Abstract
Using comparative genomics and in-silico analyses, we previously identified a new member of the prion-protein (PrP) family, the gene SPRN, encoding the protein Shadoo (Sho), and suggested its functions might overlap with those of PrP. Extended bioinformatics and conceptual biology studies to elucidate Sho's functions now reveal Sho has a conserved RGG-box motif, a well-known RNA-binding motif characterized in proteins such as FragileX Mental Retardation Protein. We report a systematic comparative analysis of RGG-box containing proteins which highlights the motif's functional versatility and supports the suggestion that Sho plays a dual role in cell signaling and RNA binding in brain. These findings provide a further link to PrP, which has well-characterized RNA-binding properties.
Keywords
Introduction
In 2003 we discovered a new gene, SPRN, which codes for a 151-residue protein (including N- and C-terminal signal sequences) with topographical similarities unique to prion protein (PrP), and which is highly conserved between fish and mammals (for an analysis of the similarities between PrP and Sho see Premzl et al. 2003). We called this new protein Shadoo (Sho; shadow of prion protein). Like PrP, Sho is most abundant in brain (Premzl et al. 2003; Uboldi et al. 2006; Watts et al. 2007). Although the functions of Sho are as yet little characterized, it has been shown by gain and loss of function experiments (RNAi and overexpression) to be essential for CNS development in zebrafish (L. Sangiorgio, University of Milan, pers. comm.) and may have a neuroprotective effect similar to PrP (Watts et al. 2007). While PrP is notorious for its association with the transmissible spongiform encephalopathies such as Creutzfeldt Jacob Disease and Bovine Spongiform Encephelopathy (Mad Cow Disease) it has become clear in recent years that PrP has a range of normal functions, including in neurogenesis and neural plasticity (Kanaani et al. 2005; Moya et al. 2005; Santuccione et al. 2005; Steele et al. 2006). In defining the natural functions of Sho we are investigating links of the protein's properties to those of other proteins, including PrP.
Here we report our finding that Sho has a conserved ‘RGG-box’ motif (Kiledjian and Dreyfuss, 1992) defined as a sequence of closely spaced Arg-Gly-Gly (RGG) repeats interspersed with other, often aromatic, amino acids. The RGG box proteins are one class of RNA-binding proteins (RBPs) involved in various aspects of RNA processing, including splicing, stabilizing, transport and translation of mRNAs (Burd and Dreyfuss, 1994). In addition to being an RNA-binding motif, the RGG box of some proteins is known to mediate interactions with other proteins; for a recent detailed example see Lukasiewicz et al. (2007).
The capacity to bind RNA constitutes another point of similarity with PrP which is known to bind RNA and DNA (Grossman et al. 2003). While it has been established that PrP is competent to bind nucleic acid, it is also known to bind many other ligands including polyanionic glycosaminoglycans (‘GAGs’). Given its propensity to bind polyanions, it is currently unclear whether the binding of nucleic acids is biologically relevant, that is, whether a normal function of PrP involves this type of interaction or whether binding observed experimentally may be a non-specific interaction. However, others have observed that PrP modifies DNA structure in a manner similar to proteins involved in transcriptional regulation (Bera et al. 2007) and have queried whether PrP may be involved in the biogenesis or transport of nucleic acid (Lima et al. 2006).
The approach we have used here is underpinned by a novel combination of comparative genomics (Hedges and Kumar, 2002) and conceptual biology (Blagosklonny and Pardee, 2002). By comparing Sho sequences from species ranging from fish to human and integrating these results with those of a comprehensive analysis of published sequence data and experimental findings, we have been able to put our observations into the broader context of RGG-box proteins. This has allowed us to formulate functional hypotheses for Sho.
Materials and Methods
The amino acid sequences of 12 Sho proteins ranging from fish to human were used in this study. Ten of these sequences are available from GenBank [Homo sapiens Np_001012526, Canis lupus familiaris CAJ43798, Bos taurus CAJ43799, Mus musculus NP_898970, Monodelphis domestica CAF43800, Gallus gallus CAJ43796, Xenopus tropicalis CAJ43801, Danio rerio CAD35503, Takifugu rubripes CAG34291, Tetraodon nigroviridis CAG30521]. The sequences for Ornithorhynchus anatinus (platypus), M. domestica (American opossum), G. gallus (chicken), and X. tropicalis were initially extracted from the genomic databases (N. Chakka, unpublished work of this group). The sequences for X. tropicalis and X. laevis were also verified experimentally (T. Vassilieva and N. Chakka, unpublished work of this group). The sequences were aligned using ClustalW (Chenna et al. 2003). Subsequent manual adjustments in the N-terminal region were made to the alignment.
