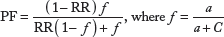Abstract
Apolipoprotein E (APOE) is a glycosylated protein with multiple biological properties. APOE gene polymorphism plays a central role in lipid metabolism and has recently been suggested to regulate inflammation. Our objective is to evaluate whether APOE polymorphism affects susceptibility to SLE. APOE genotyping was performed using ApoE StripAssay™ kit. Results indicated significantly higher frequencies of allele ∊4 and genotype ∊3/∊4 and lower frequencies of allele ∊3 and genotype ∊3/∊3 in SLE patients than controls. APOE ∊2 allele was found in three patients, whereas it was absent in controls. The frequencies of allele ∊4 and genotype ∊3/∊4 were significantly higher in SLE patients with renal involvement and those of alleles ∊2, ∊4 and genotypes ∊2/∊3, ∊3/∊4 were higher in patients with neuropsychiatric symptoms. It is concluded that APOE allele ∊4 is associated with susceptibility risk/clinical manifestations of SLE and ∊2 may increase its severity while ∊3 is protective for SLE in Saudis.
Introduction
Systemic lupus erythematosus (SLE) (MIM 152700) is a systemic autoimmune disease, characterized by the production of a large number of autoantibodies that potentially drive immunocomplex-related inflammation in various tissues and organs. 1 Nearly every organ system of the body can be affected by SLE. The clinical manifestations vary according to the organs attacked. It primarily affects premenopausal women and is characterized by a number of clinical manifestations.2–4
SLE is a chronic disease course marked by remissions and relapses, which is characterized by sudden flare-ups and repeated life-threatening infections, mild-to-severe kidney impairment, repeated miscarriages, strokes, and sometimes permanent disability and death.5–7 Incidence of SLE varies in different parts of the world and often varies with age, sex, and race. 8 The reported prevalence of SLE in various populations is 20–150 cases per 100,000.8–10 The prevalence of SLE in the central region of Saudi Arabia has been estimated to be 19.28 per 100,000 population. 11 The etiology of SLE is multifactorial, and many reports suggest a role for genetic, hormonal, immunological, and environmental factors.12,13 The abnormal lymphocyte apoptosis, decreased clearance of activated T cells, and involvement of multiple cytokines have been demonstrated with the pathogenesis of SLE.14–16 SLE is currently believed to be a T cell-driven condition, and indeed, targeting molecules expressed on T cells and their signaling pathways can be one of the potential therapeutic strategies in SLE. 17 Apolipoprotein E (APOE) is a 34.2-kDa glycosylated protein with multiple biological properties. As originally described in the context of cholesterol metabolism, APOE plays a key role in inhibiting the proliferation of T lymphocytes, regulating immune reactions, and interacting with several cytokines.18–20 Recently, Song et al. 21 reported a correlation of APOE with disease activity and related cytokines in SLE. A polymorphism in the fourth exon of the APOE gene located on the long arm of chromosome19 determines the three common alleles (∊2, ∊3, and ∊4) in human population. The presence of three alleles leads to the formation of six different genotypes, namely ∊2/∊2, ∊2/∊3, ∊2/∊4, ∊3/∊3, ∊3/∊4, and ∊4/∊4. The frequency distribution of these alleles and genotypes of APOE varies in different ethnic populations.22–24 APOE alleles/genotypes have also been associated with various autoimmune diseases.25–29 Pullmann et al. 30 reported an association of APOE ∊4 allele with neuropsychiatric SLE in Caucasians from Slovakia. In view of scarcity of data on the association between APOE alleles/genotypes and SLE, this study has been designed to investigate the association of APOE polymorphisms with SLE susceptibility in Saudi patients.
Methods
Subjects
A total of 318 subjects including 118 SLE patients and 200 age- and sex-matched healthy controls were recruited from the outpatient rheumatology clinic of Prince Sultan Military Medical City, Riyadh, Saudi Arabia, between 2007 and 2010. All the subjects were biologically unrelated Saudis. The patients were diagnosed for SLE using the revised criteria of American College of Rheumatology. The diagnostic reliability was achieved by conducting a systemic search into the case notes of the patients. The confirmed 118 SLE cases included 106 females and 12 males with a mean age of 35.19 ± 14 years and a disease duration of 1–23 years (mean 7.23 ± 6.3 years). Age of onset of SLE ranged from 22 to 60 years. The control group consisted of 150 females and 50 males with a mean age of 36 ± 10 years. SLE patients with neuropsychiatric manifestation were classified according to the criteria of the American College of Rheumatology research committee. 31 Controls subjects less than 21 years old and having a first- or second-degree relatives with SLE or any rheumatic/autoimmune disorders were excluded. This study was approved by the ethical committee of the Prince Sultan Military Medical City and written informed consent was obtained from each participant. The research was conducted in accordance with the principles of the Declaration of Helsinki.
