Abstract
Lung cancer is the second most commonly occurring non-cutaneous cancer in the United States with the highest mortality rate among both men and women. In this study, we utilized three lung cancer microarray datasets generated by previous researchers to identify differentially expressed genes, altered signaling pathways, and assess the involvement of Hedgehog (Hh) pathway. The three datasets contain the expression levels of tens of thousands genes in normal lung tissues and squamous cell lung carcinoma. The datasets were combined and analyzed. The dysregulated genes and altered signaling pathways were identified using statistical methods. We then performed Fisher's exact test on the significance of the association of Hh pathway downstream genes and squamous cell lung carcinoma.
395 genes were found commonly differentially expressed in squamous cell lung carcinoma. The genes encoding fibrous structural protein keratins and cell cycle dependent genes encoding cyclin-dependent kinases were significantly up-regulated while the ones encoding LIM domains were down. Over 100 signaling pathways were implicated in squamous cell lung carcinoma, including cell cycle regulation pathway, p53 tumor-suppressor pathway, IL-8 signaling, Wnt-β-catenin pathway, mTOR signaling and EGF signaling. In addition, 37 out of 223 downstream molecules of Hh pathway were altered. The
Numerous genes were altered and multiple pathways were dysfunctional in squamous cell lung carcinoma. Many of the altered genes have been implicated in different types of carcinoma while some are organ-specific. Hh signaling is implicated in squamous cell lung cancer, opening the door for exploring new cancer therapeutic treatment using GLI antagonist GANT 61.
Keywords
Introduction
Cancer is the third leading cause of death worldwide, and lung cancer is the second most commonly occurring non-cutaneous cancer with the highest mortality rate among all cancers. There are 1.38 million new cases of lung cancer every year, along with 1.23 million deaths due to lung cancer. 1 Based on the histological types of the cells, lung cancer is categorized into two major types, small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC). SCLC occurs almost exclusively in heavy smokers, and makes up about 14% of all lung cancers. NSCLC is a general term for several different types of lung cancer, including squamous cell carcinoma, in which the malignant cells appear larger in size in comparison to SCLC. Collectively, NSCLC subtypes contribute to about 86% of all lung cancers. 2 The average survival time for patients with SCLC is about 10 months. The survival time for patients with NSCLC varies depending on the stage of the disease. Patients with an advanced stage of NSCLC also have an average survival time of 10 months. 3 The 5-year survival rate for SCLC is only 6%, and for NSCLC, the rate is merely 16%, making the average rate 14%, one of the lowest rates of all cancers.3,4
Similar to that of many other cancers, the molecular basis of lung cancer is genetic mutation. When mutation occurs, the expression levels of related genes and their products are altered. Genes in a cell interact with each other in pathways that regulate cell growth, division, DNA repair, and apoptosis.5,6 Unduly expressed or under expressed genes dysregulate biochemical pathways including metabolic pathways and signaling pathways. Lung cancer is a very complex and heterogeneous disease. To develop better cancer treatments, one needs to understand the molecular mechanisms of carcinogenesis, cancer progression and metastasis and to understand how the cancer related genes interact in networks and pathways. The development of microarray technology has made it possible to describe the cellular and molecular states of cells quantitatively and to study the expression levels of many genes simultaneously.7–10 Microarray gene expression profiling of cancer tissues and other disease tissues has been performed by many researchers.8–16 Several groups of researchers conducted gene expression study exploring mRNA gene expression differences in normal cells and different types of lung cancer cells including several non-small cell lung cancer cells.14–16
In this study, we utilized three squamous cell lung cancer microarray datasets14–16 generated by previous researchers to identify dysregulated genes and their associated signaling pathways and to assess the involvement of hedgehog pathway in squamous cell lung cancer.