The Swiss-Prot protein database was searched using the program Prosite (Hofmann et al. 1999) http://au.expasy.org/ for known motifs within the Sho sequences. We also searched Swiss-Prot for all proteins that have an RGG-box motif, which we defined as being a sequence of at least 3 RGG repeats with no more than 6 residues between the repeats. This search produced 10 archaeal, 229 bacterial, 14 viral and 1632 eukaryotic sequences, within which there are 607 fungal, 300 plant and 70 human sequences. Examination of the human sequences showed that some well-known RGG-box proteins had not been picked up by this search. The search was then broadened to include proteins with 2 RGG repeats separated by 9, 8, 7, 6 or 5 residues. The results were visually inspected and those proteins with at least one ‘RG’ between the RGG repeats were included in our list. All uncharacterized proteins or redundant sequences were excluded. The remaining human proteins are collected in Table S1. We have only recorded the sequence beginning and ending with an RGG repeat. It should be noted that the functional RGG box may extend beyond the sequence denoted in Table S1. The RGG sequences were subsequently aligned using ClustalW.
Results and Discussion
Sho—RGG box
A sequence alignment of the N-terminal segment from residue 25 to 42 (the mature protein starts at residue 24) of Shos from different species (Fig. 1) reveals a strictly conserved arginine methylation site (GGRGG) (Lee and Bedford, 2002) at the beginning of a cluster of RGG repeats.
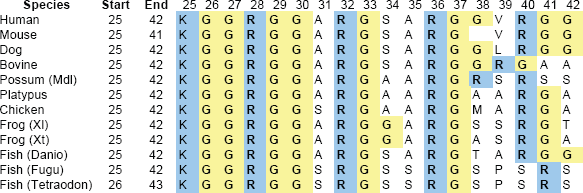
Alignment of the RGG-box sequence at the N-terminal end of Shos from fish to mammals. LHS are sequence numbers. Mdl, Monodelphis domestica; Xl, Xenopus laevis; Xt, Xenopus tropicalis; Danio, Danio rerio; Fugu, Fugu rubripes, Tetraodon, Tetraodon nigroviridis. Note that region starts with completely conserved KGG triplet. Complete RGG triplets are bolded.
In Shos from human and most other Eutherian mammals there are three RGG repeats, with the first and third separated by 9 residues (
Comparative Analysis of Rgg-Box Proteins—Structure and Composition
Proteins with an RGG-box motif, as defined for the purpose of this study (Methods), are presented in Supplementary Information Table S1. Most (#2–#34) are known to have an RNA-binding function. The subset of proteins highlighted in this paper is presented in Table 1. Analysis of all the proteins listed in Table S1 reveals that the RGG box is generally found at the end of the protein sequence, particularly at the C-terminus (Fig. 2A) and is mostly 10–19 residues in length (Fig. 2B). We found a slight preference for RGG repeats to be separated by 9 intervening residues (RGG-X9-RGG), as in Sho, but overall the spacing is variable (Fig. 2C).
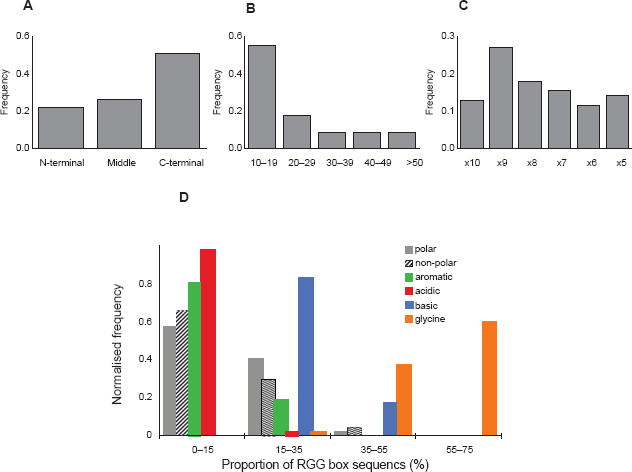
Frequency histograms of structural and compositional features of the 45 RGG sequences surveyed (Table S1).