Genotyping
Genomic DNA was extracted from the blood of SLE patients and controls using QIAamp¯ DNA mini kit (Qiagen CA). The genotypes of the APOE polymorphisms were determined using ApoE StripAssay™ kit based on polymerase chain reaction (PCR) and reverse-hybridization technique (ViennaLab Labordiagnostika GmbH). To cross-check the results of the genotyping, the APOE polymorphism was also determined by PCR and restriction fragment length polymorphism (RFLP) technique. PCR was performed using PuRe Taq™ Ready-To-Go™ PCR Beads (GE Healthcare) with primers as described elsewhere.32,33
Statistical analysis
Frequencies of various alleles and genotypes of APOE polymorphisms were compared between SLE patients and controls and analyzed by Fisher's exact test, and the P ≤ 0.05 value was considered as significant. The strength of the association of disease with respect to a particular allele/genotype is expressed by odds ratio interpreted as relative risk (RR) according to the method of Woolf as outlined by Schallreuter et al. 34 Chi-square test was used to calculate 95% confidence interval. RR was calculated only for those alleles/genotypes, which was increased or decreased in SLE patients when compared with the control group. The RR was calculated for all the subjects using the formula given below:
a is the number of patients with expression of allele or genotype; b is the number of patients without expression of allele or genotype; c is the number of controls with expression of allele or genotype; and d is the number of controls without expression of allele or genotype.
The etiological fraction (EF) indicates the hypothetical genetic component of the disease. EF values of >0.00–0.99 are significant. It is calculated for positive associations (RR > 1) using the following formula proposed by Svejgaard et al. 35
where
Preventive fraction (PF) indicates the hypothetical protective effect of one allele/genotype for the disease. It is calculated for negative associations (RR < 1) using the following formula 35 :
Values of <1.0 indicate the protective effect of an allele/genotype against the manifestation of the disease.
Results
The demographic and clinical features of SLE patients have been summarized in Table 1. Among the SLE patients, a wide range of manifestation were observed including renal involvement, fever, neuropsychological disorder, malar/butterfly rash, lung involvement, photosensitivity, cardiac involvement, and oral ulcers. The results of the laboratory investigation showed that SLE patients suffered from high-erythrocyte sedimentation rate and high C-reactive protein (CRP), thrombocytopenia, leucopenia, and lymphopenia.
Demographic and clinical features of SLE patients.

including bacteria, virus and fungi.
The results of APOE genotyping in the SLE patients and the control subjects are shown in Tables 2–4. We found that frequencies of various alleles and genotypes of APOE were significantly different between the healthy controls and SLE patients (Fig. 1). The frequency of the ∊3 alleles was significantly higher in the control subjects compared to patients (P = 0.002). On the other hand, the frequency of the ∊4 allele was significantly higher in the SLE patients compared to the controls (P = 0.0001). The frequency of allele ∊2 was 1.27% in patients while completely absent in control groups (P = 0.05). In both SLE patients and controls, the genotype distributions were in Hardy–Weinberg equilibrium. Our results for various genotypes of APOE also revealed variations in patient and control groups (Table 2, Fig. 1). The frequencies of genotype ∊3/∊4 and ∊3/∊2 were significantly higher in patients compared to the respective frequencies in controls (Ps ≤ 0.05). Although the frequency of ∊3/∊3 genotype was high in both the test and control, analysis of Saudi population data indicted significant difference in ∊3/∊3 genotype frequencies between patients and controls. Significantly lower frequency of ∊3/∊3 genotype was found in SLE patients compared with that in control (P = 0.001). Other genotypes ∊2/∊4, ∊2/∊2, and ∊4/∊4 were absent in both the SLE patients and control subjects. Upon stratification of SLE patients with or without neuropsychiatric symptoms and renal involvement, significant differences were noted in the alleles and genotype distributions (Tables 3 and 4, Figs. 2 and 3). The frequencies of ∊4 allele and ∊3/∊4 genotype were significantly higher in SLE patients with renal involvement than those without renal complications (Ps = 0.02). On the other hand, significantly higher frequencies of ∊2, ∊4 alleles and ∊2/∊3, ∊3/∊4 genotypes were found in SLE patients with neuropsychiatric symptoms as compared to the patients without any neuropsychiatric involvement (Ps ≤ 0.05). In contrast, the SLE patients without renal or neuropsychiatric involvement had significantly higher frequencies of ∊3 allele and ∊3/∊3 genotype as compared to patients showing these clinical complications (Ps ≤ 0.01). The number of male patients was very small as compared to females; hence, the comparison between the frequencies of APOE genotypes and alleles in male and female SLE patients could not yield significant result.