Materials and Methods
This research utilizes three microarray datasets generated by previous researches for NSCLC. The first dataset was generated by a group of researchers to identify a list of genes that can be used to classify different types of lung cancer. 14 The dataset includes the microarray gene expression profiles of 127 adenocarcinoma specimens, 21 squamous cell lung carcinomas, and 20 pulmonary cardinoids, as well as 17 normal cell samples. Affymetrix HG-U95A oligonucleotide probe microarray chips were used in the experiment. Each chip represents about 12,625 human gene probes.
The second dataset was created by another group of researchers who worked more specifically with squamous cell lung carcinomas. Their work paired normal lung tissue and squamous cell lung cancer tissue from 5 patients, and analyzed their results to find which genes were expressed differently in squamous cell cancer compared to normal cells. 15 The relations and interactions between the altered genes were also investigated. Affymetrix HG-U133A chips were used. Each chip represents 14,500 well-characterized probes.
The third dataset was produced by a group aimed at identifying highly dysregulated genes with biological significance in tumor progression. 16 The dataset has 46 tumor samples, including 13 adenocarcinomas and 32 squamous cell lung carcinomas, and 45 paired nontumor control samples. In the study, gene expression levels were investigated as a function of tumor type, stage and differentiation grade. Affymetrix HG-U133A chips were used. Each chip represents 54,675 mRNA probes.
The focus of this study was on squamous cell lung cancer. We used only the relevant subsets from the three datasets. The raw data (cel files) were provided for the first dataset, and Affymetrix's MicroArray Suite 5 (MAS5) implemented in GeneSpring software version 11.0 (Silicon Genetics, Redwood City, CA) was used to normalize the expression levels of genes for the dataset. The 75 percentile shift was conducted for the second dataset. The median of each sample was transformed to the median of all samples for both datasets. We then carried out statistical analysis to identify the differentially expressed genes between squamous cell lung cancer cells and normal cells. The gene expression levels were log-transformed. For the first dataset, two sample Welch
The set of commonly altered genes was analyzed using the software IPA (Ingenuity® Systems, http://www.ingenuity.com). Related molecular functions and canonical pathways were identified. Top functional groups were identified using Fisher's exact test
20
with
In addition, we examined the expression levels of genes that are associated with hedgehog (Hh) signaling pathway. Hh signaling plays important roles in a variety of normal cellular processes including embryogenesis, patterning of organ structures, tissue repair, cellular proliferation and survival.21,22 The analysis of the connection between the altered genes with Hh pathway was performed using IPA and Fisher's exact test.
Results and Discussion
Differentially expressed genes and their biological functions—
Sorting all the
Top up-regulated genes (common to all datasets).

Top down-regulated genes (common to all datasets).


Heatmap showing fold change patterns of most altered genes.
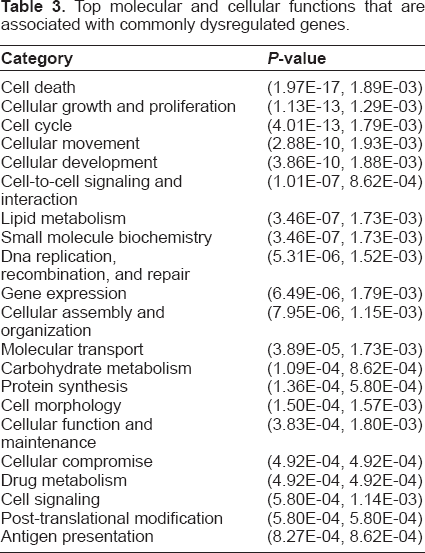
The biological functions associated with the 395 commonly dysregulated genes were obtained using IPA, as they are most likely to be connected to the mechanism of cancer growth. The results show that the genes are associated with many molecular and cellular functions as well as different types of diseases including cancer and genetic disorder. The top molecular and cellular functions are presented in Table 3.
Top molecular and cellular functions that are associated with commonly dysregulated genes.
Canonical signaling pathways associated with differentially expressed genes—
To examine dysfunctional signaling pathways, we utilized all altered genes (without FDR correction) that are common to all three datasets. FDR correction was not conducted for identifying genes since the focus was on individual pathways.
