Abstract
Genes encoding proteins that contain the universal stress protein (USP) domain are known to provide bacteria, archaea, fungi, protozoa, and plants with the ability to respond to a plethora of environmental stresses. Specifically in plants, drought tolerance is a desirable phenotype. However, limited focused and organized functional genomic datasets exist on drought-responsive plant USP genes to facilitate their characterization. The overall objective of the investigation was to identify diverse plant universal stress proteins and Expressed Sequence Tags (ESTs) responsive to water-deficit stress. We hypothesize that cross-database mining of functional annotations in protein and gene transcript bioinformatics resources would help identify candidate drought-responsive universal stress proteins and transcripts from multiple plant species. Our bioinformatics approach retrieved, mined and integrated comprehensive functional annotation data on 511 protein and 1561 ESTs sequences from 161 viridiplantae taxa. A total of 32 drought-responsive ESTs from 7 plant genera
Keywords
Introduction
Environmental stresses can negatively impact agricultural crop yield and quality.1,2 As an adaptive strategy, plant genomes encode genes that produce proteins that function in stress response and tolerance.3–5 Despite substantial research on response to abiotic and biotic stresses by plants, there are still knowledge gaps regarding the molecular mechanisms that regulate the diverse functions of environmental stress-associated plant genes and proteins. 3 The increasing availability of genomic sequences of members of the viridiplantae (green algae and land plants) in combination with high-throughput bioinformatics tools and databases4,5 provide new opportunities for examining understudied gene families that could be central to stress response in plants.
Genes encoding proteins that contain the conserved 140–160 residues Universal Stress Protein (USP) domain (Pfam Accession: PF00582) are known to provide bacteria, archaea, fungi, protozoa, and plants with the ability to respond to a plethora of environmental stresses.6–9 Nutrient starvation, drought, high salinity, extreme temperatures and exposure to toxic chemicals are examples of conditions that induce expression of genes with the USP domain. Proteins containing domain PF00582 are often collectively referred to as universal stress proteins. In
Kerk et al
13
examined the sequence and structure of 44
Water-limiting condition (drought) is one of the key abiotic stresses that can adversely affect the growth, development and yield of crop and tree plants. 20 Drought induces biochemical and physiological responses in plants 21 including reduced photosynthetic carbon and energy metabolism 22 leading to oxidative stress. High salinity is also accompanied by drought. 20 Furthermore, wood production from forest trees can be hampered by drought.32,33 The ability to respond and tolerate drought stress is a desirable phenotype especially in plants that have to survive in environments with insufficient water. The molecular and cellular mechanisms for response and tolerance have been investigated using a range of powerful high-throughput genomic and proteomic techniques to dissect gene networks response to drought. 22 Examples of drought-responsive USP genes have been reported in cotton 18 and cowpea. 23 The identification of drought responsive USP genes from multiple plants species will present an array of research tools for genetic manipulation of plants for drought tolerance. Therefore, we sought to develop a bioinformatics screening strategy to identify drought-responsive USP genes and transcripts from comprehensive protein and gene transcript databases.
There continues to be an increase in number and diversity of bioinformatics resources storing functional annotation of protein-coding sequences including those containing the USP domain.
24
The Pfam database of protein families represented by alignments and Hidden Markov Models contains at least 550 protein sequences from the viridiplantae (green algae and land plants) annotated to contain at least one USP domain.
25
These sequences have identifiers of the Universal Protein Resource (UniProt), which is the most comprehensive catalog for protein sequence and functional annotation data.
26
The UniProt entries have valued-added cross-references to external databases that provide diverse annotation including structural, gene expression, literature and sequence diversity. In addition, there are specialized plant databases not yet linked to UniProt. For example, the Phytozome resource page (http://www.phytozome.net/Phytozome_resources.php) provides links to resources for general plant genomics; gene expression; gene indices and Expressed Sequence Tags (ESTs);
Among the ESTs and cDNA resources listed in Phytozome, we observed that the TIGR Plant Transcript Assemblies database (Plantta) 27 had a wide collection of 254 plant species (as of July 2007). The ESTs and full-length cDNA are being used for discovery of genes in plant species as well as evidence of gene expression in conditions as well as anatomic parts. The identification of ESTs encoding universal stress proteins could facilitate further studies on selection of markers for comparative mapping, plant breeding and forward genetics.28,29
The Plantta resource contains simple sequence repeats (SSR) or microsatellite annotation for some transcripts. Microsatellites are 1–6 bp tandemly repeated DNA sequences that occupy a significant fraction of the nuclear genome of all eukaryotes. 30 Microsatellites in protein-coding genes can inactivate or activate genes or truncate protein. 31 In plants, microsatellites derived from EST sequences (EST-SSRs) have been proposed to be better candidates for gene tagging and are preferred over genomic-SSR markers for plant improvement programs owing to their higher interspecific transferability rate. 32 Thus, we investigated the presence of SSR on transcript assemblies and singleton sequences in Plantta. Furthermore, since our primary interest was on drought-responsive genes, we sought to identify USP-annotated Plantta ESTs that contain text relevant to drought in their dbEST 33 entries. The keyword search provided an indication of the experimental condition for generating the cDNA libraries. Finally, we determined the overlap of EST dataset containing SSR entries with the EST dataset annotated with drought or water stress.