Alleles and genotypes of APOE polymorphism in SLE patients and controls.

Alleles and genotypes of APOE polymorphism in SLE patients with and without renal involvement (RI).

Alleles and genotypes of APOE polymorphism in SLE patients with and without neuropsychiatric symptoms (NP).
Apolipoprotein E alleles and genotypes in SLE patients and matched controls.

statistically significant
95% confidence interval = 2.54 (1.28–5.02)
95% confidence interval = 2.34 (1.22–4.52)
data for EF.
Apolipoprotein E genotypes and alleles in SLE patients with and without renal involvement.
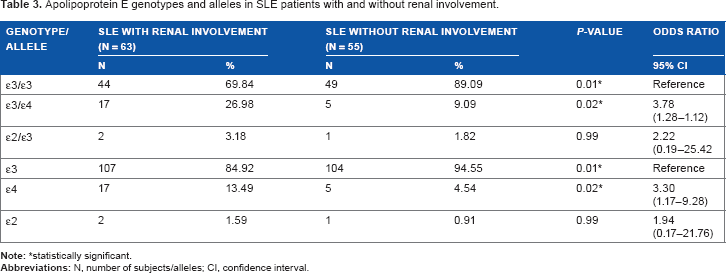
statistically significant.
Apolipoprotein E genotypes and alleles in SLE patients with and without neuropsychiatric symptoms.

statistically significant.
Discussion
In the present study, the significantly higher frequency of allele ∊4 in SLE patients as compared to controls indicated that allele ∊4 is associated with SLE and can be a risk factor (RR = 2.316, EF = 0.429). Our results also showed that allele ∊4 and genotype ∊3/∊4 were significantly associated with both renal involvement and neuropsychiatric symptoms in SLE patients. Similarly, Pullman et al. 30 reported that APOE ∊4 allele and genotype distribution were significantly higher in neuropsychiatric patients with SLE as compared to controls and suggested that allele ∊4 is associated with early onset of the disease. In contrast, Orlacchio et al. 36 reported that the distribution of APOE genotypes in SLE was not significantly different from that of the healthy controls and APOE genotypes are not associated with SLE but suggested that APOE ∊2 might interact with other disease-related factors to accelerate the onset of CAD in patients with SLE and might be an additional marker of risk in North American population. In our study, three SLE patients had APOE ∊2, whereas it is completely absent in controls. Absence of APOE ∊2 has been reported in some populations including Saudis. 33 It is possible that one of the parents in the ancestral pedigree of these three patients might be of some foreign origin.
APOE ∊4 allele is a robust and independent risk factor for a number of diseases, and ∊4 allele has been involved in the pathogenesis of various autoimmune/neuropsychiatric diseases.27,29,37,38 APOE ∊4 allele is believed to be strongly linked to inflammation and lipid levels in related rheumatic disorder, rheumatoid arthritis, 27 and our results are in agreement with it. APOE ∊4 allele has also been associated with the severity in autoimmune diseases such as multiple sclerosis, 39 psoriasis,29,40 and with early onset of Sjogren's syndrome. 41 Recently, Suhail et al. 42 suggested that APOE ∊4 allele may be susceptibility risk factor for myasthenia gravis. APOE ∊4 allele has also been associated with the positive symptoms in schizophrenia. 24 APOE ∊4 allele and genotypes containing ∊4 are also potent risk factors for coronary artery disease43,44 and type 2 diabetes mellitus.45,46
The presence of APOE ∊4 allele is responsible for increased prevalence of cardiovascular diseases in SLE patients by interfering with the lipid metabolism.47,48 The implication of APOE ∊4 in lipid metabolism and developing immunologic responses to lipid antigens may contribute to the development of SLE as reported earlier for psoriasis, another autoimmune disorder. 29 Dyslipidemia is a significant comorbidity of SLE patients with multiple negative effects in the long term. It is believed to decisively affect the long-term prognosis of lupus patients, not only with regard to cardiovascular events but also by influencing other manifestations, such as lupus nephritis. 49 Dyslipidemia in SLE may be driven by inflammation and APOE genetic factors. 27 APOE polymorphisms are major determinants of serum lipid levels in the general population, and APOE genotypes are strongly linked to inflammation and lipid levels. 27 Patients with SLE have an altered lipid profile characterized by increased triglycerides and decreased HDL cholesterol concentrations. Deleterious changes in lipid profile are independently associated with higher concentrations of markers and mediators of inflammation and disease activity and damage in patients with SLE. 50
Our results also indicated that APOE ∊4 is significantly associated with SLE with renal complications. Genetic variation in the APOE gene has also been reported to affect the degree of inflammation by influencing serum CRP levels in dyslipidaemic middle-aged individuals. 51 APOE also exerts a major role in the pathogenesis and the progression of a variety of renal diseases, as well as in the atherosclerotic complications associated with them. Some studies have provided evidence that the APOE ∊4 allele is a risk factor for the progression of renal failure, 52 whereas others have shown that APOE ∊2 allele carries a genetic risk for renal complications.53,54 Liberopoulos et al. 55 reported that the APOE ∊2 allele is independently associated with increased serum uric acid levels in healthy individuals. Recently, APOE ∊2 allele is associated with increased serum creatinine levels and decreased glomerular filtration rate in healthy subjects. 56 However, the reason for this inconsistency in results is not clear.
