Abstract
Epitope mapping allowed the location of antigenic determinants on a protein macromolecule to be identified. In particular, pepscan techniques that utilize a series of overlapping peptides, help detect key amino acid residues that are important in antibody recognition and binding. In a previous study, we employed 15-mer peptides spanning the entire length of IgG1Fc to ascertain successfully the target epitopes of isotypic/allotypic monoclonal reagents.
As an extension to this work we have used these peptides to evaluate the location of epitope targets of five IgM rheumatoid factor antibodies (RFAbs). Overall, 2 antibodies, RFAb TS2 and TS1, detected a similar epitope within the CH3 domain (360-KNQVSLTCLVKGFYP-374), whilst 1 (RFAb SJ1) recognised an epitope in the CH2 domain (294-EQYNSTYRVVSVLTV-308). In contrast, 2 RFAbs, PRSJ2 and PRTS1 detected four and five epitopes respectively within the Fc region. RFAb PRSJ2 recognised epitopes detected by RFAB TS2 and TS1 but also further epitopes in the CH2 domain (256-TPEVTCVVVDVSHED-270) and CH3 domain (418-QQGNVFSCSVMHEAL-432). Similarly, RFAb PRTS1 detected all four epitopes plus a fifth in the CH3 domain (382-ESNGQPENNYKTTPP-396). In essence there was a consensus of target epitopes identified by these rheumatoid factor antibodies. Interestingly, two epitopes (256–270, CH2 domain and 360–374, CH3 domain) were novel in that they had not been identified in previous pepscan studies. The other epitopes recognised, either overlapped or were immediately adjacent to previous epitopes detected by poly/monoclonal rheumatoid factor antibodies.
Molecular modelling (PCImdad) of IgG1Fc showed that all five epitopes were exposed and surface accessible for antibody interaction. In addition, a bioinformatics analysis of the Fc region using ExPASy was employed to identify key antigenic determinants. This ‘in silico’ approach may provide a means of determining key regions without the need to develop overlapping peptides spanning the entire length of a macromolecule.
Introduction
Rheumatoid arthritis (RA) is an autoimmune disorder, with chronic inflammation of the synovial membrane culminating in the pathological destruction of cartilage and bone in affected joints. Overall the resultant pain, loss of function and disability causes significant morbidity and mortality. The aetiology of RA remains unclear, although a genetic predisposition, together with other triggers including viral and bacterial agents are considered to be contributory factors.
Rheumatoid factors (RFs), are detected in a large proportion of RA patients and provide a key serological marker for diagnosis [1]. RF antibodies have been shown to target the Fc region of IgG [6], typically the CH2/CH3 inter-domain region by inhibition studies, enzyme degradation and site-directed mutagenesis. They are mainly low affinity IgM isotype, but may also be high affinity IgG and IgA isotypes. RFs are naturally-occurring autoantibodies that are also found in healthy individuals where they are considered to facilitate competitive tolerance [3], increase the cytotoxicity of antiviral antibodies [4] and aid immune complex formation and presentation/elimination by phagocytic cells [5].
Epitope mapping studies have contributed to the identification of potential antigenic sites of rheumatoid factors in the pathological mechanisms of rheumatoid arthritis. Crude techniques such as cyanogens bromide cleavage and enzyme degradation, have permitted the localisation of antibody reactivity to given immunoglobulin fragments. But the development of pepscan methodologies, which use a series of overlapping synthetic peptides, have provided a systematic approach for identifying key amino acids in antibody recognition and binding. Table 1 summarises a number of eloquent studies [7–11] that have used polyclonal/monoclonal rheumatoid factors which highlight CH2 and CH3 target epitopes on IgG1 Fc. Moreover, Table 1 gives details of three antigenic regions on IgG1 Fc previously identified through pepscan by monoclonal anti-IgG reagents MAbs: G7C, PNF69C and A57H) [7,9]. Interestingly, these epitopes when visualised through molecular modelling [1] were adjacent to, or overlapped with, amino acids identified by polyclonal rheumatoid factors.
Rheumatoid factor and anti-Fc target epitopes on IgG1.

