Abstract
Two problems now threaten the future of anticancer drug development: (i) the information explosion has made research into new target-specific drugs more duplication-prone, and hence less cost-efficient; and (ii) high-throughput genomic technologies have failed to deliver the anticipated early windfall of novel first-in-class drugs. Here it is argued that the resulting crisis of blockbuster drug development may be remedied in part by innovative exploitation of informatic power. Using scenarios relating to oncology, it is shown that rapid data-mining of the scientific literature can refine therapeutic hypotheses and thus reduce empirical reliance on preclinical model development and early-phase clinical trials. Moreover, as personalised medicine evolves, this approach may inform biomarker-guided phase III trial strategies for noncytotoxic (antimetastatic) drugs that prolong patient survival without necessarily inducing tumor shrinkage. Though not replacing conventional gold standards, these findings suggest that this computational research approach could reduce costly ‘blue skies’ R&D investment and time to market for new biological drugs, thereby helping to reverse unsustainable drug price inflation.
Introduction
The development of massive online text archives and biomedical data warehouses over the last decade has created new opportunities to mine useful information from fuzzy datasets comprising both statistical and semantic information.1–3 Association rules have been developed to extend fuzzy set theory to text-mining; at the simplest level, identification of a given set (e.g. both A and B not C) at higher-than-expected frequency provides support for the probability of a causal link between A and B in the absence of C, albeit with varying confidence levels quantifiable as membership functions or degrees of truth. The non-digital algorithms so derived may generate value from masses of noisy data, in effect representing Bayesian interpretations of semantic information. 4 Examples include tracking of influenza epidemics from search engine patterns, 5 the epidemiologic surveillance of AIDS, 6 monitoring of the blogosphere, 7 academic publication analysis, 8 and assessment of scientific merit based on impact factors. 9
The causality of such associations gleaned from text-mining of the oncology literature is readily illustrated
New anticancer drug development, by virtue of its target-specific nature, should readily lend itself to such semi-digital analysis. 10 Returns on investment in applied cancer research have been declining in recent years due to a decline of blockbuster drug frequency. 11 This has led in turn to progressive escalation of the cost for bringing a therapeutic to market, which now approaches US$1bn per FDA-licensed product; 12 such expense is in turn handed on to the health-care consumer. 13 Part of this cost relates to the inefficiency of the modern clinical trial system, for which problem many investigators are seeking fresh approaches such as multi-arm designs, 14 personalized medicine or pharmacodiagnostics. 15 A related problem specific to the cancer field is that selection of drugs for costly phase III trials remains based exclusively on demonstration of drug ‘activity’ quantified as tumor shrinkage, or response. 16 This endpoint, although convincingly associated with short-term therapeutic benefit, may lack sensitivity for detecting the metastasis-inhibiting activity of non-tumorilytic drugs—which could well correlate strongly with survival benefit, and be better predicted by biomarker expression. 17 Hence, the present study seeks to test how low-cost text-mining may improve applied cancer research feasibility while reducing investment risk.
Methods
Searches were undertaken using the latest text-based search-and-retrieval version of PubMed—a service of the National Library of Medicine developed by the National Center for Biotechnology Information used to integrate the major databases (including PubMed Central, Journals, Books, OMIM). The Entrez cross database search page was used to access the Entrez Global Query database search engine. A search across the Entrez database was performed by entering one or more search term(s) or phrase(s) to execute the search.
Using this approach, the PubMed database was serially interrogated using the terms representing both the primary set of interest (P) and the secondary sets of interest (S1, S2, etc.), resulting in identification of the common set of interest (C1, C2, etc.). Arithmetical correction was made for different frequencies of S1, S2, etc., permitting calculation of the expected value of C for a given S intersection given knowledge of previous values of C and assuming the null hypothesis. The ratio of C2 to C1 was then considered the multiplier (M2) by which the null hypothesis for S2 (compared to S1) was tested. Non-parametric statistical analysis was performed by chi-square calculation.
Results
An initial example of how disease biology or phenotype can be predicted by text-mining is presented in Figure 1 and Supplement S2, which illustrate that the clinical complication of finger clubbing is more often associated with lung tumors of either squamous cell carcinoma or adenocarcinoma histology than with small-cell tumors (χ
2
= 37.96, df = 2,
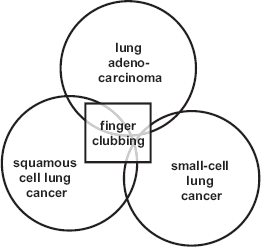
schematic illustration of a text-mined association. The extent to which a phenotype such as finger clubbing is associated with a specific lung cancer histology may be estimated by association (see supplement S2), as visualized here.