The bioinformatics strategy described can be adapted for analyzing a set of viridiplantae protein sequences defined by a Pfam protein domain. Furthermore, plant transcripts from other abiotic and biotic stress conditions can be mined and analyzed. In summary, we identified diverse plant universal stress proteins and transcripts responsive to drought including those that contain microsatellite markers that may regulate their function.
Methods
Construction of Dataset of Viridiplantae Universal Stress Proteins
Viridiplantae proteins annotated in Pfam database 25 with Pfam domain PF00582 were downloaded and computationally processed with a suite of UNIX and PERL scripts to retrieve their respective UniProt Identifiers. Subsequently, for non-obsolete or deleted UniProt entries, the protein domain architecture, organism source of sequence, protein sequence length and protein molecular weight were extracted from XML-formatted UniProt entries (UniProt release 2010_10—Oct 5, 2010). These selected annotations are typically available for UniProt entries. Overview of the USP dataset construction is illustrated in Figure 1. Analysis of the protein domain architecture annotation provided a prediction of the number of USP domains as well as additional types of protein domain(s) present.

Flowchart for constructing dataset of viridiplantae universal stress proteins.
Orthologous Viridiplantae Drought-Responsive Genes Encoding Universal Stress Proteins
A UniProt entry for a protein sequence contains value-added cross-references to other databases (http://www.uniprot.org/docs/dbxref). The cross-referenced databases for each viridiplantae USP entry was computationally extracted from the XML formatted files. A non-redundant list of the databases was assembled and used to construct a presence-absence matrix consisting of rows of UniProt protein identifiers and columns of selected databases. A zero (0) was used to encode absence of cross-referencing to a database and one (1) for presence of cross-reference to a database. This matrix was then searched for USP entries with cross-reference to the Gene Expression Atlas (a subset of ArrayExpress) 45 and Ortholog MAtrix Project (OMA) Browser. 34 The matrix was visualized using a Linux version of matrix2png. 35 The Gene Expression Atlas (GXA) stores microarray and other gene expression data and was selected because it had annotation for “Experimental Factors”, which included a subsection on “Environmental Stresses” such as drought. Furthermore, the OMA Browser allows for exploration of orthologous relations between protein sequences for 1000 species (Release of May 2010).
A combination of the data from GXA and OMA allowed us to identify orthologous plant proteins in which a member has been demonstrated to be responsive to drought. Additional homologous sequences for the identified drought up-regulated USPs were retrieved from PLAZA—a resource for plant comparative genomics 36 and their multiple sequence alignment generated using ClustalW2 at http://www.ebi.ac.uk/clustalw/.
Viridiplantae Universal Stress Protein Transcripts Derived from Drought Conditions
The TIGR Plant Transcript Assemblies (Plantta; http://plantta.jcvi.org/) 27 consists of a collection of transcripts (assembled ESTs and singletons) for at least 215 plant species. The content of webpage for each USP transcripts in the Plantta resource was also parsed to identify those with microsatellite (SSR) annotation. We sought to identify universal stress protein ESTs from cDNA library source derived from drought stress. The first step involved retrieving from Plantta, transcripts annotated with the text “universal stress protein”. In the second step, all the ESTs identifiers in dbEST 33 associated with the Plantta transcripts were retrieved and the entries in GenBank downloaded and searched for text “drought”. Another search strategy, the dbEST entries were searched for text “water” and then the retrieved subset searched with text “stress”. The assumption was that the presence of “drought” or combination of “water” and “stress” was indicative of a cDNA library derived from drought stress conditions. This mining of text in the dbEST entries was done to help identify universal stress protein ESTs as research tools for understanding stress response in a large number of plant species of agricultural, economic, ecological or industrial importance but without complete genome sequences.