Amino acids presented in single amino acid code plus Eu reference numbering.
As an extension to our work, we have undertaken pepscan analysis of five Fc-binding rheumatoid factor antibodies [12] and compared their reactivity against previously identified RF epitopes. A molecular modelling programme (IMDAD) [7] was used to locate the potential target epitopes of these antibodies as antigenic ‘hot spots’ within the IgG1 Fc macromolecule. It is recognised that pepscan remains an expensive investigative tool. Consequently, we evaluated a bioinformatic approach [13,14] to investigate potential antigenic targets. Overall, ‘in silico’ analysis favourably highlighted epitopes identified by pepscan studies.
The rationale for these analyses has been to identify possible unique or overlapping regions of antigenicity on IgG Fc region with the view to facilitating possible peptides for therapeutic uses.
Materials and Methods
Rheumatoid Factor Antibodies
Rheumatoid factor IgM antibodies (RFAbs) were kindly provided by Professor JB Natvig (Rikshospitalet University Hospital, Institute of Immunology, University of Oslo, Norway). Details of the characterisation and purification of these antibodies TS2, TS1, SJ1, PRSJ2 and PRTS1 has previously been reported [12]. In particular RFAbs TS2, TS1 and SJ1 were monospecific with regard to their reactivity against native IgG, whilst RFAbs PRSJ2 and PRTS1 were polyspecific.
Synthetic Peptides
Ninety four overlapping 15-mer peptides corresponding to the Fc region of IgG1 NIE were kindly provided by Mimotopes Pty Ltd. Peptides were initially reconstituted in DMSO and then diluted in citrate phosphate buffer as previously described [7]. Peptides spanned the CH2 and CH3 domains, beginning 238-PSVFLFPPKPKDTLM-252 and terminating 424-FSCSVMHEALHNHYT-438, with an offset of 2. All amino acids were related to the Eu protein numbering index [15].
Pepscan Assay
An ELISA based assay for each overlapping peptide was used as previously described [7]. In brief, biotinylated peptides were coated onto streptavidin coated plates, blocked with bovine serum albumin in 0.1 M PBS pH 7.2 and incubated with purified antibodies PRSJ2, PRTS1, TS2, TS1 and SJ1 diluted to the following optimal working concentrations (7.3, 3.1, 46, 38 and 2.1 μg/ml) respectively. Following washing and addition of anti-human HRP-conjugate and then substrate, plates were read at 405 nm (Titertek Multiscan). Optical density values were expressed as 0–2 OD Units. The OD values obtained from control RFAbs (SJ3, TS3, KL2 [12]) that exhibited minimal activity in pepscan, were used to determine a mean value for each peptide which was subtracted from all other pepscan profiles. Pepscan profiles for controls highlighted background activity: mean 0.05 OD Units. Control isotype matched RFAbs have similarly been shown to bind to native IgG [12].
Pcimdad Display
A PC interactive molecular display and design programme (PCImdad) was kindly provided by Dr. Michael Levitt and used as previously described [7]. The software utilised the Brookhaven Protein Data Bank (PDB files) for the coordinates of macromolecules. In brief, the PDB file 1Fc1 (Deisenhofer IgG1 Fc sequence [16]) was selected and displayed in space-filling mode. Key amino acids derived from pepscan, bioinformatic analysis and published data were then highlighted using the command language provided by the manufacturer (M. Levitt, Molecular Applications Group, Stanford University, U.S.A). Overall both Deisenhofer IgG1 Fc and protein Fc NIE were broadly similar in sequence but with variation in allotype; IgG1 Deisenhofer Fc: non-G1m(a), NIE Fc: G1m(a).
Bioinformatic Analysis
The protein sequence of IgG Fc (accession number AF150959-1) was extracted from the NCBI/Genbank online database (www.ncbi.nlm.nih/). In silico analysis was performed using the biocomputing software programmes available online at http://www.expasy.ch/cgi-bin/protscale.pl. The algorithms used included hydrophilicity, residue polarity, solvent accessibility and flexibility index for primary structure. Threshold values were determined as the top 25%, 30% and 40% of peaks as suggested by Alix 1999 [17]. In addition, propensity for β-turns was evaluated using BetatPred2 available at http://imtech.res.in/raghava/betaturns. Using these parameters a region was considered antigenic when four or more of these properties were above threshold values or showed evidence of β-turns.