Molecular biological issues may also be addressed by text-mining which shows, for example, that the p53 signalling (apoptotic) pathway is 70% more strongly associated with the

Phenotypic distinction of ‘adenocarcinoma’ vs. ‘squamous carcinoma’ using text-mining; for each phenotype shown, the PubMed database was exclusively interrogated (e.g. adenocarcinoma NOT squamous cell carcinoma, and vice versa). The figures on the horizontal axis represent the multiplier (M) by which each phenotype is preferentially associated with the histology labelled on the adjacent vertical axis.
This strategy can be further applied to questions relating to clinical trial design. Imagine, for example, that there are two novel candidate anticancer therapeutics: one is an antagonist to insulin-like growth factor-1 (IGF1), and the other an inhibitor of signalling via the hepatocyte growth factor (HGF, scatter factor, c-met ligand) pathway. A simple text-mining search shows that HGF is eight-fold more strongly associated with tumor metastasis, whereas IGF1 is three times more strongly associated with drug resistance (χ
2
= 564.15, df = 1,
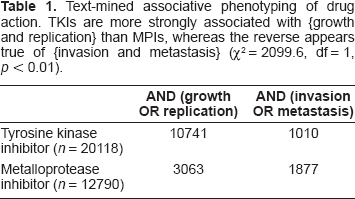
Table 1 provides another illustration of hypothesis generation relevant to clinical trials using similar secondary data sets. In this case, a company has drugs of two classes, either tyrosine kinase inhibitors (TKIs) or metalloprotease inhibitors (MPIs). The question is again raised: is there any difference in the respective likelihoods that one drug class will be more active in causing tumor responses (i.e. antagonizing replication) while the other may be less active in its antimitogenic action but more active in suppressing metastasis? As shown, the text-mined association of MPIs with antimetastatic (vs. antiproliferative) key words is 6.5-fold that of TKIs, whereas the reverse trend is true for antiproliferative terms (χ
2
= 2099.6, df = 1,
Text-mined associative phenotyping of drug action. TKIs are more strongly associated with {growth and replication} than MPIs, whereas the reverse appears true of {invasion and metastasis} (χ
2
= 2099.6, df = 1,
This approach could also be applied to prediction of targeted drug efficacy for different disease subtypes prior to the advent of a validated biomarker. Consider the scenario in which a company has a promising mTOR pathway inhibitor and is wanting to prioritise its phase 2–3 clinical trial strategy. Since the Akt proto-oncogene is an important node in the mTOR signalling network, we may begin by characterising the text-mined associations of different tumor subtypes with Akt; this analysis shows that the strongest association is with prostate cancer (Table 2).
Association of tumor subtypes with a putative surrogate biomarker for the mTOr pathway, Akt, showing that the strongest association is with prostate cancer. As expected, the much stronger correlation of the mTOr inhibitor temsirolimus, which is already licensed for use in renal cell cancer, is with the latter malignancy. P, primary set size; C, intersecting set size.

In contrast, when assessing the associations of the mTOR pathway inhibitor temsirolimus, the strongest association is with its licensed indication in renal cell carcinoma. Although not surprising, this difference raises the commercially relevant hypothesis that a ‘second use’ for mTOR inhibitors could well be confirmed for other malignancies including, though not limited to, prostate cancer.
The associations of different treatment approaches with text-mined costs and benefits may also be compared using this seamless approach. Figure 3 shows the comparative quantitative associations of the secondary variable {overall survival benefit) with the primary variables of either {adjuvant cancer therapy} in column 1, {palliative cancer therapy} in column 2, {maintenance cancer therapy} in column 3, or {preventive cancer therapy} in column 4, and reveals a steady decline in that order. In contrast, this trend is reversed when {cost-effectiveness} is substituted as the secondary variable. This striking difference suggests that third-party payers could scrutinize the value of adjuvant treatment claims more intensively, while subsidizing preventive interventions for healthy individuals with greater enthusiasm. For companies under pressure to prove survival benefits for product licensing, on the other hand, these data suggest that drugs used in palliative or maintenance settings may deliver more easily defended ratios of QALY to unit cost than adjuvant studies (as well as being cheaper to conduct in the first instance). The possibility is also raised that biologically-targeted preventive agents could become the backbone of the blockbuster drug market.