Results
Construction of Dataset of Viridiplantae Universal Stress Proteins
A total of 511 viridiplantae proteins annotated with universal stress protein domain (PF00582) from 43 unique taxa (NCBI Taxonomy IDs) were downloaded from UniProt on October 24, 2010 (Table 1). The protein count per taxa ranged from 1 to 88. The protein counts for Liliopsida (monocotyledons), dicotyledons, and other viridiplantae including green algae were 235, 203 and 73 respectively. Furthermore, land plants with at least 50 USP records in UniProt from the Pfam dataset were
Dataset of viridiplantae universal stress proteins entries in UniProt.


Distribution of sequence length of 511 viridiplantae universal stress proteins.
A total of 17 Pfam protein domains arranged in 17 architectures were associated with the dataset (Table 2 and Fig. 3). Ten of the 17 protein domains occurred only in one protein, most of which are uncharacterized as with sequences from
Distribution of protein families in viridiplantae universal stress proteins.

Description of protein domains available at http://pfam.sanger.ac.uk/.
Viridiplantae universal stress proteins with tandem USP domains.
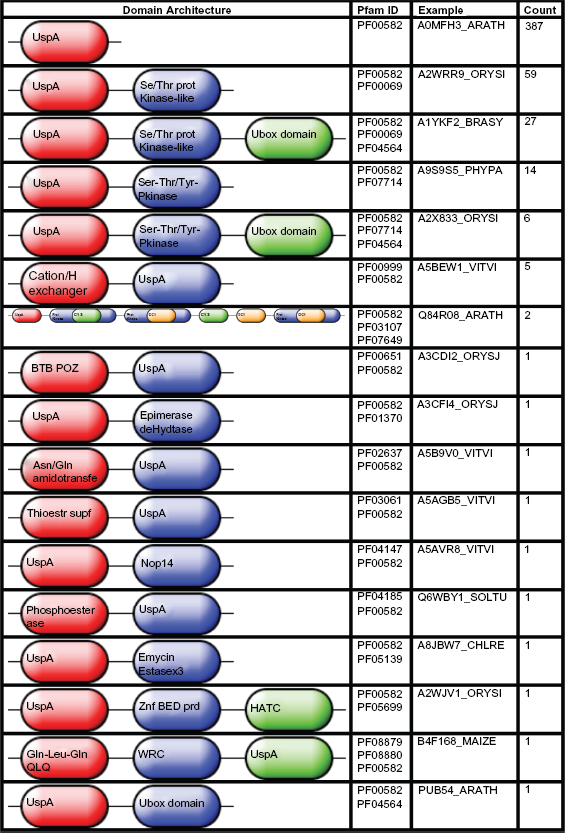
Protein domain architectures, examples and counts in dataset of plant universal stress proteins. Architecture images obtained from InterPro (www.ebi.ac.uk/interpro), an integrated database of predictive protein “signatures” for protein annotation and classification. The examples are UniProt identifiers with abbreviations for the plant taxa as follows—ORYSI:
Orthologous Viridiplantae Drought-Responsive Genes Encoding Universal Stress Proteins
The UniProtKB database cross-references for each viridiplantae USP entry stored in the XML format were extracted to determine the availability of each database annotation across the dataset of entries. Table 4 shows databases that were used to annotate at least 100 USPs. The complete list of 45 cross-references is available in Supplementary File 1. The Gene Ontology, InterPro, NCBI Taxonomy, and Pfam were found in all the 511 UniProt entries. In order to construct a matrix, 40 of the cross-references were selected with references present in all entries removed as well as RefSeq, which had an identical number of entries with Entrez Gene database. The matrix is available in the Supplementary File 1.
Selected UniProt cross-reference resources linked to plant universal stress proteins.

Twelve USP sequences were annotated with both the ArrayExpress and Ortholog Matrix Project (OMA) Browser (Fig. 4). Three

Visualization of matrix of availability of annotation with 40 external database references for selected plant universal stress proteins in UniProt. Description of column headings is documented in Supplementary File 1. Notes: Red, presence of database annotation; Green, absence of database annotation.