Results
Pepscan Profiling
All five rheumatoid factors (RFAbs) generated pepscan profiles when using IgG1 Fc overlapping peptides. RFAb TS2 (Fig. 1a) showed reactivity to an antigenic region within the CH3 domain encompassed by peptides 61 and 62 (358-LTKNQVSLTCLVKGFYP-374) (Table 2). A maximum signal intensity of 0.2 OD units was observed to peptide 62. In the case of RFAb TS1, an identical reactivity profile was shown (Fig. 1b) but with a weaker signal intensity of 0.12 OD units to peptide 62. Furthermore it was noted that RFAb TS1 showed reactivity (0.25 OD units) to peptide 85: residues 406-LYSKLTVDKSRWQQG-420. RFAb SJ1 (Fig. 1c) showed reactivity to a single region within the CH2 domain. Our data highlighted that RFAb SJ1 identified a potential antigenic region bound by peptides 28–31 that included residues 292-REEQYNSTYRVVSVLTVLHQD-312. The highest signal intensity (0.14 OD units) was observed for peptide 29 (Table 2). Overall, antibodies TS2, TS1 and SJ1 appeared to be specific for a single epitope.

Pepscan profile of RFAbs TS2 (a), TS1 (b), SJ1 (c), PRSJ2 (d), PRTS1 (e). Vertical axis: optical density (0–2 OD) units (note scale on figures highlight 0–2000), horizontal axis: peptide number.
Comparison of RFAb reactivity against IgG1 peptides.

Peptides are numbered according to the EU protein index. + highlights reactivity against peptides ascertained through pepscan, + indicates highest OD value exhibited by a given rheumatoid factor antibody (RFAb). Overall five peptide regions are observed (highlighted in grey).
In contrast, RFAb PRSJ2 showed antibody reactivity to four potential antigenic regions (Fig. 1d) that encompassed residues 252-MISRTPEVTCVVVDVSHEDPEVK-274 (CH2 domain), 290-KPREEQYNSTYRVVSVLTVLHQDWLNG-316 (CH2 domain), 360-KNQVSLTCLVKGFYPSD-376 (CH3 domain) and 418-QQGNVFSCSVMHEALHN-436 (CH3 domain). For this antibody, maximum signal intensities of 0.74, 0.15, 0.34 and 0.94 OD units were observed to peptides 10, 29, 63, and 91 respectively. Likewise RFAb PRTS1 broadly exhibited a similar reactivity profile to these four regions (Fig. 1e), but with generally lower OD values: 0.44, 0.29, 0.42 (for peptide 62) and 0.14 OD units. However, RFAb PRTS1 also recognised a fifth antigenic region. The latter was detected in peptide 73: 382-ESNGQPENNYKTTPP-396, with a signal intensity of 0.12 OD units.
When taken together our data (Table 2) shows a broad consensus of 4 potential antigenic regions or epitopes that were identified by our RFAbs. Clearly a fifth epitope was solely identified by RFAb PRTS1. In summary, the following peptides were considered potential target epitopes and designated (i) 256-TPEVTCVVVDVSHED-270, (ii) 294-EQYNSTYRVVSVLTV-308, (iii) 360-KNQVSLTCLVKGFYP-374 (vi) 382-ESNGQPENNYKTTPP-396 and (v) 418-QQGNVFSCSVMHEAL-432.
