Abstract
Riboswitches are regulatory RNA that control gene expression by undergoing conformational changes on ligand binding. Using phylogenetic analysis and comparative genomics we have been able to identify the class of genes/operons regulated by the purine riboswitch and obtain a high-resolution map of purine riboswitch distribution across all bacterial groups. In the process, we are able to explain the absence of purine riboswitches upstream to specific genes in certain genomes. We also identify the point of origin of various purine riboswitches and argue that not all purine riboswitches are of primordial origin, and that some purine riboswitches must have originated after the divergence of certain Firmicute orders in the course of evolution. Our study also reveals the role of horizontal transfer events in accounting for the presence of purine riboswitches in some gammaproteobacterial species. Our work provides significant insights into the origin, distribution and regulatory role of purine riboswitches in prokaryotes.
Introduction
Riboswitches are a type of regulatory ribonucleic acid (RNA) that modulates expression of downstream genes involved in ligand biosynthesis by undergoing a conformational change on ligand binding. They are typically found in the 5' untranslated regions (UTRs) of primarily prokaryotic messenger (m)RNAs. However, some riboswitches have been found in a few eukaryotes as well.1,2 Riboswitches are composed of two domains, an aptamer domain that contains the site for ligand binding, and an expression platform, which can switch between two conformations depending on whether the ligand is bound or unbound to the aptamer domain. They can control gene expression either by termination of transcription3,4 or by inhibiting translation initiation 5 by sequestering the ribosomal binding site in the mRNA. Several classes 6 of riboswitches have been discovered, with the classes being distinguished by the metabolite that binds to the riboswitch. The aptamer domain is highly conserved both at the sequence and the structural level for riboswitches belonging to the same class, whereas the expression platform is highly variable even among riboswitches of the same class. In a recent work, 7 we exploited the high level of sequence conservation of the aptamer domain for a given class of riboswitches in order to develop a fast and accurate method of riboswitch classification based on profile Hidden Markov Models (pHMMs).
Riboswitches regulate various important biochemical pathways in response to the intracellular metabolic concentration, and are widespread in pathogenic bacteria, which makes them a promising drug target.8,9 Natural as well as rationally designed structural analogs that mimic a riboswitch binding ligand have the potential to bind riboswitches and regulate gene expression. Such compounds with antimicrobial action have already been designed for TPP, 10 lysine 11 and purine12,13 riboswitches. The possibility that riboswitches are promising drug targets is further emphasized by the finding that even natural antibiotics can act by targeting riboswitches.14,15 Various riboswitch-based artificial regulatory systems have also been engineered to modulate ligand-dependent gene expression.16,17 Complex riboswitches, which act according to Boolean logics and quick-responding two-way gene control systems have also been designed.18,19 As more classes of riboswitches are discovered, 20 a detailed understanding of the riboswitch structure and function will be crucial in designing riboswitches that can precisely control the concentration of corresponding metabolites and thereby affect the functioning of an organism.
Riboswitches are also speculated 21 to be the remnants of RNA-based metabolite sensors that may have been present in an RNA world. Evidence for that hypothesis requires identifying the point of origin of riboswitches. Moreover, analyzing the distribution pattern of riboswitches across different prokaryotic genomes and the genes they regulate would shed light on the evolution of riboswitches.
The work of Rodionov et al22,23 on the comparative genomics of thiamine biosynthesis and vitamin B12 metabolism has provided considerable insight into the nature of thiamin and cobalamin biosynthesis genes that are regulated by their corresponding riboswitches. More recently, Barrick and Breaker 24 carried out a study of the distribution of various classes of riboswitches across different bacterial groups. However, in order to better understand the origin and evolution of riboswitches, it is essential to first obtain a detailed picture of riboswitch distribution across all prokaryotic genomes. This has to be done not just for each riboswitch class, but also for each distinct gene (or operon) that the riboswitch of a given class regulates. By analyzing this distribution pattern and identifying the genes (or operons) regulated and their role in the metabolic pathways of ligand biosynthesis, it is possible to acquire a better understanding of the role played by riboswitches in gene regulation.
The aim of this paper was to carry out a comprehensive analysis of purine riboswitch distribution across prokaryotic genomes. In the process, we were able to identify the point of origin of the various purine riboswitches, correlate the presence of purine riboswitches with the presence and nature of genes that they regulate, as well as the metabolic pathways to which these genes belong. In some instances, we even found evidence of horizontal transfer of purine riboswitches across distant prokaryotic phyla, which nevertheless share the same environmental niche. Our work provides the first detailed analysis of the origin, evolution and comparative genomics of purine riboswitches.