Subset of RGG-box proteins (see Table SI in Supplementary Information for full list).

number of protein as appears in Table S1.
RNA-binding motifs in addition to the RGG box.
RRM = 80–90 amino acid sequence containing a RNP-1 (octapeptide) and RNP-2 (6 amino acid) consensus sequences.
K homology region as in hnRNP K.
RNA binding.
Protein binding.
The amino acid composition of the sequences was analysed by calculating the proportion of basic (Arg, Lys and His), acidic (Glu and Asp), aromatic (Phe, Trp and Tyr), polar (Ser, Thr, Asn, Gln and Cys), Gly and the other non-polar amino acids (Ala, Val, Leu, Ile, Met and Pro) which make up each sequence and then producing a frequency distribution for the entire set of proteins (Fig. 2D). As expected, a majority of sequences is Gly rich, with peaks in frequency at 50%–60% Gly composition while basic residues peak at 20%–30%. Although a significant number of sequences do not contain an aromatic acid between the RGG repeats, it is possible that there are aromatic residues in close sequence or spatial proximity to this domain. Very few sequences contain acidic residues.
The Sho RGG sequence conforms to these general structural and compositional parameters. It is found at the end of the protein (N-terminus), is 15 residues long and is comprised of 47% Gly, 27% basic, 20% non-polar and 7% polar residues, and has no acidic or aromatic residues. We aligned the RGG sequence of Sho against other sequences with RGG-X9-RGG spacing in order to identify those most similar to Sho (Fig. 3). Several sequences have 50% or more residues identical to those in the Sho RGG box. Experimental studies have demonstrated that the Fragile X Mental Retardation Protein (FMRP) (#32, Table S1) (Zanotti et al. 2006) and the Herpes Simplex protein ICP27 (Mears and Rice, 1996) bind RNA with their RGG boxes which, like Sho, consist of 2 RGG repeats separated by 9 residues.

Alignment of the RGG box of proteins with RGG-X9-RGG spacing. The number of the residue at the start and end of the sequence is given, as well as the total number of exact residue matches (#) to Sho.
Overall, our comparative analysis supports the prediction that the RGG box of Sho is competent to bind RNA.
Sho—Predicted Arginine Methylation and Phosphorylation Sites
The Arg methylation site in Sho is completely conserved in all species from fish to human, suggesting functional importance. Arginine methylation is a common post-translational modification in RGG-box domains (Liu and Dreyfuss, 1995) which affects protein-protein interactions (Boisvert et al. 2005) and RNA binding (Dolzhanskaya et al. 2006). It influences diverse cellular processes, including cellular location of proteins (Passos et al. 2006) transcription, processing and transport of mRNAs (Yu et al. 2004) and signaling pathways (Boisvert et al. 2005).
Phosphorylation is another common post-translational modification found in RBPs. Methylation and phosphorylation mechanisms co-regulate a number of RGG-box proteins, possibly including Sho; again for a detailed example see Lukasiewicz et al. (2007). We identified 3 potential protein kinase C (PKC) phosphorylation sites (SAR (34–36 huSho), SLR (63–65 huSho) and SYR (119–121 huSho)) for Sho. One of these, SAR34–36, is within the RGG box and is found in all the Eutherian mammal sequences analysed (Fig. S1 in Supplementary Information). Phosphorylation of Ser34 would have a direct affect on the structure of the RGG box and most likely affect its function. Although the phosphorylation-site motifs are patterns with a high probability of random occurrence it is interesting to note that the presence of at least one phosphorylation site has been experimentally confirmed in 70% of the RGG-box proteins surveyed (Table S2 in Supplementary Information). This is a high proportion even taking into account the over-representation of nuclear proteins in the phosphoproteome (Olsen et al. 2006) and leads us to suggest that phosphorylation is particularly prevalent in RGG-box proteins. The finding of potential methylation and phosphorylation sites in Sho is another point of similarity with other RGG-box proteins. The existence of phosphorylation sites within Sho raises the possibility that Sho may be involved in a signaling pathway that is regulated by phosphorylation.