Pcimdad Display of Key Antigenic Sites
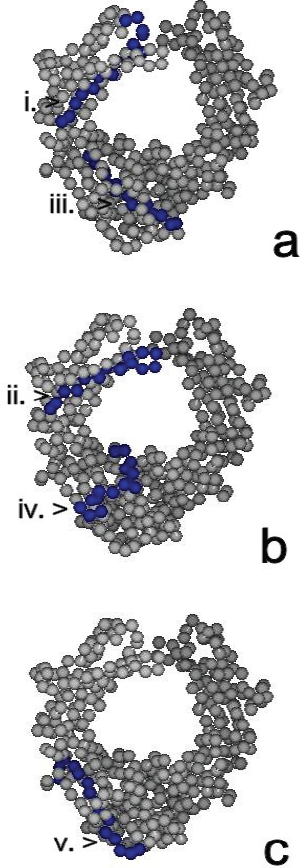
The location and surface topography of RFAb target epitopes identified through pepscan in this study were visualised on an IgG molecule using the space-filling mode (Fig. 2a, b, c). The sequence (i) 256-TPEVTCVVVDVSHED-270, was present on an exposed segment of the fx2 β-pleated face and b2 bend (or loop) of the CH2 domain whilst sequence (ii) 294-EQYNSTYRVVSVLTV-308, was located on the b4 loop that bridged the fx3 and fx4 faces of the CH2 domain. Both sequences were solvent accessible. Sequences (iii) 360-KNQVSLTCLVKGFYP-374, (iv) 382-ESNGQPENNYKTTPP-396 and (v) 418-QQGNVFSCSVMHEAL-432, were present on exposed segments on the CH3 domain. Sequence (iii) was located on part of the b1 loop and to a large extent on the fx2 face whereas sequence (iv) was located largely on the b3 loop and a small segment of the fx3 β-pleated face. Sequence (v) was located at the C-terminal of the IgG1 Fc macromolecule and included part of the fy2 face and b6 segment of the CH3 domain. All regions were thus capable of interaction with antibody/rheumatoid factors.
Bioinformatic (In Silico) Analysis
In silico analysis of IgG1 Fc highlighted a total of 11 epitopes based on four out of five parameters being satisfied. Overall the threshold applied for plots of hydrophilicity, accessibility, flexibility and polarity determined the number of additional peptides that could be highlighted as potentially antigenic (Table 3). At 25% (high stringency) (Alix, 1999 [17]), 5 antigenic regions were identified whereas at 30%, 3 further epitopes were identified. An additional 3 epitopes were highlighted when a low threshold was applied (40%).
Bioinformatic analysis of IgG1Fc.

Peptides are numbered according to the EU protein index.
Discussion
Rheumatoid arthritis (RA) affects 0.5%–1% of the British population with about 75% of patients being rheumatoid factor (RF) positive [1,18]. Clearly RF is not exclusive to RA but found in other diseases such as Sjogren's syndrome and mixed cryoglobulinemia, and shown to be transiently increased in a number of infectious diseases [19,20]. Here, it is possible that RF may have a beneficial role in the resolution of disease. In the acute stage of RA, IgM RF is produced that is of low affinity and as the disease progresses, class switching occurs (to IgG or IgA) together with somatic mutation of complimentary determining residues yielding high affinity RF antibodies. Overall, these facts suggest an antigen-driven immune response. The binding of RFs to IgG has been located to the CH2/CH3 interdomain region that is also associated with the docking of proteins such as Staphylococcus protein A. In the case of RF-AN (derived from an EBV transformed B cell from an RA patient), the reactivity is isolated to IgG1, 2 and 4 suggesting the importance of histidine 435 in the Ga target epitope [21].