Methods
Genomic sequence data retrieval and categorization
The Refseq database was downloaded from the National Center for Biotechnology Information (NCBI) FTP site. A total of 646 completely sequenced bacterial genomes were extracted from the Refseq database. Since in this study we are looking at the distribution and evolution of purine riboswitches across different taxonomic groups, the genomes were categorized into different phylums on the basis of taxonomy. Perl scripts were used to extract and categorize the genomic data.
Riboswitch Identification
A profile Hidden Markov Model (pHMM)-based method 7 of riboswitch identification was used to identify riboswitches in the bacterial genomes. A purine riboswitch-specific pHMM 7 was used to screen the bacterial genomes. A systematic analysis of the genomic context of purine riboswitches was carried out to identify the genes upstream to which riboswitches occur. The genomes with missing riboswitches were analyzed in detail to determine the precise cause of the missing riboswitches. The UTRs of the genes with missing riboswitches were also scanned using other riboswitch detection tools like Riboswitch Finder, RibEx, and Covariance Model to make sure that the riboswitches were really absent in such cases and not missed by the pHMM based detection method. This analysis did not reveal any instances of a purine riboswitch that was not detected by our pHMM method.
Phylogenetic analysis
Purine riboswitch identification in bacterial genomes pertaining to different taxonomic groups reveals that they are present predominantly in Firmicutes. With an aim to gain insight on the evolution and point of origin of different purine riboswitches, the riboswitch occurrence information was mapped onto the phylogenetic tree of the organisms belonging to the phylum Firmicutes. Out of the 646 completely sequenced genomes obtained from the Refseq database, 136 belonged to Firmicutes. For phylogenetic tree construction twenty protein families were selected. The choice of the protein sequences was made on the basis of the work of Ciccarelli and colleagues 25 that was aimed at building a highly resolved tree of life. The list of proteins is given in Supplementary file 1. The protein families selected for phylogenetic construction was the subset of the proteins used by Ciccarelli and colleagues. 25 Only those proteins that were found in all the 136 Firmicute species were used to build the sub-set. The twenty clusters of orthologous groups were selected in such a way so as to exclude any lateral transfers, 25 which is essential for obtaining a highly resolved tree.
The protein sequences corresponding to each COG were extracted from all 136 species and aligned using MUSCLE. 26 After alignment, the poorly-aligned regions with more than 20% gaps were trimmed using trimAl. 27 The aligned sequences were concatenated to produce a super-gene alignment of 4975 positions which was then used to build a phylogenetic tree using neighbor-joining (NJ) as well as maximum likelihood (ML) methods. The NJ tree was generated with Phylip version 3.69 28 using a series of sequentially executed programs. Seqboot was used to generate the data sets replicates from the alignment file. Then Protdist generated a distance matrix using the JTT model. The program Neighbor, was used to construct phylogenetic trees from each data set using the Neighbor-Joining method and the consensus tree, was built using the Consense program. The NJ tree was generated for 100 (see Figure 1) as well as 1000 bootstrap replicates (see Supplementary file 2). The two trees are consistent with one another except for some rearrangements in some of the late branches that do not affect our conclusions. This is evident from the mapping of the points of origin of the various purine riboswitches onto the NJ tree with 100 and 1000 (see Supplementary file 2) bootstrap replicates respectively.

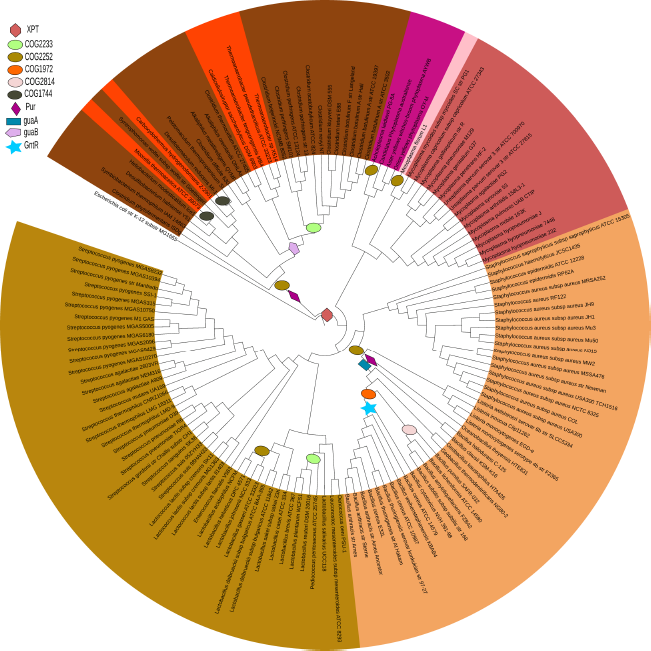
Phylogenetic distribution of purine riboswitches in Firmicutes.
