomic regions to identify their interacting molecules. Detailed protocols of iChIP and enChIP are attached as Supplementary Files and found in our published methods papers.27,29,40 Updated versions of these protocols in the future can be downloaded from our homepage (http://www.biken.osaka-u.ac.jp/lab/microimm/fujii/iChIP_protocols/english.html).
Author Contributions
Wrote the first draft of the manuscript: TF, HF. Contributed to the writing of the manuscript: TF, HF. Jointly developed the structure and arguments for the paper: TF, HF. Made critical revisions and approved final version: TF, HF. Both authors reviewed and approved of the final manuscript.
Supplementary Material
Supplementary File 1
Procedure to purify genomic DNA with iChIP/enChIP.
Supplementary File 2
iChIP/enChIP SILAC procedures.
Supplementary File 3
iChIP/enChIP MS procedures.
Supplementary File 4
iChIP/enChIP RNA-Seq procedures.
Supplementary File 5
Preparation of retroviral vector plasmids.
Supplementary File 6
Design of gRNA and construction of gRNA expression vectors.