Functional Significance
Sho differs from most of the other proteins surveyed in that it has no other RNA-binding motifs. This is unusual but not unique as hnRNP U (#12; Table S1) has no other RNA-binding motif apart from the RGG box. The RGG box is typically associated with binding to single-stranded nucleic acids, (Zhang and Grosse, 1997) whereas additional RNA-binding motifs may allow binding of a broader range of RNA targets as is the case for nucleolin (#33, Table S1) (Ghisolfiet al. 1992). The inherent flexibility of the RGG box (Ramos et al. 2003) can also enable binding to several RNA targets, as has been shown for FMRP (Darnell et al. 2004) which binds to many RNA targets but an affinity for RNA that forms a stable G-quartet structure (Menon and Mihailescu, 2007; Ramos et al. 2003). As Sho lacks other RNA-binding motifs, we expect it to bind single-stranded nucleic acid, and potentially a range of such targets, as for FMRP.
The RGG box is a positively charged domain known to interact electrostatically with other proteins and anionic molecules. A well-characterized example is the RGG box of the yeast protein Npl3p which docks with the kinase Sky1 (Lukasiewicz et al. 2007). A non-protein example is provided by the intracellular hyaluronan binding protein (HAPB4) (#21, Table S1) which has high sequence similarity to Sho (Fig. 3). The RGG domain of HAPB4 also constitutes a glycosaminoglycan (‘GAG’) binding motif (R/K–X(7)-R/K) (Yang et al. 1994) and has been found to bind strongly and specifically to hyaluronan and weakly to RNA (Huang et al. 2000). Although it is not surprising to find this motif in an Arg-rich sequence (in fact it is present in most of the proteins included in Table S1), it has particular relevance in the case of Sho, given its cellular location.
The cellular location of Sho will determine its opportunities to bind RNA and whether this is its primary function. We originally predicted Sho to be a GPI-anchored protein (Premzl et al. 2003). This has now been confirmed in mouse (Watts et al. 2007) and for a Sho-like protein (Sho2) (Premzl et al. 2004; Strumbo et al. 2006) in zebrafish (Miesbauer et al. 2006). However, some GPI-anchored proteins, including PrP, undergo anchor cleavage (‘shedding’), (Parkin et al. 2004; Zhang et al. 2005) resulting in formation of soluble proteins which can relocate to other cellular destinations and are capable of performing multiple functions (Campana et al. 2005). While the cell surface is one likely location for Sho, it may be a multifunctional protein found in other cellular locations as well, as for PrP. If Sho sheds its GPI anchor or undergoes proteolytic cleavage before attachment to the cell membrane (Watts et al. 2007), the RGG-box domain would be available for functional roles intracellularly. Other RGG-box proteins are known to have multiple cellular locations, for example, nucleolin and the Ewing Sarcoma (EWS) protein (#26, Table S1) are found on the cell surface as well as in the nucleus and cytoplasm. In fact, there is growing evidence that some RNA-binding proteins have additional roles as cell surface receptors (Bajenova et al. 2003; Belyanskaya et al. 2003; Hirano et al. 2005) and in signaling pathways as noted for the ras GTPase activating protein binding protein 1 (Kennedy et al. 2001).
Attached to the cell surface, Sho would be positioned to act as a receptor for ligands found at the cell surface, including nucleic acids, as suggested for EWS (Belyanskaya et al. 2001). Sho may, therefore, have a role in cell signaling, similar to PrP which binds the neural cell adhesion molecule and thus participates in the tyrosine kinase fyn signaling pathway leading to neurite outgrowth (Santuccione et al. 2005). Alternatively, in this location Sho may bind other anionic ligands such as the GAG, hyaluronan, which is known to bind another GPI-anchored protein, brevican, and is involved in the structural plasticity of neural tissue (Rauch, 2004). It is interesting to note that PrP also binds GAGs including hyaluronan and heparin (Pan et al. 2002) and that GAGs may facilitate the conversion of the normal cellular PrP to the isoform found in prion disease (Yin et al. 2007).
If Sho were to shed its GPI anchor and re-enter the cell or if a segment of the N-terminal region incorporating the RGG domain was cleaved off prior to expression at the cell surface, the RGG box would be available to interact with cellular RNA. Indeed, as a small protein of no more than 123 residues, Sho would be capable of diffusing in and out of the nucleus (Cyert, 2001) and shuttling RNA from the nucleus to the cytoplasm. This is a function normally performed by RNA-binding proteins involved in neural plasticity, which participate in the biogenesis of mRNA, its transport to dendrites and repression of translation pending appropriate neural stimulation (Ule and Darnell, 2006).