Epitope mapping studies have provided a substantial list of Fc-related peptides that could be important for RF binding [7–11]. In the present study, we have used 15-mer overlapping peptides of IgG1 Fc to explore the reactivity of five IgM RFs and identify a consensus of potential RF amino acid regions/epitopes. Three antibodies exhibited specificity to a single domain: both RFAbs TS2 and TS1 to a region in the CH3 domain, epitope (iii) 360-KNQVSLTCLVKGFYP-374 and RFAb SJ1 to a region in the CH2 domain, epitope (ii) 294-EQYNSTYRVVSVLTV-308. Overall, this data was consistent with the monospecificity of the rheumatoid factors [12]. Interestingly epitope (iii) was adjacent to amino acids previously cited RF antigenic sites but novel in terms of its position (Fig. 3) and was present in IgG1, 2, 3 and 4 subclasses. In contrast epitope (ii) highlighted amino acid variations between IgG subclasses and was located in a region previously identified by polyclonal and monoclonal rheumatoid factors. RFAb PRSJ2 showed strong reactivity for epitopes (ii) and (iii), but also highlighted reactivity for epitopes (i) 256-TPEVTCVVVDVSHED-270 and (v) 418-QQGNVFSCSVMHEAL-432. Again epitope (i) was novel in terms of its CH2 position in comparison to previously cited RF epitopes and was distal to lower hinge region residues associated with Fc-receptor (FcγRI) binding (234-LLGG-237) [22,23] (Fig 3). Epitope (v) was located at the C-terminal region of the CH3 domain and in a zone previously shown as a potential site for RF activity. RFAb PRTS1 showed lower reactivity in terms of OD units as compared to RFAb PRSJ2 and exhibited cross-reactivity to epitopes (i), (ii), (iii), (v) plus epitope (iv) 382-ESNGQPENNYKTTPP-396. Again the pepscan profiles of both IgM antibodies were consistent with the polyspecific nature identified by Soltys et al. 1994 [12]. It is plausible that these antibodies are of low affinity and may explain the cross-reactivity exhibited in our study. Epitope (iv) has not previously been identified by rheumatoid factor antibodies but by a monoclonal antibody, A57H, generated from a mouse immunized with a purified IgG1 Fc myeloma protein [9]. Clearly a caveat of pepscan is that it provides a picture of antibody reactivity to linear stretches of amino acids. Evidently a given antibody may recognise adjacent peptides (possessing a particular amino acid motif), however there also examples of antibodies (e.g. monoclonal antibody HP6069 [11]) that recognise more that one region. This could be due to a number of factors including low affinity interactions, recognition of similar structures (despite different amino acids) [24] and the recognition of epitopes not usually exposed or hidden in the native protein. It is also possible that the assay system and orientation of peptides could contribute to unexpected reactivity or indeed poor reactivity for some peptides.

Epitopes identified through bioinformatics and Pepscan. Key: bioinformatic epitopes at 25% threshold XX (boxed, shaded) and at 30% YY (boxed, non-shaded). XX = reactivity of RFAbs in this paper highlighted on protein NIE only. X (italic) = nuance between Eu and NIE. ∗∗ reactivity of polyclonal and ++ monoclonal RFs from previous studies [7–10]. IgG proteins: Cγ1a reference protein Eu IgG1 Fc accession number AF150959–1, Cγ1b protein NIE (used for pepscan).
Molecular modelling shows that all five RFAb epitopes were located on exposed surfaces on IgG Fc and accessible for antibody interaction (Fig. 2). The opportunity to highlight key residues provides a useful adjunct to epitope mapping in providing their location in a three-dimensional setting. Interestingly epitope (ii) possessed the amino acid asparagine 297 that is associated with glycosylation of IgG [25] (Fig. 3). The fact that this antibody bound to IgG1 in vitro suggests that the epitope was not masked or sterically hindered by the attachment of carbohydrate moieties. Epitope (iii) was proximal to the G1m(a) allotypic region and epitope (v) included the non-G1m(x) residue, alanine 431, and was on the same β-strand as histidine 435 (Fig. 3). Both the G1m(a) motif, 356-DELTK-360, alanine/glycine 431 and histidine 435 residues are solvent accessible [7,26]. Overall, for all five RFAb epitopes there were few amino acid variations between isotypes suggesting that these antibodies would bind to all four subclasses.