For the ML tree, the best-fit amino acid evolution model was selected for the alignment using the ProtTest3 29 program, which uses PhyML 30 for likelihood calculations with Akaike Information Criterion (AIC). The LG 31 substitution model with invariant sites (I) and four gamma distribution (G4) rate categories was determined to be the most appropriate for the given alignment. Maximum likelihood phylogenetic trees were generated with the substitution model, LG+I+G4 using PhyML version 3. 30 The reliability of the trees was evaluated using 1000 bootstrap replicates. Finally the consensus tree was build using the Consense program from the Phylip package.
The ML tree groups the
The Firmicute tree constructed here separates the different orders clearly with sufficiently high bootstrap values except for the class Clostridia. One reason for this may be that the members of this group are paraphyletic and do not form a phylogenetically coherent group. 33 Some of the late branches have low bootstrap support as can be seen from Figure 1. However, these pertain to evolutionary relationships within specific families, and do not affect the interpretation of our results.
To demonstrate that some of the riboswitches found in Gammaproteobacteria were horizontally transferred from the Firmicute phylum, phylogenetic analysis of the
The mapping of the riboswitch distribution on the phylogenetic tree of Firmicutes was visualized using iTOL. 34
Results
A search of the Refseq database for purine riboswitches found 263 candidates. The distribution of purine riboswitch indicates that they are widespread in Firmicutes but rare in other bacterial groups. In Firmicutes, all classes except Mollicutes use the riboswitch mode of regulation extensively. In certain genera like Bacillus and Clostridium, riboswitches occur multiple times per genome. Figure 1 gives the phylogenetic distribution of all of the purine riboswitches in Firmicutes, and Table 1 indicates the nature of the gene/operon that the purine riboswitches regulate. Analysis of the upstream genes around which purine riboswitches are found reveal that the riboswitch regulation for the purine salvage pathway genes and the transporter genes is widespread in Firmicutes. However, in a few cases, purine riboswitches have also been found to regulate transcription factors as well as the genes belonging to the de-novo synthesis of inosine monophosphate (IMP). Such instances are found primarily in the family Bacillaceae, which has the largest number of purine pathway genes under the riboswitch mode regulation. The riboswitches in these organisms are distributed upstream to a diverse set of genes, which include salvage and de-novo pathway genes as well as permeases, transcription factors and transporters. Organisms from the class Clostridia also regulate a variety of genes comprising transporters, salvage and de-novo genes for IMP synthesis via riboswitch. Riboswitches also regulate guanine monophosphate (GMP) synthesis in many organisms from the Bacillales and Clostridia groups. Streptococcaceae, Lactobacillaceae, Leuconostocaceae,
Gene/operon under riboswitch regulation.

Outside Firmicutes, the presence of purine riboswitches was restricted to few members of the families of Thermotogaceae, Fusobacteriaceae, Shewanellaceae, Vibrionaceae and Bdellovibrionacea. The presence of purine riboswitches in a few organisms belonging to these families can be attributed to horizontal transfer of the purine riboswitch, along with the gene it regulates from the members belonging to the Firmicute group where purine riboswitches are widespread.
In the sub-sections below, we analyze the phylogenetic distribution of riboswitches found upstream to different genes involved in purine metabolic pathway and purine transportation. The riboswitches are named after the gene upstream of which they occur. If a riboswitch occurs upstream of an operon, then it is named after the first gene in the operon.