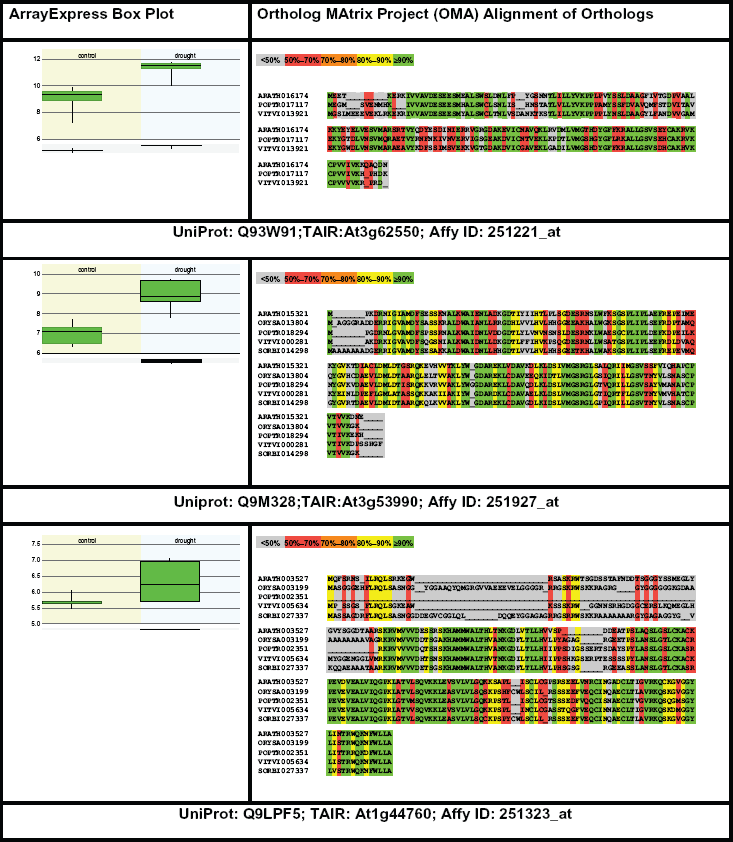
Gene expression and protein sequence alignment of
Visual inspection of the alignments showed that the G-2X-G-9X-G (S/T) motif for small phosphoryl/ribosyl-binding residues of Adenosine Triphosphate (ATP) 49 was present in Q9M328 and Q93W91 but absent in Q9LPF5. Additional homologous sequences for the drought-responsive proteins provided by PLAZA 36 and ClustalW2 generated sequence alignments can be found in the Supplementary File 2. The multiple sequence alignment for 16 homologous sequences including drought responsive ATP-binding motif containing At3g62550 is presented in Figure 6. The conserved Aspartate (D) residue in position 12 of At3g62550 is known to be involved in adenine binding in ATP-binding USPs.15,50

Multiple sequence alignment of drought-responsive
Viridiplantae Universal Stress Protein Gene Transcripts Derived from Drought Conditions
A total of 1561 ESTs clustered into 360 singletons and 185 Transcript Assembles from 137 unique viridiplantae members (82 genera) and annotated with text “universal stress protein” were obtained from the TIGR Plant Transcript Assemblies (Supplementary File 1).
Plant genera represented in universal stress protein gene transcripts dataset.

A dataset of 80 simple sequence repeats (SSRs) linked to 20 singletons and 47 transcript assembles was constructed (Supplementary File 1). A total of 31 types of SSRs (3 uninucleotide; 7 dinucleotides; 16 trinucleotides; 1 tetranucleotide; 3 pentanucleotides; and 1 hexa-nucleotides) were retrieved (Table 6). The transcript count associated with each SSRs was also determined to identify potential unique EST-SSR markers. For example, the dinucleotide TA was unique for singleton DY959747 from
Simple Sequence Repeats (SSR) linked to universal stress protein gene transcripts.