Bioinformatic analysis uses computational algorithms to predict regions of hydrophilicity, accessibility, polarity, flexibility and β-turns that in turn may identify potential antigenic regions of proteins. This in silico approach has been successfully used to determine likely antigenic ‘hot spots’ on auto-antigens such as cardiac myosin [27], and antigenic sites on viruses [13]. In adopting this approach, we noted a degree of plasticity in peptide regions selected using ExPASy depending on the stringency (or threshold) set. At a threshold of 25%, as initially highlighted by Alix (1999), 5 peptides were identified of which 3: 286-NAKTKPREEQYNSTY-300, 349-YTLPPSRDELTKNQVS-364 and 263-VVDVSHEDPEVKFNW-277 either overlapped or were a few residues upstream or downstream to RFAb epitopes (ii), (iii) and (i) respectively. At a threshold of 30%, 3 additional epitopes were identified that overlapped with RFAb epitopes (ii) and (v). Again a number of epitopes selected through bioinformatics were identified as key epitopes by mouse MAbs G7C, PNF69C and A57H. When a low threshold was applied (40%), further epitopes (391-YKTTPPVLDSDGPFF-405 and 433-HNHYTQKSLSLSPGK-447) were identified that overlapped to some extent with poly/monoclonal RFs from previous studies. In a recent study on the human immunodeficiency virus (HIV-1), bioinformatics analysis was used to identify key peptides that were synthesised and assessed against HIV-1 positive serum. In this particular study, the sera of HIV-1 patients bound to a large proportion of peptides, although a few peptides were negative. Unfortunately no mention of the stringency applied in the selection of peptides was reported. In our proof of principle study, a threshold of 30% appeared to be an optimal ‘benchmark’ for highlighting crucial epitopes on IgG1 Fc. With less stringency, other peptides may be highlighted that could be of less importance in vivo e.g. epitope 324-SNKALPAPIEKTISK-338 exhibited no overlap with RF epitopes. Overall, bioinformatics may provide a guide of antigenic regions that are likely to be found in vivo with the caveat of a suitable threshold to exclude unnecessary peptides. This approach could provide a way forward in selecting key epitopes on large macromolecules e.g. autoantigenic epitopes in RA and systemic lupus erythematosus, without the need (and expense) for numerous overlapping peptides spanning the entire length of a protein [28,29].
In essence, epitope mapping and molecular modelling provide a useful means of identifying key epitopes for antibody-antigen interaction and binding. Our study differed from some previous pepscan studies in employing 15-mer (as compared to 7-mer) overlapping peptides. It is plausible that longer peptides may provide better opportunities for displaying epitopes since some conformation, in the context of the native protein, could be adopted. In addition we used the strategy of biotinylated peptides. Again this ensured maximum display of peptide as opposed to using hydrophobic interactions of peptide and plastic plate that could mask the salient amino acid residues. Clearly further work using shorter peptides and inhibition studies could help further refine the RFAb epitopes identified. It is possible that the peptides identified may have some potential as therapeutic agents [30] should they prove to modulate RF production [31]. It would be interesting to evaluate IgG RFAb's to evaluate an increased reactivity/affinity to similar or other epitopes. Evidently the spectrum of epitopes might alter on IgG, and other macromolecules in the pathogenesis of RA, due to epitope spreading [32,33]. What causes the initial antigenic insult in RA remains unknown, although there is a possible mechanism of molecular mimicry [34]. Preliminary alignment analysis (using ExPASy) shows that some RFAb epitopes (i), (ii) and (iv) share some homology with human endogenous retroviruses (HERVs): TPEAT epitope (i), NGQP epitope (iv) with HERV-K10 and NQNSLTC epitope (iii) with HERV-L. Interestingly HERV-K10 has been implicated in RA and SLE [35,36,37]. This area warrants further investigation plus an assessment of cross-reactive epitopes with microbes and autoantigens. In addition, the application of bioinformatics to identify key antigenic regions on host proteins and possible microbes may prove useful in other diseases such as multiple sclerosis and systemic lupus erythematosus.
Footnotes
Acknowledgements
The authors would like to thank Gordon Tribbick and Mimotopes Pty Ltd. for the provision of synthetic peptides, P. Rajani for laboratory assistance and the James Beattie Charitable Trust, Wolverhampton, The Lions Clubs of District 105 M (West Midlands) U.K, South Staffordshire Medical Foundation, Wolverhampton, and the Research Institute in Healthcare Science, University of Wolverhampton for the provision of laboratory space and reagents.
