Abstract
The transcription factor family intimately regulates gene expression in response to hormones, biotic and abiotic factors, symbiotic interactions, cell differentiation, and stress signalling pathways in plants. In this study, 170 AP2/ERF family genes are identified by phylogenetic analysis of the rice genome (Oryza sativa l. japonica) and they are divided into a total of 11 groups, including four major groups (AP2, ERF, DREB, and RAV), 10 subgroups, and two soloists. Gene structure analysis revealed that, at position-6, the amino acid threonine (Thr-6) is conserved in the double domain AP2 proteins compared to the amino acid arginine (Arg-6), which is preserved in the single domain of ERF proteins. In addition, the histidine (His) amino acid is found in both domains of the double domain AP2 protein, which is missing in single domain ERF proteins. Motif analysis indicates that most of the conserved motifs, apart from the AP2/ERF domain, are exclusively distributed among the specific clades in the phylogenetic tree and regulate plausible functions. Expression analysis reveals a widespread distribution of the rice AP2/ERF family genes within plant tissues. In the vegetative organs, the transcripts of these genes are found most abundant in the roots followed by the leaf and stem; whereas, in reproductive tissues, the gene expression of this family is observed high in the embryo and lemma. From chromosomal localization, it appears that repetition and tandem-duplication may contribute to the evolution of new genes in the rice genome. In this study, interspecies comparisons between rice and wheat reveal 34 rice loci and unveil the extent of collinearity between the two genomes. It was subsequently ascertained that chromosome-9 has more orthologous loci for CRT/DRE genes whereas chromosome-2 exhibits orthologs for ERF subfamily members. Maximum conserved synteny is found in chromosome-3 for AP2 double domain subfamily genes. Macrosynteny between rice and Arabidopsis, a distant, related genome, uncovered 11 homologs/orthologs loci in both genomes. The distribution of AP2/ERF family gene paralogs in Arabidopsis was most frequent in chromosome-1 followed by chromosome-5. In Arabidopsis, ERF subfamily gene orthologs are found on chromosome-1, chromosome-3, and chromosome-5, whereas DRE subfamily genes are found on chromosome-2 and chromosome-5. Orthologs for RAV and AP2 with double domains in Arabidopsis are located on chromosome-1 and chromosome-3, respectively. In conclusion, the data generated in this survey will be useful for conducting genomic research to determine the precise role of the AP2/ERF gene during stress responses with the ultimate goal of improving crops.
Introduction
Plants encounter various natural affronts, from pests, diseases, and ecological changes, including those affecting temperature and water. These factors greatly impinge on a plant's endurance, growth, and production. In order to stay alive and grow in diverse environments, plants have developed a complex signalling network at the molecular, cellular, and system levels. Gene co-expression at the transcription level is the foremost regulatory mechanism in biological processes. Transcription factors (TFs) control the majority of multiple stress response genes in a synchronized manner and are characterized as attractive targets for application in molecular plant biology. Among different TFs, the ethylene responsive transcription factors (ERF) family plays a vital role in plant growth and enables plants capable of fighting ambiance changes.1–4 Therefore, it is important to understand the function of these genes in order to ameliorate crop yield and allow them to grow in sundry environmental conditions.
In the plant Kingdom, APETALA2/ethylene-responsive element-binding protein (AP2/EREBP) is a large family of TFs that contain AP2, RAV, and ERF family genes. This super family is characterized by the conserved AP2/ERF DNA binding domain, comprising of 60–70 amino acid residues.5,6 This domain, first reported in the homeotic gene AP2 of Arabidopsis, is involved in flower development. 7 The ERF domain was first described as a conserved motif in four DNA-binding proteins (EREBP1, 2, 3, and 4; presently renamed ERF1, 2, 3, and 4, respectively) in tobacco plants. This domain binds to a GCC box, which is a DNA sequence involved in the ethylene-responsive transcription of genes. 8 In tomatoes, three proteins (Pti-4, Pti-5, and Pti-6) are reported to interact with disease resistance proteins, determined using yeast two-hybrid assays, 9 and each protein has a domain homologous to the AP2/ERF domain. Based on the number of AP2/ERF domains and their gene function, the AP2/ERF gene family is divided into four subfamilies: AP2, RAV, dehydration-responsive element-binding protein (DREB), and ERF. 10
The AP2 family genes possess two repeats of the AP2/ERF domain and the ERF family proteins contain a single AP2/ERF domain whereas RAV (related to VP1/ABI3) family proteins have an additional B3 DNA binding domain. The AP2 subfamily is further subdivided into two monophyletic groups: AP2 and AINTEGUMENTA (ANT). 11 Based on ERF domain binding to DNA sequences, the ERF family is further split up into two subfamilies: ERF and CBF/DREB. 6 Proteins encoded by ERF subfamily genes bind to the core motif AGCCGCC;8,12 whereas, CBF/DREB subfamily genes containing C-repeats recognize the cis-acting element, A/GCCGAC. 13
ERF subfamily genes have conserved alanine (Ala) at position-14 and aspartic acid (Asp) at position-19, while CBF/DREB subfamily genes have valanine (Val) at position-14 and glutamine (Glu) at position-19. 14 The three-dimensional structure of the AP2/ERF domain from AtERF1 is portrayed using hetero-nuclear multidimensional Nuclear Magnetic Resonance and is comprised of three anti-parallel β-sheets and an α-helix. 15 Moreover, tryptophan (Trp) and arginine (Arg) residues present in the β-sheets come into contact with DNA during transcription. 14
The AP2/ERF family is identified in the Arabidopsis,6,10 grapevine, 16 poplar, 17 and rice genomes with 145, 132, 200, and 163 genes, respectively. 18 Using genetic and molecular approaches, AP2/ERF proteins were determined to play an imperative role in the transcriptional regulation of diverse biological processes. These processes are related to plant growth, development, and response to environmental stimuli.2,19 The genes in this family participate in different pathways in response to hormones and biotic and abiotic stresses, such a jasmonate,20,21 abscisic acid (ABA),22,23 drought,24,25 salinity,26,27 cold,28,29 disease,21,30,31 and flooding stress. 32 The AP2 family contributes to the regulation of developmental processes, such as flower development,7,33 early floral meristem identity, 34 crown and lateral roots,35,36 and somatic embryogenesis. 37 In addition, the involution of RAV family members is reported in the AP2/ERF family in response to ethylene, 38 brassinosteroid,39,40 and biotic and abiotic stresses.30,41,42
Determination of the phylogenetic relationship is a crucial step in elucidating the evolution of crop species. In the past, AP2/ERF genes were considered plant specific but, 5 recently, this domain was reported in non-plants for the first time in the ciliate Tetrahymena thermophila, 43 cyanobacterium Trichodesmium erythraeum, and viruses Enterobacteria phage Rb49 and Bacteriophage Felix 01. 44 The non-plant AP2-domain containing proteins have HNH domains edging the AP2/ERF domain. These are known as HNH endonuclease encoded by the His-Ans-His sequence motif. 45 It is hypothesized that AP2/ERF TFs may have originated by horizontal transfer of the (HNH)-AP2 endonuclease from bacteria/viruses into plants via transpositions and homing processes. 44
The above cited work was done on different plants whereas the present work is reported on rice. Belonging to the gramene family, rice is the world's most important staple crop that ensures food security. In addition to its agricultural utility and small genome size (394Mb), rice provides an excellent model plant system to study plant-pathogen interactions, hormonal pathways, and resistance to environmental stresses. Some AP2/ERF family genes have been cloned from rice, but the function of the majority of them remains unclear. The completion of high-quality sequencing of the rice genome has furnished an excellent opportunity for elaborate gene analysis.
In this study, 170 OsAP2/ERF rice genes are obtained from different database searches and are classified into their respective clades according to their homology with known genes. The functional analysis of each transcriptional factor belonging to the AP2/ERF family is carried out keeping in mind their functional redundancy. In addition, phylogenetic analysis of AP2/ERF genes, complete alignment of each subfamily gene, and distribution of the conserved motifs outside the AP2/ERF domain exhibiting the specificity for gene function is performed. Moreover, to generate a clear picture of AP2/ERF genes, the position of each gene on the chromosomes is determined and gene expression profile canvasing is executed to determine tissue specificity. Comparative mapping between a monocot (wheat) and a eudicot (Arabidopsis) is performed to identify homologs/orthologs among the genomes. The data brought forth in this canvas will help in the selection of appropriate genes for further functional characterization and understanding of precise regulatory checkpoints operating during development and stress responses in crop plants.