Xanthine phosphoribosyltransferase (XPT) riboswitch
The

Phylogenetic distribution of the
All the
All the
The distribution of the XPT riboswitch is more continuous in the organisms belonging to the order
Transporter riboswitch
In this section, we discuss the phylogenetic distribution and possible origin of riboswitches that regulate the transporter genes. Transporter proteins responsible for purine uptake are ubiquitous in Firmicutes. Five different transporters were found to be under riboswitch regulation in Firmicutes, namely

Phylogenetic distribution of transporter riboswitches in Firmicutes.
In
The
The distribution of the
Another class of transporter under riboswitch regulation belongs to the COG2252. The genes classified as belonging to the COG2252 are widespread in Firmicutes. COG2252 includes permeases for diverse substrates such as xanthine, uracil and vitamin C. Many members of this family are functionally uncharacterized and may transport other substrates also. All organisms of the order
All Thermoanaerobacterales, except
The organisms from the family
Riboswitches upstream to transporter genes belonging to COG1744 and COG2814 are relatively rare, as is evident from Figure 3.
Pur operon riboswitch
The de-novo biosynthetic pathway for purine nucleotides is highly conserved and well-represented in all the three domains of life, thereby leading to the suggestion that it was present in the last common ancestor.
37
However, the organization of the genes encoding the enzymes for the pathway and their regulation vary.
38
The pathway has been well studied in organisms like
The genes for the de-novo pathway encode the enzymes for the inosine monophosphate (IMP) biosynthesis. IMP acts as the common intermediate in the inter-conversion between adenine and guanine. 40 This enables the cell to maintain the desired composition of the nucleotide pool.
In Bacillus, the de-novo genes for purine biosynthesis are organized as 12-member
The distribution of the

Phylogenetic distribution of the
In
guaA and guaB riboswitches
The de-novo purine biosynthesis reactions convert PRPP to IMP, which is the first purine nucleotide and acts as a common purine precursor. Inosine monophosphate can either synthesize

Phylogenetic distribution of
Clostridiales possess
In Thermoanaerobacterales, only the
GMP synthase gene is subject to the riboswitch mode of regulation in all the organisms belonging to the
GntR riboswitch
GntR is a family of bacterial transcription regulators. The transcription factors of this family have an N-terminal DNA-binding domain, a C-terminal effector-binding domain, and/or an oligomerization (E-b/O) domain. The DNA-binding domain is well conserved. However, the effector-binding domain is variable and heterogeneous, on the basis of which GntR regulators are classified into different subfamilies.
44
The distribution of riboswitches that regulate GntR is shown in Figure 6. In Bacillales, a specific subclade comprising of

Phylogenetic distribution of rare riboswitches in Firmicutes.
Other riboswitches
There were some riboswitches that were not widespread but appeared in a few organisms across the Firmicutes, as shown in Figure 6. All of these riboswitches regulate the genes categorized as the salvage-pathway genes.
Purine
Riboswitches were also found upstream to amidohydrolase (AMH) family protein in
The riboswitch upstream to the
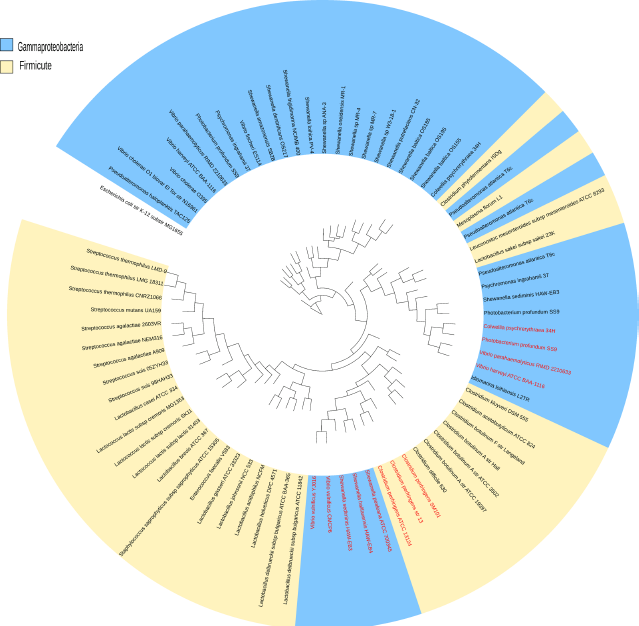
Phylogenetic tree of the adenosine deaminase gene found in Firmicutes and Gammaproteobacteria.