There are 1122 differentially expressed genes with 506 commonly up-regulated and 616 down-regulated. The core analysis using IPA suggests that over 100 canonical signaling pathways are implicated in squamous cell lung cancer (Supplemental Table 3). The top altered pathways include cell cycle: G2/M DNA damage checkpoint regulation, RAR activation, Wnt/β-catenin signaling, mTOR signaling, HIF1α signaling, as well as EGF signaling. The top 40 signaling pathways implicated in squamous cell lung carcinoma are shown in Figure 2.
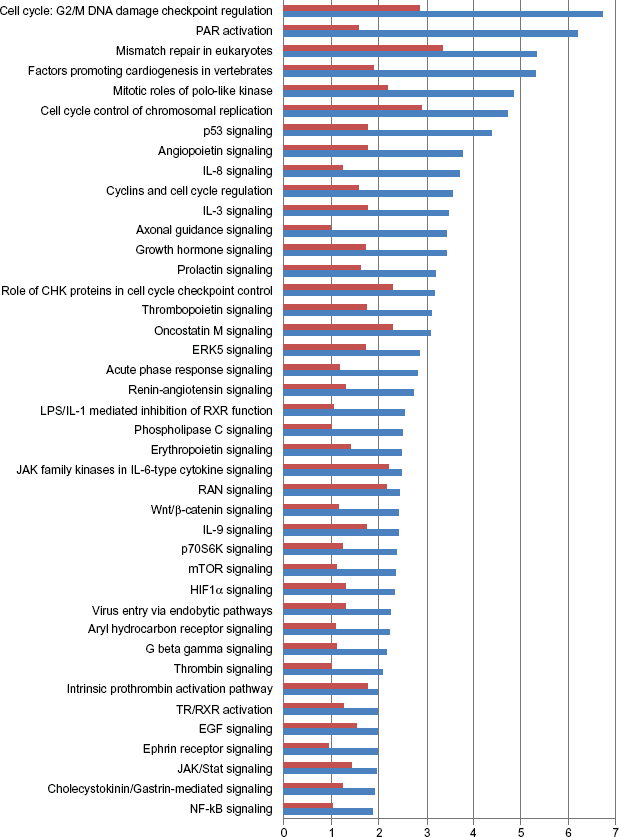
Altered signaling pathways linked with squamous cell lung carcinoma. The blue bars stand for –log(
Downstream molecules of hedgehog signaling pathway—
The hedgehog signaling pathway has been implicated in the development of different human cancers including prostate cancer, colon cancer and basal-cell carcinoma. GLI proteins are the effectors of hedgehog signaling. GLI1 and GLI2 are the two key members of GLI family and together they have 223 downstream target molecules collected in the IPA knowledge database. We analyzed the GLI target molecules for their connection with squamous cell lung cancer. Interestingly, 37 out of 223 are significantly altered. The 37 associated genes are presented in Table 4. 21 of the 37 were up-regulated. 15 of them were considerably differentially expressed in squamous cell lung cancer cells with FDR corrected
GLI1/GLI2 downstream genes commonly differentially expressed in all three datasets.

Up-regulated genes.
The Fisher's exact test on the 37 genes yields a
Contingency table for GLI1/GLI2 downstream molecules.
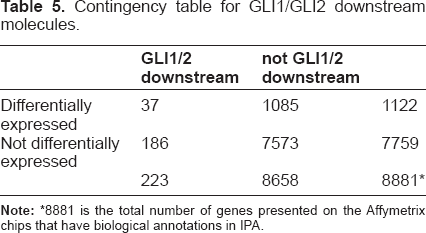
8881 is the total number of genes presented on the Affymetrix chips that have biological annotations in IPA.