Materials and Methods
Identification of AP2/ERF genes in rice genome
First, rice AP2/ERF genes were identified in the genome of Oryza sativa l. japonica cultivar Nippon-bare using ESTs and cDNA sequences. The data were downloaded from various public repositories, including The National Centre for Biotechnology Information (NCBI), 46 The Database of Rice Transcription Factors (DRTF), 47 The MSU Rice Genome Annotation Project Database, 48 Knowledge-based Oryza Molecular Biological Encyclopedia (KOME), 49 and Plant Genome Database (PlantGDB). 50 Next, all retrieved sequences were subjected to the BLAT online tool available on the RAP-DB website to find homologous sequences in the rice genome. 51 The sequences showing more than 80% coverage areas were expanded approximately 2000 bp on both sides of the hit to find the open reading frame (ORF) using the GENSCAN online tool. 52 Data assembly was performed using a DNA Assembly Sequence Programme CAP3. 53 In addition, Simple Modular Architecture Research Tool (SMART) is used to confirm the presence of the AP2/ERF domain in the resulting sequences. 54
Phylogenetic and MEME motif analysis
The AP2/ERF domain comprising protein sequences obtained from various sources are aligned using ClustalX 2.0 54 and redundant entries are removed. 55 A combined un-rooted neighbor-joining (NJ) tree was generated in MEGA 4.0 56 with the following default parameters: 56 poisson correction, pairwise deletion, and bootstrap (5000 replicates). Conserved motifs in rice AP2/ERF protein sequences were identified using a motif based sequence analysis tool, MEME Suite version 4.7.0, 57 with following parameters: optimum width 6–200 amino acids, any number of repetitions of a motif, and maximum number of motifs set at 25. The BLAST search for the resulting motifs in NCBI and MS-Homology databases was carried out to determine their significance.
Intron/exon size distribution of AP2/ERF family genes
Intron positions in genes are ascertained through the identification of gaps in alignment of full-length cDNA transcripts with genomic sequences using the online tool Gene Structure Display Server (GSDS). 58 Concisely, for a single full-length cDNA aligned against a conterminous stretch of genomic sequence, exons are proximal blocks of homologous sequence between full-length cDNA and genomic sequences, whereas introns are gaps between exons consisting entirely of genomic sequence. The general distribution of introns within each coding DNA sequence (CDS) is analyzed by the distribution of exon sizes. The mean exon size for a full length cDNA containing ni introns (regardless of pattern of distribution) is calculated as [Length of coding DNA sequence/(ni + 1)]. 59 The total length of a gene is calculated by adding the length of all of the exons of each gene. Moreover, exon size and total gene length of each gene are plotted graphically to clarify the range and size of AP2/ERF family genes.
Gene expression analysis
To further investigate and confirm AP2/ERF gene expression, the rice expression profile database (RiceXPro), which is a repository of gene expression data derived from microarray experiments encompassing the entire life cycle of the rice plant from germination, seedling, tillering, stem elongation, panicle initiation, booting, heading, flowering, and ripening stage, 60 was used. This tool generates a heat map of normalized signal intensity values for each plant tissue for each gene and provides a quantitative measure of the transcript of particular genes.
In Silico identification of AP2/ERF gene family orthologs/homologs in rice, wheat and Arabidopsis genomes
Comparative genetic mapping among closely related (wheat) and distantly related (Arabidopsis) genomes divulges a high level of synteny in gene content and gene function. The Arabidopsis AP2/ERF genes were retrieved from the supplementary file provided by a genome wide survey of ERF family genes in Arabidopsis and rice, 10 and were confirmed using a database of genetic and molecular biology for the model plant Arabidopsis thaliana (TAIR). 61 Regarding wheat (Triticum aestivum), AP2/ERF sequences were retrieved from NCBI, 46 Plant Genome Database (PlantGDB), 50 and Chinese-Spring draft genome assembly or raw sequence reads using BLAST available online at CerealsDB, University of Bristol. Wheat and Arabidopsis AP2/ERF consensus gene sequences were used to BLAST against rice annotated genes and rice full length (FL) cDNAs with BLASTx and BLASTn, respectively, to identify orthologs/homologs between the genomes.
Results and Discussions
To ascertain the AP2/ERF family genes in the rice genome, sequences obtained from various resources as mentioned in the materials and methods were assembled using the CAP3 online tool. After confirmation of the AP2/ERF domain using the SMART online tool and removing the redundant sequences by alignment using ClustalX 2.0, it was determined that 170 sequences encode the AP2/ERF domain (Supplementary File 2). It was also discovered that 143 genes encode a single AP2/ERF domain. Twenty-three genes are anticipated to encode proteins comprising double AP2/ERF domains whereas four genes are prognosticated to encode a single AP2/ERF domain together with one B3 domain. Based on these results, the former and latter genes are appointed to the AP2 with double domain and the RAV families, respectively.
These results are an improvement on the AP2/ERF super family described by Nakano and his co-workers as they do not use these two subfamilies in their investigation under genome-wide analysis of the ERF gene family in Arabidopsis and rice (Supplementary File 2). 10 Regarding the importance of the AP2 subfamily genes with double domains, the ANT gene plays a critical role in the development of gynoecium marginal tissues (eg, stigma, style, and septa), and in the fusion of carpels and medial ridges that lead to ovule primordial. 62 BABY BOOM (BBM) promotes cell proliferation, differentiation, and morphogenesis, especially during embryogenesis. 63 WRINKLED (WRl) genes are involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways. These are involved in the activation of a subset of sugar-responsive genes and the control of carbon flow from sucrose import to oil accumulation in developing seeds. 64 The PLETHORA (PLT) gene acts as a transcriptional activator and functions as a master regulator of basal/root fate; it is essential for root quiescent centre (QC) and columella specification, stem cell activity, as well as for the establishment of the stem cell niche during embryogenesis. PLT also modulates root polar auxin transport by regulating the distribution of PIN genes and plays an essential role in specifying pattern and polarity in damaged roots. 65 Besides, genes with floral homeotic proteins promote early floral meristem identity and are subsequently required for the transition of an inflorescence meristem into a floral meristem; they play a central role in the specification of floral identity, particularly for the normal development of sepals and petals in the wild-type flower. 66
Other subfamily genes with one B3 domain function as negative regulators of plant growth and development. These genes are expressed in all tissues: roots, rosette leaves, cauline leaves, inflorescence stems, flowers, and siliques. They have highest expression in roots and rosette leaves and low expression in flowers. 39
Phylogenetic relationships between AP2/ERF family transcription factors in rice
To dissect the phylogenetic relationships between rice AP2/ERF family proteins, multiple alignment analyses were executed using the amino acid sequences of the AP2/ERF domains. The alignment predicts that Gly-4, Val-5, Gly-11, Ile-17, Arg-26, Trp-28, Leu-29, Gly-30, Ala-38, Ala-39, Asp-43, Gly-51, Asn-58, and Phe-59 are completely conserved in all AP2/ERF single domain family members in rice (Fig. 2A). In addition, more than 95% of ERF members have conserved Arg-3, Arg-6, Arg-8, Glu-16, Arg-18, Ala-35, Ala-45, Ala-46, and Ala-55 amino acid residues. The AP2/ERF proteins with a single domain in the AP2/ERF subfamily contain conserved Alanine (Ala) at position-14 and Aspartic acid (Asp) at position-19, whereas the CBF/DREB subfamily genes have Valine (Val) at position-14 and Glutamine acid (Glu) at position-19 (Fig. 1). 14 Instead, few gene domains are without Glu-19 and exhibit significant similarity with the rest of the domain although genes in the RAV family with a B3 domain have Gly-14 instead of (Val/Ala-14) (Fig. 2A).