Riboswitch distribution outside Firmicutes
The purine riboswitches outside Firmicutes are scarce and restricted to a few species in the
The purine riboswitch is found upstream to the
Thermotoga lettingae and Petrotoga mobilis from Thermotogaceae family possess ariboswitch upstream of the
A few organisms belonging to the Shewanellaceae and Vibrionaceae family of Gammaproteobacteria also carry the purine riboswitch.
Discussion
It is tempting to attempt to infer the evolutionary origin of the various purine riboswitches, given our knowledge of their detailed phylogenetic distribution. It has been argued21,48 that riboswitches are remnants of primordial regulatory machinery that may have been operational in an RNA world. Therefore, it seems interesting to ascertain whether the origin of the riboswitches can be traced back to the root of the tree of life.
In Firmicutes, the
Possible point of origin of the various purine riboswitches.
The distribution of riboswitches upstream to transporter genes belonging to COG1972, COG2814 and COG1744 indicates they are relatively rare and are likely to have originated in specific clades of the Bacillus genus or a few organisms belonging to Clostridiales and Thermoanaerobacteriales (see Fig. 8) quite late in the evolution of Firmicutes. Riboswitches upstream to the transporter genes belonging to COG2233 and COG2252 are more widespread. Even so, Figure 8 shows the multiple independent origins of the riboswitch upstream to some COG2252 genes, two of which can be placed at the root of the
The purine de-novo genes are found in almost all the organisms belonging to the phylum Firmicutes (except Mesoplasma, Phytoplasma and Mycoplasma). However, only a small fraction of them carry a purine riboswitch upstream to them. It is difficult to infer the origin of the
For the purpose of determining the point of origin of the
The distribution of the riboswitch upstream to the
The presence of the riboswitch upstream to the
Horizontal transfer of riboswitches along with the regulated genes has been well documented.47–50 In our analysis, we found examples of horizontal riboswitch transfer (HRT) from Firmicutes to other phyla. It appears that the
Some of the purine salvage pathway genes, de-novo pathway genes and transporter genes belonging to COG2233 and COG2252 are ubiquitous in all orders of Firmicutes (with the exception of parasitic orders like Mycoplasmatales). Hence, while discussing the evolutionary origin of the corresponding riboswitches, it is difficult to rule out the possibility that these riboswitches originated (along with the genes or operons they regulate) at the root of the Firmicute phylum (or perhaps even earlier) but were eventually lost, sometimes along with the gene (or operon), in some groups of organisms during the subsequent evolution of Firmicutes.
Another question that is raised by our analysis deals with the extent to which riboswitches are essential for the functioning of the organism. If a purine riboswitch was lost in several lineages or groups without having an adverse effect on the viability of the organisms, then it is likely that those organisms possessed alternative means of regulating purine metabolism, and did not need to rely exclusively on riboswitches.
Conclusions
Our work on the detailed distribution of purine riboswitches reveals that it regulates a wide variety of genes, ranging from purine biosynthesis genes to transporters and transcription factors. Hence, the evolutionary origin of the purine riboswitches has to be considered in the context of the many different types of genes that are regulated by these riboswitches. Our analysis suggests that the origin of the purine riboswitch upstream to the
Funding
The work was funded in part by a grant given to SS by the Department of Biotechnology (DBT), Government of India.
Competing Interests
Author(s) disclose no potential conflicts of interest.
Author Contributions
PS contributed to the design of the study, wrote the programs, analyzed the data and wrote the manuscript. SS contributed to the design of the study, analyzed the data and wrote the manuscript. All authors read and approved the final manuscript.
Supplementary Data
gi|116491818:1818151-1819476 is the identifier indicating the gene coordinates of the pbuX gene that possesses the riboswitch. gi|116491818:c1427741-1426431 is the identifier indicating the gene coordinates of
(a) Sequence alignment of COG2252 genes in
Footnotes
Acknowledgements
We thank Sudha Bhattacharya and L. Aravind for valuable discussions.
As a requirement of publication author(s) have provided to the publisher signed confirmation of compliance with legal and ethical obligations including but not limited to the following: authorship and contributorship, conflicts of interest, privacy and confidentiality and (where applicable) protection of human and animal research subjects. The authors have read and confirmed their agreement with the ICMJE authorship and conflict of interest criteria. The authors have also confirmed that this article is unique and not under consideration or published in any other publication, and that they have permission from rights holders to reproduce any copyrighted material. Any disclosures are made in this section. The external blind peer reviewers report no conflicts of interest.