Genes in a cell interact with each other through a sequence of orchestrated events in pathways. Aberrant cell signaling along the pathways breaks the organized events, promotes proliferation, inhibits apoptosis, induces angiogenesis, and facilitates the cancer to spread. Our study combined three microarray datasets to explore more stable biomarkers or drug targets for squamous cell lung cancer and the pathways implicated. The results indicate that the expression levels of many genes were altered and multiple pathways were dysfunctional in squamous cell lung carcinoma (Supplemental Tables 1 and 2, Fig. 1).
The altered genes encode cytokines, enzymes, growth factors, kinases, transporters transmembrane/nuclear receptors and transcription regulators. They participate in variety of molecular activities in different biological processes. The molecules were located in multiple cellular locations in extracellular space, cytoplasm, nucleus, and plasma membrane.
As we can see from Table 1, the top up-regulated genes include members of the keratin family, keratin 6A, keratin 15, keratin 16, and keratin 17. Their expression levels were significantly elevated. Keratins are intermediate filament–forming proteins that provide mechanical support and perform a diversity of other functions in epithelial cells.23–25 All four of them are epithelial keratin genes. They are associated with molecular functions such as structural constituent of cytoskeleton and protein binding and participate in cytoskeleton organization, cell differentiation, regulation of cell proliferation, epidermis development, intermediate filament cytoskeleton organization and ectoderm development.
The proteins encoded by keratin 6A are type II cytokeratins (neutral-basic). These cytokeratins organize in pairs of heterotypic keratin chains, which are co-expressed during differentiation of simple and stratified epithelial tissues. Keratin 15, 16 and 17 are acidic. Keratin 16 and 17 are paired with keratin 6 while keratin 15 is paired with keratin 5 in squamous epithelial cells. It has been reported that keratins can be used as diagnostic markers in human carcinomas. 25 A recent review on keratin summarized the evidence for active keratin involvement in cancer cell invasion and metastasis, as well as in treatment responsiveness. 25 However, their involvement in squamous cell lung carcinoma requires further investigation.
Also significantly over-expressed was cyclin-dependent kinase 1 (CDK1). The protein encoded by this CDK1 is a member of the Ser/Thr protein kinase family. It is a catalytic subunit of the highly conserved protein kinase complex known as M-phase promoting factor, which is essential for G1/S and G2/M phase transitions of eukaryotic cell cycle. The kinase activity of this protein is controlled by cyclin accumulation and destruction through the cell cycle. The phosphorylation of this protein also plays important regulatory roles in cell cycle control.26–28 CDK1 is involved in many signaling pathways including apoptosis signaling; ATM signaling; cell cycle: G2/M DNA damage checkpoint regulation; cyclins and cell cycle regulation; role of CHK proteins in cell cycle checkpoint control; and sonic hedgehog signaling. In addition, several other cell-division cycle proteins including CDC20 and CDC45 were up-regulated.
The top down regulated genes are presented in Table 2. Several LIM domain proteins, FHL1, LIMCH, and LMO3, are in this group. Also included is LDB2 (Supplemental Table 2). The LIM domain is a double-zinc finger motif found in a diversity of proteins. The zinc fingers have highly conserved cysteines and joined by a 2-amino acid spacer. LIM proteins include transcription factors and cytoskeletal proteins. It is reported that the primary role of LIM domains is in protein-protein interaction, through the formation of dimers with identical or different LIM domains or by binding distinct proteins. LIM-only proteins are implicated in the control of cell proliferation.29,30
FHL1 that encodes a member of the four-and-a-half-LIM-only protein family was significantly down regulated. Expression of these family members occurs in a cell and tissue specific mode and the proteins are involved in many cellular processes. The reduced expression of FHL1 in lung cancer has been recently reported. It was suggested that the inhibitory effects of FHL1 on lung cancer cell growth were associated with both the G1 and the G2/M cell cycle arrest associated with a marked inhibition of cyclin A, cyclin B1 and cyclin D as well as the induction of the cyclin dependent kinase inhibitors p21 and p27. 30 Conversely, several members of cyclin family (cyclin A2, cyclin B1, cyclin B2 and cyclin dependent kinase I) were found up-regulated in lung cancer in this study (Supplemental Table 1) while cyclin dependent kinase inhibitors CDKN1A and CDKN1B were consistently down-regulated in all three datasets (although the down-regulation levels were not above the statistical significant threshold).