The conserved amino acid residues found in ERF (A) and CBF/DREB (B).

Alignment of the AP2/ERF domains from Rice (O. sativa) cv. japonica ERF proteins. (A and B) displayed the alignment of single AP2/ERF and double AP2/ERF double domain proteins respectively.
The multiple alignments of the amino acid sequences of AP2/ERF double domain proteins were determined. The amino acid residues were as follows: Gly-4, Val-5, Gly-12, Glu-15, His-17, Asp-20, Gly-38, Ala-46, Ala-47, Arg-48, Ala-49, Asp-51, Ala-53, Ala-54, Lys-56, Gly-59, Asn-67, and Phe-68. These were found to be conserved in the first domain of AP2/ERF double domain proteins. It is of note that at position-6, Thr-6 is conserved in the double domain AP2 genes compared to the Arg-6 conserved in single domain ERF proteins. Moreover, the His-amino acid is found in both the domains of double domain AP2/ERF proteins that are missing in the single domain AP2/ERF proteins. In the second domain of AP2/ERF double domain proteins, the alignment indicated that the amino acid residues Tyr-2, Arg-3, Gly-4, Val-5, Gly-12, Arg-13, Trp-14, Ala-16, Arg-17, Gly-19, Ala-44, Tyr-45, Asp-46, Ala-48, Ala-49, Ile-50, Gly-54, Thr-59, Asn-60, and Phe-61 are highly conserved (Fig. 2B).
Based on these observations, a phylogenetic tree of 170 AP2/ERF genes was constructed using bootstrap analysis (5000 replicates) based on the multiple sequence alignments of their protein domains (Fig. 3). The bootstrap values for the nodes in this phylogenetic tree are not high in every case, similar to the results of the phylogenetic analysis done on Arabidopsis ERF proteins. 10 The phylogram is alienated into a total of 11 groups with four main groups, namely group-I to group-IV. The members of the ERF gene family are classified into 10 groups in Arabidopsis. 10 Group-I is distinguished into three sub groups: I-a, I-b, and I-c. The proteins categorized in group-I are characterized with a double domain. These proteins play important roles in Arabidopsis, rice, Brassica napus (Rape), and Medicago truncatula for vegetative to embryonic growth, somatic embryogenesis, seed development, lateral roots formation, petal cell identity, floral meristem identity and transition to flowering time.33,36,37,63,64,67–69 The genes in this group are expressed more in the flower, leaves, and stem as opposed to the roots. 70
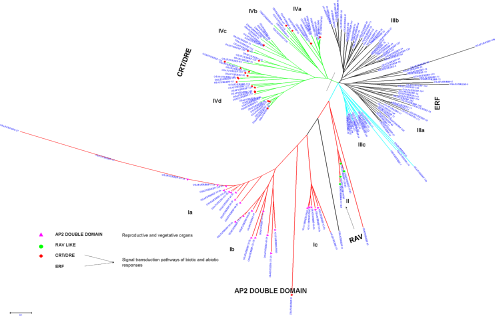
An unrooted phylogenetic tree of Rice AP2/ERF proteins is constructed using NJ method.
AP2/ERF genes in Group-II consist of one additional B3 domain and are distinguished as RAV proteins. The function of these proteins in Arabidopsis are as negative regulators of plant growth and development and in the tomato play a pivotal modulator role in defense mediated pathways.39,30 The remaining genes fall in into group-III and group-IV and have a single AP2/ERF domain. The genes in group-III are part of the ERF subfamily that are used in crop plants to defend against biotic resistance, such as pathogens and diseases. The role of these genes have been studied extensively in Arabidopsis, rice, tomato, and tobacco plants with respect to hormone response (jasmonic acid, salicylic acid, absicic acid, and ethylene), 10 pathogen/disease's resistance,4,12,31,71–77 and insect damage (Supplementary Table S2). 78 Some family members play crucial roles in transcription inhibition due to the presence of an EAR motif.10,79 It is reported that ERF is over expressed in shoots due to low temperature, but expressed little in cultured cells and roots.3,80,81
Finally, the genes in Group-IV are characterized into CBF, DREB, and TINY proteins, which are used for abiotic factors, such as cold, salt, and drought resistance. These genes are characterized in Arabidopsis, rice, maize, rape seed, sunflower, tomato, and tobacco plants for their response to abiotic factors (Supplementary Table S2).6,25,82–93 The inclusion of these genes in distinct groups based on their protein similarities need to be characterized according to different stress regimes, which may help in correlating function within their phylogenetic placement.
Distribution of conserved motifs
Complete AP2/ERF protein sequences of rice were analyzed for the presence of conserved motifs using the MEME Suite version 4.7.0. Overall, 19 conserved motifs were identified and named 1–19. The consensus sequences of these motifs are provided in Supplementary File 1. Motifs 1, 2, 3, 4, 5, 6, and 9, correspond to the AP2/ERF domain region. The remaining 18 motifs were found to characterize specific clades in the phylogentic tree. The distribution of these conserved motifs in proteins of relevant clades in the phylograms is laid out in Figure 4.
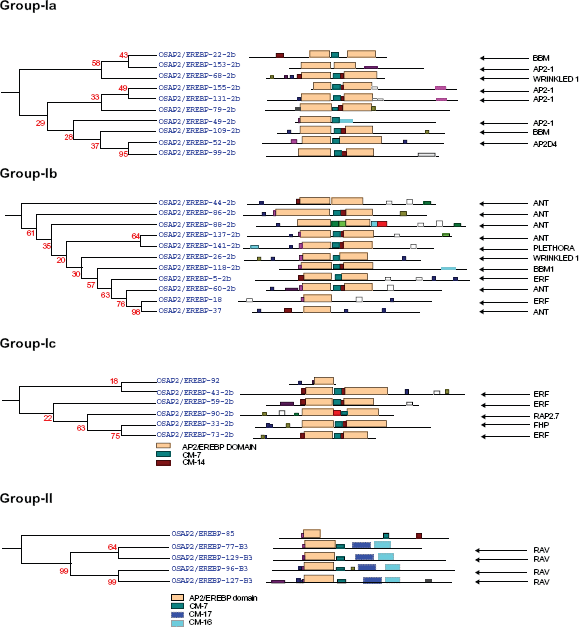
The phylogenetic relationship in the AP2/ERF family genes. Group-Ia, Ib and group-Ic represent the AP2/ERF double domain proteins with their conserved motifs. Group-II exhibited the RAV family genes with one B3 domain. Their conserved motifs determined by the MEME online tools are given the Supplementary File 1.
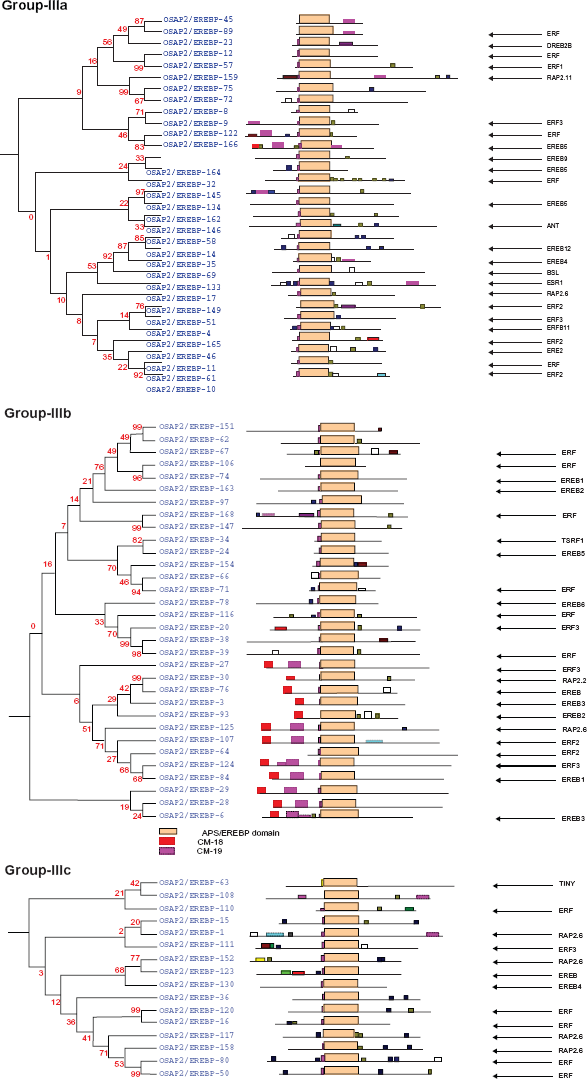
The phylogenetic relationship among the AP2/ERF family genes in rice. group-IIIa, IIIb and IIIc corresponds to the AP2/ERF single domain proteins. group-IIIa, IIIb and IIIc falls in the ERF. Their conserved motifs determined by the MEME online tools are given the Supplementary File 1.