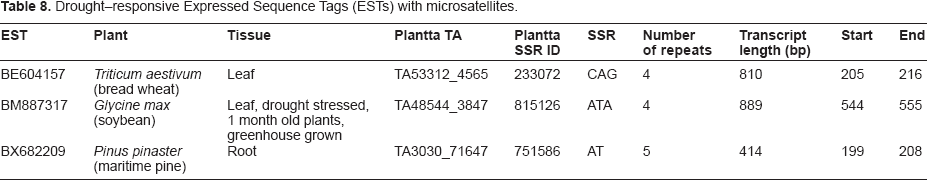
The bioinformatics strategy retrieved 32 drought-responsive ESTs from 7 plant genera
Drought-annotated plant Expressed Sequence Tags (ESTs)
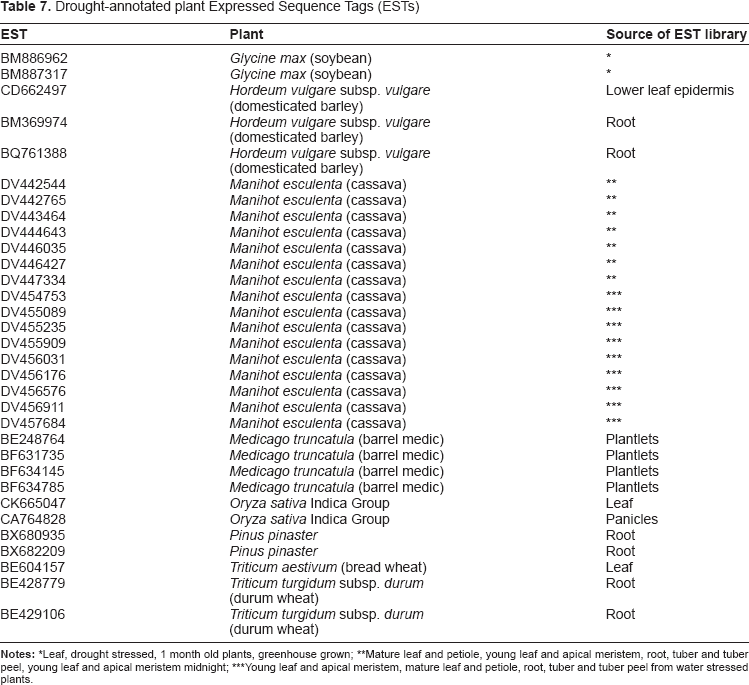
Leaf, drought stressed, 1 month old plants, greenhouse grown;
Mature leaf and petiole, young leaf and apical meristem, root, tuber and tuber peel, young leaf and apical meristem midnight;
Young leaf and apical meristem, mature leaf and petiole, root, tuber and tuber peel from water stressed plants.
Discussion
Plants are continuously exposed to abiotic and biotic stresses that require adaptation for survival. The availability of genomic sequences from a variety of viridiplantae has facilitated the dissection of the molecular, cellular and developmental responses to environmental stresses including drought. 37 Our investigation demonstrates the benefits of integrating data on universal stress proteins from comprehensive protein and transcript databases. The value-added and prioritized datasets produced presents new opportunities to better investigate the function of universal stress proteins from diverse plants. According to the focus of the investigation, the protein and gene transcript datasets are discussed in the context of response to drought and salt stress.
Construction of Dataset of Viridiplantae Universal Stress Proteins
We have retrieved, mined and integrated comprehensive functional annotation data on 511 universal stress protein and 1561 ESTs sequences from the viridiplantae. A total of 161 plants with unique NCBI Taxonomy Identifier were associated with the sequences. Thus, we have provided a catalog of protein and gene transcripts from model and non-model plant species those of importance in agriculture, ecology, industry and alternative energy. A catalog limited to
The bioinformatics strategy extracted functional annotation data from comprehensive public domain protein and gene transcript databases. The Pfam protein family database 36 served as the source of protein sequences for which their functional annotation data in the UniProt protein resource 26 were extracted and integrated with other specialized databases including those storing data on gene expression 38 and protein sequence evolution. 34 We also extracted functional annotation data from the Plantta EST resource, since ESTs are a source of genomic information especially for plants without complete genome sequencing projects. The bioinformatics approach presented could be useful for other researchers interested in other protein families.
The particular function of a protein depends on its combination of domains. In general, the presence of the USP domain may provide the ability for the function of the other domain to be expressed under stress conditions. The USP domain appears as a single domain in small USP proteins (~14–15 kDa), as two domains arranged in tandem in larger USP proteins (~30 kDa), or as one or two USP domains together with other functional domains.9,13 Our analysis extracted and organized the domain combinations present in the 511 plant USPs thereby providing function-categorized subsets of the dataset. The categories can be investigated for shared function and regulation. Protein phosphorylation by kinases is a known pathway utilized by plants to response to osmotic stress.52,53
Five proteins had annotation for the sodium/hydrogen exchanger family domain (PF00999), a domain for transport of sodium ions either out of cell or organelles in exchange for hydrogen ions to prevent toxic accumulation of sodium ions.54,55 The
Nine of the 12 protein sequences with tandem USP domains were from green algae. There are currently a limited number of reports on functional characterization of proteins with tandem USP domains.10,42,43 In
Orthologous Viridiplantae Drought-Responsive Genes Encoding Universal Stress Proteins
Cross-referencing of specialized databases to a protein sequence entry in UniProtKB provides additional functional annotation that can help accelerate selection of plant USPs for characterization. The UniProtKB provides links to at least 126 specialized resources including plant bioinformatics databases such as The Arabidopsis Resource (TAIR),
45
Gramene,
46
and EnsemblPlants.