Down-regulated RHOB is a member of the RHO GTP-binding protein family. The RHO family plays a crucial role in the dynamic regulation of the actin cytoskeleton. Studies suggested that the proteins of the family also have a diversity of functions in membrane pathways. The RHOB was shown to localize to vesicles of the endocytic compartment, suggesting a potential function in regulation of endocytic traffic. The expression of active RHOB causes a delay in the intracellular trafficking of the epidermal growth factor (EGF) receptor through regulating PRK1.31–33
The signaling pathway investigation in this study reveals that many of the cancer related pathways reported in the literatures are also responsible for squamous cell lung carcinoma. They include cell cycle regulation pathway (
Cell cycle: G2/M DNA damage checkpoint regulation pathway is on the top of the altered pathways. G2/M checkpoint is the second checkpoint within the cell cycle. It prevents cells from entering mitosis when DNA is damaged, providing an opportunity for repair and stopping the proliferation of damaged cells and helping to maintain genomic stability. 26 14 out of 49 molecules on the pathway were significantly altered in lung cancer including the cyclin-dependent kinase 1 (CDK1) which is essential to drive cells into mitosis (Supplemental Tables 1 and 2).
p53 activation is induced by a number of stress signals, including DNA damage, oxidative stress and activated oncogenes. The p53 protein is employed as a transcriptional activator of p53-regulated genes. This results in cell cycle arrest, cellular senescence or apoptosis. 17 out of 49 molecules on the pathway were significantly altered, including up-regulated SERPINB5 and TP63. Serpin peptidase inhibitor, clade B (ovalbumin), member 5 (SERPINB5), also known as maspin, is a tumor suppressor gene. It was reported to block the growth, invasion, and metastatic properties of mammary tumors. Its expression was down-regulated in breast tumors, prostate cancer and uterine cervix cancer.34–36 It was reported that SER-PINB5 over-expression increased the rate of apoptosis of both preneoplastic and carcinomatous mammary epithelial cells and reduced tumor growth through a combination of reduced angiogenesis and increased apoptosis. 32 However, the role of SERPINB5 in the inhibition of tumor growth is rather complex. In contrast to its reduced expression levels in breast cancer and other types of cancer, SERPINB5's over-expression in pancreatic carcinoma was also reported. 37 A recent study shows how it works in regulating cell growth when tightly coupled with the thrombin receptor protease activated receptor-1 (PAR-1).38 Our study indicates that SERPINB5 is significantly over-expressed in squamous cell lung carcinoma in all three datasets (Table 1). The tumor protein TP63 is another up-regulated gene (Table 1) that encodes a member of the p53 family of transcription factors. It acts as a sequence specific DNA binding transcriptional activator or repressor. Its over-expression has already been documented in lung carcinoma. 39
EGFR encodes a receptor for members of the epidermal growth factor family. Binding of the protein to a ligand induces receptor dimerization and tyrosine autophosphorylation and leads to inappropriate activation of the anti-apoptotic Ras signaling cascade, eventually leading to uncontrolled cell proliferation. Both EGFR and the signal transducer and activator STAT1 were up-regulated in the three datasets (based on the
Another important cancer related pathway is Wnt/β-catenin signaling pathway. Wnt signaling pathway is a network of proteins best known for their roles in embryogenesis and cancer, but also involved in normal physiological processes in adult animals.46,47 Wnt proteins are secreted morphogens that are required for basic developmental processes, such as cell-fate specification, progenitor-cell proliferation and the control of asymmetric cell division, in many different species and organs.49,50 Wnt ligand binds to its receptor and stabilizes cytoplasmic β-catenin through inhibition of the β-catenin degradation complex. β-catenin is then free to enter the nucleus and activate Wnt-regulated genes through its interaction with TCF family transcription factors and associated recruitment of coactivators. 20 out of 174 Wnt signaling related genes are altered in the datasets. Although conflicting roles of Wnt-5 A in cancer have been reported,
51
our data shows it is significantly up-regulated and the Wnt inhibitor WIF1 is down-regulated. The
mTOR pathway has been extensively explored in some other cancers, including renal cell carcinoma, and large clinical trials have reported promising progression-free survival advantage with mTOR inhibitor drugs in this disease. Our report suggests that mTOR signaling may be involved in squamous cell lung carcinoma as well, and exploring this pathway in more depth in squamous cell carcinoma of the lung may be needed.52,53
Many hurdles remain in the race to find a better treatment to cancer. Recently a cellular screen for antagonists of GLI-mediated transcription revealed that a small molecule GANT61 was able to selectively inhibit the GLI-mediated gene transactivation, which is the last step of the Hh pathway.