The phylogenetic relationship among the AP2/ERF family genes in rice. Group-IV symbolises the AP2/ERF single domain proteins. Group-IVa, IVb, IVc and IVd are predicted to relate CBF/DREB genes. Their conserved motifs determined by the MEME online tools are given the Supplementary File 1.
Conserved motifs outside the AP2/ERF domain
In general, transcription factors comprise functionally important conserved motifs outside the DNA binding domain, which are related to transcriptional activity, nuclear localization, and protein-protein interactions. 10 Such functional amino acid sequences motifs are often conserved among members of a subgroup in large families of transcription factors in plants; proteins with these motifs in their subgroups are likely to have similar functions. 94
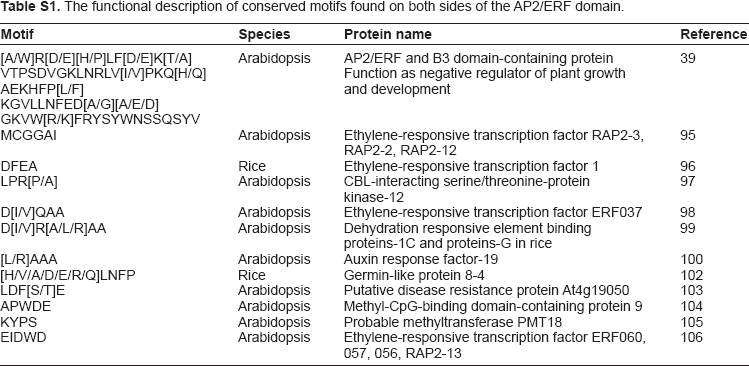
An investigation of the conserved motifs in the proteins of each clade in the AP2/ERF family of rice was accomplished using multiple alignment analysis with ClustalX 2.0. 55 The conserved motifs found in the OsAP2/ERF family are summarized in Supplementary Table S1. Most of the motifs are selectively distributed among the specific clades. For instance, in Arabidopsis, the conserved motifs CM-16 and CM-17 found in the C-terminal region of group-II with consensus sequences KGVLLNFED[A/G] [A/E/D]GKVW[R/K]FRYSYWNSSQSYV and [AW] AR[ED][HP]LF[DE]K[AT]VTPSDVGKLNRLV [IV]PKQ[HQ]AE[KR]HFP[LF], respectively, function as a transcriptional repressor for plant growth and development (Fig. 5 and Supplementary Table S1).39,49 CM-18, MCGGAI[LI]AD[LF]IP, and CM-19, D[DE] D[FW]EA[AD]F[ER]EF[EDL][DSV][DR][DS] [DGH]D[DS][DE]D[ED], are found in the N-terminal region of members of group-III. They have two small blocks of conserved amino acid's sequences, eg, MCCGAI and DFEA, which play a role in ethylene transcriptional activation (Fig. 6, Supplementary Table S1).95,96 CM-11 in the members of group-IVc and IVd have, at the C-terminal, the consensus sequence [LR]PR[PA]A[ST]A[SA]PKD[VI][QR]AAAA[LA] AAA[AM]ARPPP. The alignment of the members predicted four small blocks of conserved amino acid residues: LPR[P/A], D[I/V]QAA, D[I/V]R[A/L/R] AA, and [L/R]AAA. These are the essential signatures in Arabidopsis for CBL-interacting serine/threonine-protein kinase-12, Ethylene-responsive transcription factor ERF037, dehydration responsive element binding proteins-1C, and proteins-1G and Auxin response factor-19, respectively (Fig. 7, Supplementary Table S1).97–100 The [L/R]AAA motif is a homologue of auxin response factors (ARFs). It binds specifically to the DNA sequence 5′-TGTCTC-3′ and is discovered in the auxin-responsive promoter elements (AuxREs). This motif acts as a transcriptional activator and is involved in ethylene responses and it regulates lateral root formation through direct regulation of LBD16 and/or LBD29. 101 The conserved amino acid sequence LPR[P/A] detected in the CBL proteins (serine-threonine protein kinases) binds to the regulatory NAF domain of the CIPK protein and activates the kinase in a calcium-dependent pathway. 97 The [H/V/A/D/E/R/Q]LNFP motif is found to be involved in disease resistance. 102