Text-mined comparison of survival vs. cost data for different cancer treatment modes, suggesting that adjuvant therapies are most strongly linked to survival benefit (open columns) but least to cost-effectiveness (solid columns), whereas preventive treatments have the opposite associations. Palliative and maintenance treatments appear intermediate for both secondary endpoints.
Discussion
The central finding of this study is an exciting one for modern researchers: namely, that the sudden availability of huge online text-based databases permits novel, cheap and potentially important research to be conducted by computer-based researchers, thereby expanding the scope and accessibility of research methodology. Moreover, by extending this approach, innovative informatics strategies may be developed and automated over time to derive value from otherwise wasted informational resources. This is true not only for curiosity-driven academic research but, as shown here, also for commercially-driven applied science.
It must be emphasized, however, that numerous pitfalls await the unwary who seek to ascribe causality to text associations without rigorous exclusion of confounders (to appreciate this, one has only to consider the famous ‘beer-and-diapers’ market research anecdote). 19 The challenges include unambiguous identification of genes or other molecules, leading to false-positive data inclusions, and failure of current text-mining systems to achieve 100% recall, leading to false-negative loss of sensitivty. Hence, unless carefully excluded, synonyms and name variations may cause significant problems with the accuracy of text-based data mining.
An additional problem relates to system ‘noise’ which, in any text-mined database, may be expected to increase as the age of the data archive increases. To address the confounding effect of ageing data will involve inevitable compromises between database size and data novelty; on the other hand, more recent data may be subject to dispute or lack of replication, so may be less reliable than more durable (and hence, text-repetitive) data. A further limitation is that text terms may change over time, with molecules or drugs being renamed, or having different names in different languages or countries. Such limitations require prior knowledge on the part of the investigator(s) to enable corrective action in anticipation.
How can a purely text-mining approach distinguish positive and negative associations? A given article examining two variables may not report a simple positive association, but rather the lack of an association or even an inverse or negative association; text-mining will not distinguish these contingencies, however. In general, however, it is reasonable to assume that positive associations will vastly outnumber negative associations when measured by text-mining, if only because negative associations tend to be less interesting to readers (and hence to journals and publishers) who may in general be keener to understand causality than exceptions or confounders.
Notwithstanding these important caveats, it is becoming clear that improved knowledge management is essential to regaining momentum in pharmaceutical R&D, 20 and this unmet need requires fresh approaches. The advantages of the text-mining approach described here are that it is simple, cheap, fast, objective, and powerful. Its disadvantages are at least as plentiful, but perhaps the main ones are (i) that it is easily confounded by overlooking nonrandom associations which may not be obvious to all investigators, and (ii) that careful thought and broad background knowledge are needed to frame a series of questions that can provide a useful association. These limitations are sufficiently serious to restrict the use of such approaches for now to hypothesis generation, but it seems reasonable to expect that methodologic refinement and practical experience will lead to far more sophisticated approaches that will eventually be incorporated into routine research practice. Indeed, it may even become possible to devise programs that scan scientific literature and experimental data in the background, generating associations and hypotheses at different thresholds of certainty and improbability (creativity). In some ways this would be an expected outcome of the information explosion which now produces more data than any individual can hope to absorb and evaluate. 21
The examples given here relate to the field of oncology, but could just as readily be applied to other therapeutic areas such as cardiovascular or neurological disease. In the same way that credit card companies and search engines now scan electronic data for commercially useful patterns, so can scientists now begin looking beyond their own laboratory benches in an effort to exploit the informatic power of modern databases. Although the challenges of text-data specificity and sensitivity are not to be underestimated, we may discover that the fruits of the genomic revolution will be greater than any of us can yet conceive.
Conflict of Interest Statement
There is no conflict of interest.
Footnotes
Acknowledgements
I thank Joyce Mak for statistical analysis, and Professors Karen Lam and Raymond Liang for support.
Supplementary Data
Text comparison of ‘kinase’ and ‘protease’ strings as an association of pro-replicative vs. pro-metastatic terminology (see also Table 1).

| AND (growth OR replication OR mitosis) | AND (migration OR motility OR metastasis) | |
|---|---|---|
| Kinase | 147941 | 23748 |
| Protease | 44028 | 15732 |
Quantification of text-mined molecular phenotype: there is a greater than 16-fold difference in the association of ‘kinase’ with pro-mitogenic vs. pro-metastatic key words when compared with the corresponding associations of ‘protease’ (χ
2
= 4889.64, df = 1,