54
Hh signaling is tightly coupled with Wnt signaling.
47
It plays important roles in a variety of normal cellular processes including embryogenesis, patterning of organ structures, tissue repair, cellular proliferation and survival.21,22 The signaling pathway is activated when hedgehog signaling ligands, such as sonic hedgehog, bind to the membrane receptor patched (PTCH), and thus stop the suppression of the signaling molecule smoothened (SMO) caused by PTCH. SMO then activates the cytoplasmic protein SUFU, which releases the GLI family of transcription factors, GLI1 and GLI2. Once the GLI molecules are released, they translocate to the nucleus, where they bind to target genes to start transcription.21,22 The signaling process is shown in Figure 3. Hh signaling pathway is an important regulator of the embryonic development and becomes reactivated in cancer. It has been activated in several human cancers including prostate cancer, colon, and small cell lung cancer although the pathway seems to be inactive in most normal adult tissues.55–58 The inhibition of this pathway could offer a selective way of controlling just the tumor cells. In our data, GLI2 was up-regulated although the significantly altered expression level of GLI1 was not observed. On the other hand, our investigation into the group of GLI1 and GLI2 downstream molecules using IPA revealed that significant number of genes was altered in the group (Table 5). The

Mechanism of hedgehog signaling.
Conclusion
By combining three microarray datasets, we created a more comprehensive and complete gene expression analysis of squamous cell lung cancer. The list of altered genes is more reliable than the ones obtained from a single setting of experiments, therefore providing a more dependable set of lung cancer biomarkers/drug targets. Equally the group of implicated signaling pathways is more trustworthy. The results of the analysis indicate numerous genes were altered and multiple pathways were dysfunctional in squamous cell lung carcinoma. As discussed above, many of the altered genes have been implicated in different types of carcinoma although some of the altered genes are organ-specific. Hh signaling is implicated in squamous cell lung cancer, opening the door for exploring new cancer therapeutic treatment using GLI antagonist GANT 61. While active in many different types of cancer, Hh pathway is believed to be mostly inactive in normal adult tissues, making it a good target for cancer treatments. Consequently, the treatment based on GANT 61 is likely to cause few side effects.
Disclosures
Author(s) have provided signed confirmations to the publisher of their compliance with all applicable legal and ethical obligations in respect to declaration of conflicts of interest, funding, authorship and contributorship, and compliance with ethical requirements in respect to treatment of human and animal test subjects. If this article contains identifiable human subject(s) author(s) were required to supply signed patient consent prior to publication. Author(s) have confirmed that the published article is unique and not under consideration nor published by any other publication and that they have consent to reproduce any copyrighted material. The peer reviewers declared no conflicts of interest.
Supplementary Materials
Supplementary tables 1, 2 and 3 can be found in: 8283 files.zip
Footnotes
Acknowledgment
Authors thank Christina Wooley for helpful discussions. This work was supported in part by the Choose Ohio First in Bioinformatics Scholarship grant.