RAV like motif sequences conserved in the C-terminal region of group-II in rice.

The conserved motif amino acid sequences in group-III are identified in the N-terminal region.

The conserved motif amino acid sequences in group-IVc and group-IVd respectively are identified in the C-terminal region.
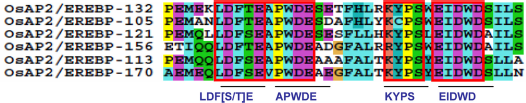
The conserved motif amino acid sequences in group-IVa are identified in the C-terminal region.
Four small preserved amino acid residues were found after alignment in the CM-10 motif with the consensus sequence PEMEKLDFTEAPWDESETFHLRKYPSWEIDWDSILS (Fig. 8, Supplementary Table S1). Among them, a unique small conserved sequence of amino acids, LDF[S/T]E at the C-terminal region, plays a key role in disease resistance. 103 The APWDE motif was found to be involved in transcriptional regulation and acts as a histone acetyltransferase, which mediates the acetylation of histone H3 and H4 of target loci. It is also implied in an auxin-independent regulation of shoot, branching, and flowering time. In addition, it is expressed in leaves, buds, flowers, stems, and over-methylated genomic DNA. 104 The KYPS motif is involved in the DNA methylation process, which plays an important role in genome management and in regulating gene expression during development. 105 A distinctive small motif, EIDWD, was also found to be involved in Arabidopsis response to ethylene. 106 It is clear that most of the conserved motifs identified in this study have plausible features in their amino acid compositions as shown in Supplementary Table S1.
Characteristics of each group in the rice AP2/ERF gene family
The characteristics of each group of the rice AP2/ERF family are described below. For reference, current information regarding the functions of the genes in the AP2/ERF family is summarized in Supplementary Table S2.
Group-I
Group-I is divided into three subgroups: Ia, Ib, and Ic (Fig. 4, Group-Ia, Ib and Ic). All genes in this group have double AP2/ERF domains except OsAP2/EREBP-049, OsAP2/EREBP-018, OsAP2/EREBP-037, and OsAP2/EREBP-092. These genes fall in group-I due to their domain similarities. At this time, the functions of the OsAP2/EREBP-079-2b, OsAP2/EREBP-092-2b, and OsAP2/EREBP-099-2b genes are unknown. All genes have the conserved motifs CM-7 and CM-14 that separates the two domains. The loci with these two conserved motifs are provided in Supplementary File 2 and are reported as ANT, BBM, and AP2 like (AIL) proteins of Arabidopsis.10,37,67 Genes grouped in group-Ia, Ib, and group-Ic are involved in seed (embryonic growth, seed development, seed germination) and flower traits (petal cell identity, flowering time, and floral meristem identity).33,63,64,67,69,
Group-II
Group-II proteins share three conserved motifs: CM-7, CM-16 and CM-17 in the C-terminal region (Fig. 4, Group-II), which are contiguous with the AP2/ERF domain. In addition to a single AP2/ERF domain, these genes have one B3 domain. The RAV family proteins act as negative regulators of plant growth and development in Arabidopsis and take part in defence mediated responses in tomatoes against different stresses.39,40,30
Group-IIIa
Group-IIIa consists of 32 genes (Fig. 4, group-IIIa). The AP2/ERF genes with the generic names OsAP2/EREBP-010, OsAP2/EREBP-011, OsAP2/EREBP-046, OsAP2/EREBP-051, and OsAP2/EREBP-057 interact as a transcriptional activator and are involved in disease resistance pathways in Arabidopsis, rice, tobacco, and tomato plants.4,31,71,72 Their domains bind to the GCC-box pathogenesis-related promoter element. They are involved in the regulation of gene expression induced by stress factors mediated by ethylene that seem dependent on a protein kinase cascade. The genes, coded as OsAP2/EREBP-009 and OsAP2/EREBP-012, are transcriptional inhibitors and may regulate the expression of other genes in a co-expression manner. The N-terminal regions of the genes OsAP2/EREBP-009, OsAP2/EREBP-122, OsAP2/EREBP-134, and OsAP2/EREBP-166 consist of the conserved motif CM-19. Furthermore, defence-related phytohormones, such as ethylene, jasmonate, and salicylic acid differentially induce the expression of this group of genes. 10
Group-IIIb
This subgroup of AP2/ERF genes is also related to biotic stresses, such as disease and pathogen resistance and responds to jasmonite and ethylene signal transduction pathways. Out of the 32 genes, the function of nine of these genes has not been studied. This group shares two motifs, CM-18 and CM-19, in the N-terminal region (Fig. 4, group-IIIb). The functional characterization of the two small conserved proteins blocks, MCGGAI and DFEA, are studied by comparing with the Arabidopsis ERF genes.95,96 The OsAP2/EREBP-074 and OsAP2/EREBP-084 (EREBP1) genes in wheat and cotton repress GCC box-mediated transcription by improving pathogen and abiotic stress tolerance in transgenic plants.73,74 Recently, genes OsAP2/EREBP-020, OsAP2/EREBP-027, and OsAP2/EREBP-124 (ERF3), are up regulated by the feeding of the rice striped stem borer on rice. 78 In addition, OsAP2/EREBP-003, OsAP2/EREBP-006, OsAP2/EREBP-024, OsAP2/EREBP-078, OsAP2/EREBP-093, and OsAP2/EREBP-163 display important roles in cultured cells and roots.80,81
Group-IIIc
Group-IIIc consists of 16 genes (Fig. 4. group-IIIc). The function of three genes, OsAP2/EREBP-015, OsAP2/EREBP-036, and OsAP2/EREBP-108 has not previously been studied. Except for the gene OsAP2/EREBP-063 (TINY), which plays an important part in abiotic stresses, the other genes have a crucial function in biotic stresses (insect resistance, responsive to stress hormones like jasmonic acid, salicylic acid, abscisic acid, and ethylene). 10
Group-IVa