47
We have integrated available database cross-references to provide a visual view of databases across the viridiplantae USPs analyzed. The utility of such view was demonstrated on a subset of proteins that were annotated with ArrayExpress
45
and Ortholog MAtrix Project (OMA) Browser.
34
This view enabled us to easily identify Q9SW11 (U-box domain-containing protein 35; At4g25160, PUB35) as an enzyme based on the presence of the Enzyme Commission (EC) number (Fig. 4: Column 4, Row 10). The U-box domain for regulated protein ubiquitination and degradation is a modified RING-finger domain involved in protein that lacks metal-binding ability.
48
Comparative structural and functional assays could reveal the interactions of the USP domain and the enzyme domains present in Q9SW11. Orthologous drought-responsive universal stress proteins could be candidates to engineer desired phenotypes in plants. Our analyses identified three
Viridiplantae Universal Stress Protein Gene Transcripts Derived from Drought Conditions
Expressed Sequence Tags generated from stress-challenged plant tissues have been used as high quality transcripts to discover genes, identify candidate stress-responsive genes/transcripts and identify functional markers such as genic microsatellites and single nucleotide polymorphisms.49–51 The effects of SSR type as well as number of repeats on gene regulation, transcription and protein function are poorly understood in plants when compared to human or animal systems. 51 In this article we report automatic extraction of information on simple sequence repeats (SSRs) associated with 1561 ESTs in the Plantta resource. 27 Our analysis identified candidate USP gene transcripts in multiple plants (Supplementary File 1 and Table 5); organized the SSRs into types (Table 6), drought-annotated USP ESTs (Table 7) and USP EST-SSRs from drought-stress tissues (Table 8). The majority (49 of 80) of the USP EST-SSRs was the trinucleotide type, which has been reported to be the most abundant in rice, wheat and barley52,53 as well as peanut 54 and citrus. 55 All together, our analyses provide a comprehensive collection of USP ESTs including those responsive to drought. We have clustered the plant genera based on the number of species to facilitate investigating the EST-SSR and EST-Single Nucleotide Polymorphisms (SNPs) in USP genes for comparative mapping, transferability, genetic diversity and plant improvement.
Drought-responsive Expressed Sequence Tags (ESTs) with microsatellites.
Conclusions
The molecular mechanisms by which genes encoding the universal stress protein domain are able to confer in plants the ability to respond and adapt to environmental changes are not well defined. We have computationally retrieved, mined and integrated functional annotations on protein and gene transcripts that encode the universal stress protein domain. The datasets from cross-database mining provide organized resources for the characterization of USP genes as useful targets for engineering plant varieties tolerant to unfavorable environmental conditions.
Disclosures
This manuscript has been read and approved by all authors. This paper is unique and not under consideration by any other publication and has not been published elsewhere. The authors and peer reviewers report no conflicts of interest. The authors confirm that they have permission to reproduce any copyrighted material.
Footnotes
Acknowledgments
Mississippi NSF-EPSCoR Award (EPS-0903787); Research Centers in Minority Institutions (RCMI)—Center for Environmental Health at Jackson State University (NIH-NCRR G12RR013459); Pittsburgh Supercomputing Center's National Resource for Biomedical Supercomputing (T36 GM008789); US Department of Homeland Security Science and Technology Directorate (2007-ST-104-000007; 2009-ST-062-000014; 2009-ST-104-000021). SSS was a Louis Stokes Mississippi Alliance for Minority Participation (LSMAMP) Fellow in 2005 and is currently a PhD Candidate in the Environmental Science PhD Program at Jackson State University. Disclaimer: The views and conclusions contained in this document are those of the authors and should not be interpreted as necessarily representing the official policies, either expressed or implied, of the funding agencies.