APOE may have an impact on various steps of multiple pathogenic processes causing variations in SLE features as it functions in both physiological and pathophysiological conditions at multiple levels. 57 For the immunoregulation, APOE affects on the immune system by suppressing T cell proliferation and neutrophil activation, and regulating macrophage function as well as modulating inflammations and oxidation. 58 APOE-suppressing proinflammatory signaling, and vice versa, indicates an intricate APOE-mediated feedback regulation of inflammatory and immune responses. 20 APOE ∊4 allele has been associated with inflammation, which is characteristic feature of SLE.58–60 Significantly higher levels of the proinflammatory cytokines tumor necrosis factor-α and interleukin-6 have been reported in animals expressing the ∊4 allele compared to those with the ∊3 allele. 59 Increased oxidative stress in the APOE ∊4 cells has been suggested to contribute to higher cytokine production by enhancing the activation of nuclear factor-KB. 61 Moreover, increased expression of interleukin-1β, macrophage inflammatory protein-1α, and tumor necrosis factor-α, as well as the transactivation of nuclear factor-κB, have been observed in APOE ∊4 macrophages. 60 APOE allele ∊4 carriers may have increased pro-inflammatory cytokines and high-oxidative stress that may contribute to the SLE. The high frequency of SLE in females of child bearing age in present study and elsewhere 62 may be attributed to the APOE ∊4 boosting effect of estradiol, 63 indicating the functional significance of APOE ∊4 in the etiology of SLE.
Absence of ∊2 allele in Saudi control subjects and its presence in SLE patients with maximum clinical manifestations and systems involvement indicated that ∊2 allele might be associated with severity of SLE in Saudis. In our study, ∊2 carriers were found to have highest CRP levels and had renal and neuropsychiatric involvement. The highest mean CRP levels have also been reported among APOE ∊2 carriers in Japanese Americans and Filipino SLE patients.64,65
Furthermore, our results indicated that allele ∊3 is protective for SLE in Saudis (RR = 0.375, PF = 0.267). Allele ∊3-containing homozygous genotype (∊3/∊3) is also protective for SLE (RR = 0.345, PF = 0.231), whereas heterozygous genotypes ∊2/∊3 and ∊4/∊3 may increase susceptibility. APOE allele ∊3 has also been reported to be protective for other autoimmune/neuropsychiatric disorders.29,66
Conclusion
The ∊4 allele of APOE gene polymorphisms is associated with susceptibility/clinical manifestations of SLE and can be a risk factor, whereas ∊2 allele may increase the severity of the disease. Allele ∊3 is protective for SLE in Saudis. This is the first report indicating the association of APOE polymorphism in the etiology of SLE in Saudi patients. However, further studies with larger population are needed to confirm this association.
Author Contributions
Performed clinical examinations, collected demographic data, and read the manuscript: HA, GH, SJ. Performed genotyping, analyzed data, and drafted the manuscript: MA. Edited and revised the manuscript critically: MT. Designed the study, supervised, and gave final approval for the manuscript to be published: AA. All authors read and approved the final manuscript.
Footnotes
Acknowledgment
The authors thank S. Sadaf Rizvi and Mohammad Al-Asmari for their help with laboratory work.