Except for the genes OsAP2/EREBP-019, OsAP2/EREBP-053, OsAP2/EREBP-103, OsAP2/EREBP-047, and OsAP2/EREBP-119, genes in this group contain the CM-10 motif in the C-terminal region of the AP2/ERF domain (Fig. 4 group-IVa). Close inspection of this conserved motif divides it into four blocks with the following protein sequences: APWDE, LDF[S/T]E, EIDWD, and KYPS. In Arabidopsis, these four conserved protein blocks play key roles in disease resistance, auxin responsiveness, and regulation of genes during expression and transcription activation (Supplementary Table S1).103–106 The overexpression of the genes OsAP2/EREBP-132 and OsAP2/EREBP-113 in Arabidopsis are characterized for high salt resistance. 6
Group-IVb
This small group consists of only six genes. Among them, the gene OsAP2/EREBP-167 is involved in water deficit tolerance in rice and sunflowers.25,82 Moreover, the OsAP2/EREBP-095 gene is a disease resistance gene in Arabidopsis. 31 The gene OsAP2/EREBP-025 requires characterization (Fig. 4 group-IVb).
Group-IVc
The roles of the genes in subgroup-IVc have been extensively studied. These genes play essential functions in response to high salinity and cold-stress. 6 Although the functions of the OsAP2/EREBP-031, OsAP2/EREBP-055, OsAP2/EREBP-070, OsAP2/EREBP-083, OsAP2/EREBP-91, OsAP2/EREBP-94, OsAP2/EREBP-101, and OsAP2/EREBP-139 genes are unknown, these genes may play a role as transcriptional activators in gene expression in response to abiotic stresses due to their placement in the phylogram (Fig. 4 group-IVc). In Arabidopsis and corn, TINY (OsAP2/EREBP-142 and OsAP2/EREBP-135) is a homolog of the group-IVc and responds to cold stress.92,93
Group-IVd
The proteins in group-IVd possess the CM-11 motif that is homologous to proteins of group-IVc. Possession of these motifs and their phylogenetic relationships indicate a strong similarity between groups-IVd and IVc (Fig. 4 group-IVd). The function of genes with the generic names, OsAP2/EREBP-002, OsAP2/EREBP-042, OsAP2/EREBP-056, OsAP2/EREBP-105, OsAP2/EREBP-136, and OsAP2/EREBP-157, need to be investigated. The CM-11 motif has a conserved [H/V/A/D/E/R/Q]LNFP amino acid residue sequence whose function is studied in rice and other homologous plant species. 102 Another conserved protein sequence, LPR[P/A] in motif CM-11, also plays an important role in serine-threonine protein kinase mechanisms to activate the kinase in a calcium-dependent manner. 97 The functional characterization of these genes for low temperature, salt, dehydration, drought resistance, and osmotic tolerance has been investigated in tomato, rice, Arabidopsis, and corn plants.
Gene length with respect to intron/exon size
Usually, the intron/exon position in the CDS furnishes clues on evolutionary relationships of genomes. 10 To explore the rice genome to find more AP2/ERF family genes, the full cDNA information is computed using the coverage of 2000 bp region on both sides of the hit to avoid any possible gene losses. It was found that gene length ranged from 781.1 to 1679.7 bp with an average gene length of 1230.4 ± 449.3 bp. These findings indicate that the average gene length is within the 2000 bp limit used in our survey to find these genes. Therefore, it is evident that every exon covered the ESTs and this makes it possible to map the rice genome and this 2000 bp limit ensures that there is no loss of genes (Fig. 9A). For furthering probing, we analyzed the size distribution of exons in our cDNA region by comparing the CDS with genomic sequences in each gene. It was found that the length of exons ranges between 138.4 to 274.79 bp with the average size of exons being 205.3 bp with a standard deviation of 68.96, which is also within the limit and indicates that all the genes are in the expanding range of 2000 bp on both side of the hit (Fig. 9B, Supplementary Table S3).
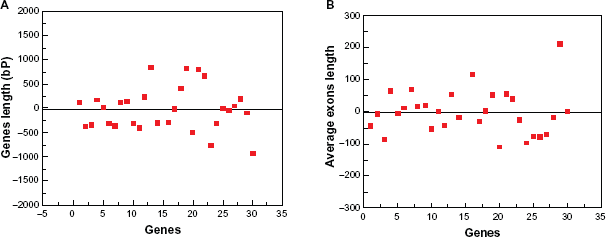
Distribution of AP2/ERF family genes with full cDNA information. (A) Distribution of gene length in 2000 bp mapping hits. (B) Exon size distribution in gene length.
Expression profile analysis of the AP2/ERF gene family
Rice, being a staple food crop, suffers yield losses as a result of environmental cues and pathogens during plant development. It is evident that AP2/ERF family genes play important roles in plant development in response to biotic and abiotic stresses and thus, they present as ideal candidates to investigate the molecular regulation of these processes. To determine the expression pattern of putative AP2/ERF family genes, the RiceXPro database was probed for 12 different kinds of rice tissues. The expression of these genes was detected in the root, stem, leaf, sheath, inflorescence, ovary, anther, embryo, pistil, lemma, palea, and endosperm. Most of the AP2/ERF genes indicate some degree of tissue specificity. 107 Among the vegetative organs, the expression of these genes was found most abundant in the root (14%), followed by the leaf (8%) and stem (6%). With regards to reproductive organs, they were most common in the embryo (12%) and lemma (11%) (Fig. 10). The expression analysis of each gene in different tissues is determined by signal intensity values derived from the RiceXPro database. Most of the genes have a similar degree of tissue specificity except for OsAP2/ERF#001, OsAP2/ERF#128, and OsAP2/ERF#048, which exhibit more expression in 12, 10, and 9 rice tissues, respectively (Fig. 11). It should be noted that the genes with higher expression levels belong to the subgroups IIIc, IVc, IVb, and Ib. The genes in group-IIIb and group-IVd have the same level of gene expression. It is apparent that the members of the subgroup Ia and group-II have the lowest gene expression in the rice tissues used in this canvas (Fig. 11).

Distribution of rice AP2/ERF family genes in vegetative (root, stem, leaf and sheath) and reproductive tissues (inflorescence, ovary, anther, embryo, pistil, lemma, palea and endosperm).
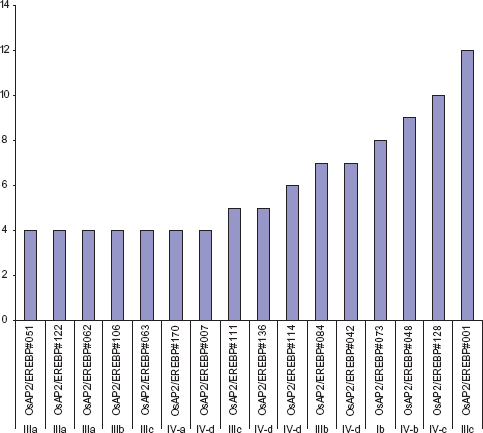
Gene's classification according to their expression level in each tissue.
Furthermore, the function of genes in each group divulges that the members of group-IIIb were highly present in the root. Gene expression during inflorescence, pistil, and anther development was high in members of group-IIIa. During embryogenesis, expression was high in group-Ia and group-IIIb genes. During the heading stage in rice, these AP2/ERF genes were expressed in the lemma and palea in members of group-IVd. Moderate gene expression in the leaf was observed in group-IIIa, whereas the lowest expression of AP2/ERF genes in the stem, sheath, and endosperm was observed in all groups (Fig. 12).

Tissue specific expression of the AP2/ERF family genes in each group.
Recently, transcriptome analysis of rice in mature root tissue by massive parallel sequencing unveiled previously unrecognized tissue-specific expression profiles and render an interesting platform to study the differential regulation of transcribed regions of root tissues. 108 Moreover, this technique foregoes canvas spotlighting in which OsDREB1B (a DREB subfamily gene) was expressed significantly in roots compared to the leaves, shoots, growing points, mature seeds, and other tissues of unstressed rice plants. 109 The subfamily gene, OsAP211, is extremely prominent in shoot tips in mature rice and immature seeds at the booting stage compared to other rice tissues. 110 The ERF subfamily gene, OsAP25, is detected in the leaves, shoots, roots, growing points, flower, immature seeds, and mature seeds of rice. 111 In wheat, 9 types of tissues are used for expression profile analysis and the transcripts of these genes are most abundant in leaves followed by roots, seeds, and stem. 107
Chromosome position of the identified rice AP2/ERF genes
To examine the genomic distribution of AP2/EREBP genes on rice chromosomes (chr), we identified their positions using a Rice TOGO Browser database search. 112 A total of 170 rice AP2/EREBP genes were localized on the 12 chromosomes with an uneven distribution. A similar situation also occurred with respect to the OsERF and AtERF family genes, thereby indicating that ERF genes are distributed widely among monocot and eudicot genomes. 10 The OsAP2/EREBP genes are present in all regions on a single chromosome (eg, at the telomeric ends, near the centromere, and in between) and are distributed individually or in clusters (Fig. 13). Chr2 has the maximum number (26) of OsAP2/EREBP genes, and chr4 and chr6 have 22 and 20 genes, respectively. The high number of AP2/ERF sequences in these chromosomes is due to the repetition of adjacent genes. Interestingly, the same tendency was found for the chromosomal location of group-II (RAV binding domain) genes. Only four OsAP2/EREBP genes were identified on chr11, all of which are found on the short arm. Two OsAP2/EREBP genes (Os07g0410300 and Os07g0410700) arranged as tandem duplications were found around the centromere on chr7. Less than 10 chromosome locations are found on each of chr10, chr11, and chr12.

The physical location of the AP2/ERF genes on rice chromosomes.
Comparative mapping between rice, wheat, and Arabidopsis genomes
To explore orthologs/homologs, wheat and Arabidopsis sequences were used to BLAST against the rice genome using BLASTx and BLASTn, respectively. The aligned sequences with more than 60% similarity to rice genes elsewhere in rice genome produced high-scoring pairs (HSPs). These HSPs were selected based on similarity percentage criteria (Supplementary Tables S4 and 5), which facilitate the differentiation of orthologous regions in the genome with high sureness and, subsequently, describes shared duplications between these genomes. These syntenic surveys facilitate finding different orthologous/homologous gene annotations in the comparative study of rice, wheat, and Arabidopsis genomes.113–117
In wheat (Triticum aestivum L.), AP2/ERF family genes were compared to rice orthologous loci in the rice genome on chr1 to chr9, and not chr10, chr11, or chr12. 118 The microcollinearity allowed a higher resolution of distribution in chr9, followed by chr3, chr2, and chr6, while the minimum predicted orthology was found in chr7. Comparison with rice chromosomes predicts 25% gene orthologs in chr9 followed by 13% on chr2 (Supplementary Table S4). CRT/DRE subfamily genes from wheat found syntenic loci on rice on chr1, chr2, chr4, chr6, chr8, and chr9. The majority of these genes were conserved on chr9 followed by chr1 and chr8. In addition, ERF subfamily gene orthlogous loci were found on chr1, chr2, chr3, chr6, and chr9 respective to their similarity, and maximum syntenic regions were mapped on chr2. Regarding the AP2 family with double domains, these genes were distributed on chr3, chr7, and chr8, and the maximum conserved loci were preserved on chr3 (Fig. 14). The comparative distribution of wheat genes to the rice genome clearly designates that rice chromosomes have passed through different phylogenies. 118
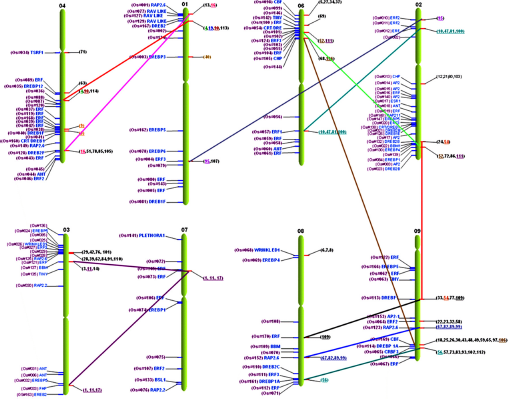
Comparative mapping of wheat AP2/ERF genes on rice chromosomes.
Comparative mapping between rice and Arabidopsis displayed 11 orthologous/homologous loci, unveiling the broad extent of collinearity between these genomes.119,120 The homologs in rice genome were found on chr1, chr2, chr4, chr5, chr6, chr9, and chr11; whereas, in Arabidopsis, this collinearity was found preserved on chr1, chr2, chr3, and chr5. The chromosomal distribution of the AP2/ERF family genes revealed that chr1 in Arabidopsis has more homologs (45%) followed by chr5 (27%). These findings are consistent with the evolution and divergence of the ERF family genes in Arabidopsis. 10 In rice, the conserved homology is found in chr2 (36%) followed by chr4 (27%) (Supplementary Table S5). The ERF subfamily genes of Arabidopsis on chr1, chr3, and chr5 have syntenic loci in rice on chr2, chr4, chr6, and chr9. The genes related to abiotic stresses, such as DRE and TINY on Arabidopsis chr2 and chr5 have their orthologs on rice chr2 and chr4, respectively. The subfamily genes with B3 domains in Arabidopsis found on chr1 have orthologs on rice chr1 and chr5, whereas the subfamily gene with AP2 double domains in Arabidopsis on chr3 is syntenic to rice chr11 (Fig. 15). It is reported that 98% of homologs of known maize, wheat, and barley proteins are found in rice. Synteny and gene homology between monocot (rice and the other cereal) genomes are extensive, whereas synteny with dicots (Arabidopsis) is limited. 121 Moreover, scant collinearity in gene order is also observed between rice and Arabidopsis genomes using comparative genomics. 122

Comparative mapping of rice and Arabidopsis AP2/ERF family.
Conclusions
Transcription factor proteins primarily regulate a web of biological processes and have egressed as a powerful tool for the manipulation of composite metabolic pathways. In this study, 170 AP2/ERF genes are identified from different rice databases available in the public domain. The results reveal much about the diversification of AP2/ERF family genes in the rice genome. Gene translocation and segmental duplication might have imparted towards the expansion of the AP2/ERF gene family. During the enlargement of the AP2/ERF gene family, many groups and subgroups evolved, resulting in a high level of functional divergence. The conserved motifs present in their respective clades suggest the specificity of the genes for function in this group. Comparative mapping revealed that the homologs/orthologs are present in rice, wheat, and Arabidopsis indicating that many of the genes in these species antedated the divergence of monocots and eudicots. The expression analysis of AP2/ERF family genes furnishes a new avenue for functional analyses in rice. During the domestication of rice, selection pressure may lead to selecting the genotypes that have specific conserved motifs and have related molecular functions against natural confronts. Modern bioinformatics and biotechnology tools may predict and characterized these genes in their respective clades. As a model plant, and having a great synteny with the grass family with respect to gene structure, the information generated about the AP2/ERF gene family in rice will also provide a platform for predicting the function of genes of crops whose genome sequences are in their infancy.
Author Contributions
Conceived and designed the experiments: M.R. Analysed the data: M.R. Wrote the first draft of the manuscript: M.R. Contributed to the writing of the manuscript: M.R. Agree with manuscript results and conclusions: M.R., G.H. and Y.G. Jointly developed the structure and arguments for the paper: M.R., J.H. Made critical revisions and approved final version: M.R., X.Y. All authors reviewed and approved of the final manuscript.
Funding
This work is supported by the National Major Specific Project of the People's Republic of China (2011 ZX 08002-004, 2009 ZX 08016-001A, 2009 ZX 08002-013B) and the International Science and Technology Cooperation project of China MoST (2009 DFB 20290).
Competing Interests
Author(s) disclose no potential conflicts of interest.
Supplementary Data
The functional description of conserved motifs found on both sides of the AP2/ERF domain.
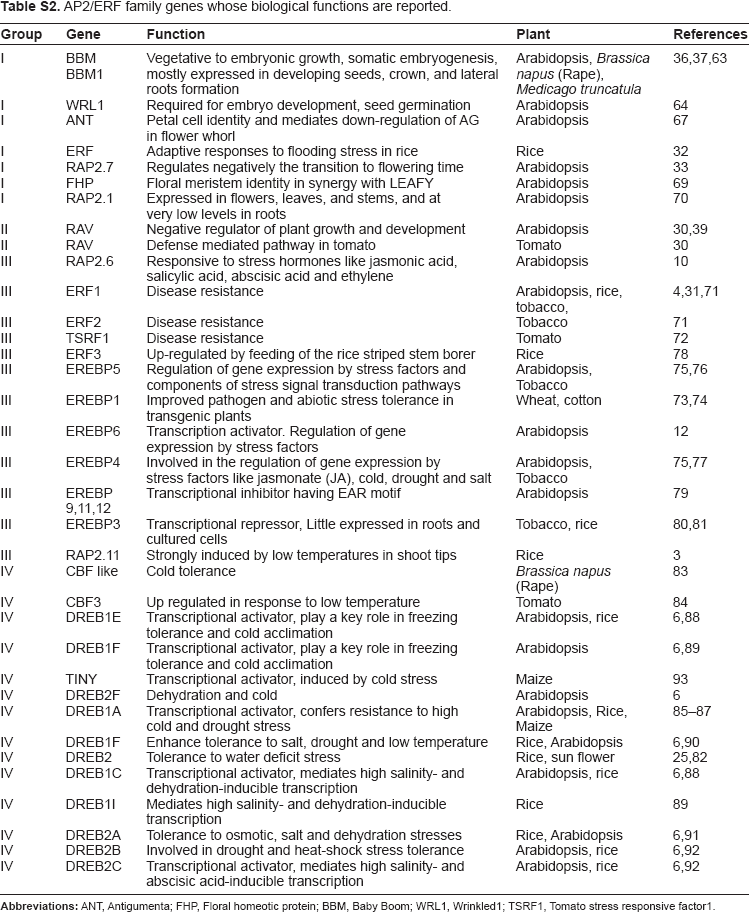
AP2/ERF family genes whose biological functions are reported.
Full length cDNA ascertained with respect to intron, exon size distribution.

Synteny between rice and wheat AP2/ERF family genes.

Synteny between rice and Arabidopsis AP2/ERF family genes.

Conserved motifs discovered using the MEME Suite version 4.7.0.

AP2/ERF family genes in rice.

The OS RAP (ID) number is mentioned because OsAP2/ERF#139 does not have the MSU locus identifier.

Footnotes
As a requirement of publication author(s) have provided to the publisher signed confirmation of compliance with legal and ethical obligations including but not limited to the following: authorship and contributorship, conflicts of interest, privacy and confidentiality and (where applicable) protection of human and animal research subjects. The authors have read and confirmed their agreement with the ICMJE authorship and conflict of interest criteria. The authors have also confirmed that this article is unique and not under consideration or published in any other publication, and that they have permission from rights holders to reproduce any copyrighted material. Any disclosures are made in this section. The external blind peer reviewers report no conflicts of interest.