Conclusion
In summary, we have observed that Sho has a conserved RGG-box domain with similar composition to other known RGG-box proteins. We predict that this domain has functional significance and may mediate some of the neural functions already indicated for Sho. Our analysis leads us to postulate that Sho is an RNA-binding protein which may also play a role in cell signaling. Our initial experiments to test the prediction have shown Sho RGG box peptide is competent to bind RNA but further work is required to characterize the interaction.
The discovery of the RGG box in Sho opens new avenues for investigating its function and potential functional overlap with PrP. It is known that PrP plays a role in neural plasticity through its involvement in neural signaling pathways. Here we suggest that Sho may bind mRNA directly and thus play a role in neural plasticity similar to other neural RBPs.
Note
Beck et al. (J. Med. Genet. online 19/9/08) have reported an association of a null allele of SPRN with variant CJD.
Disclosurez
The authors report no conflicts of interest.
Footnotes
Acknowledgements
We acknowledge support from the Australian National University Institute for Advanced Studies (IAS) block grant.
Supplementary Information
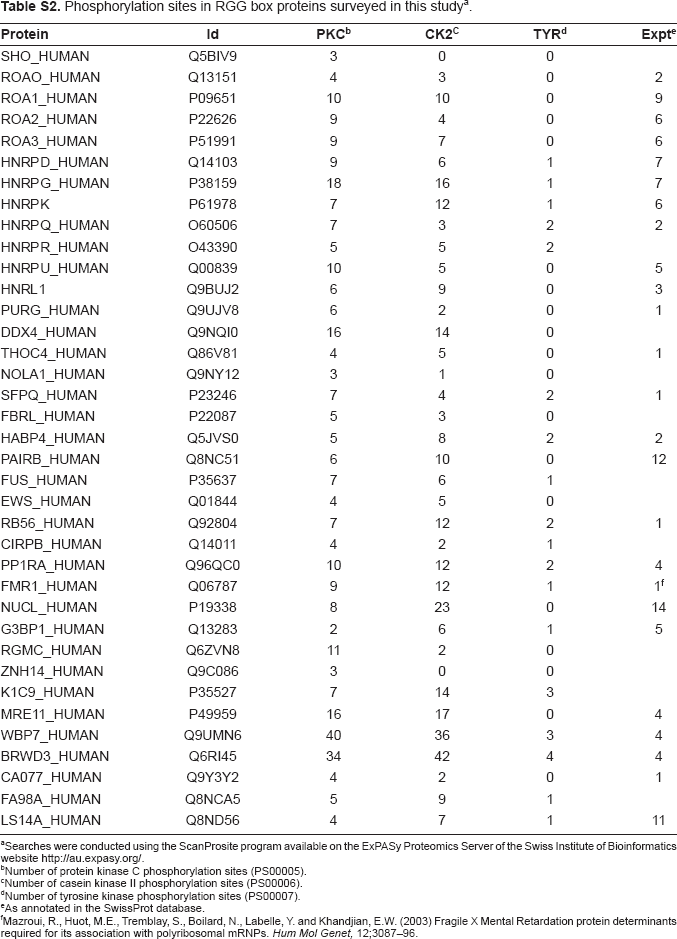
Phosphorylation sites in RGG box proteins surveyed in this studya.
| Protein | Id | PKC b | CK2 c | TYR d | Expt e |
|---|---|---|---|---|---|
| SHO_HUMAN | Q5BIV9 | 3 | 0 | 0 | |
| ROAO_HUMAN | Q13151 | 4 | 3 | 0 | 2 |
| ROA1_HUMAN | P09651 | 10 | 10 | 0 | 9 |
| ROA2_HUMAN | P22626 | 9 | 4 | 0 | 6 |
| ROA3_HUMAN | P51991 | 9 | 7 | 0 | 6 |
| HNRPD_HUMAN | Q14103 | 9 | 6 | 1 | 7 |
| HNRPG_HUMAN | P38159 | 18 | 16 | 1 | 7 |
| HNRPK | P61978 | 7 | 12 | 1 | 6 |
| HNRPQ_HUMAN | O60506 | 7 | 3 | 2 | 2 |
| HNRPR_HUMAN | O43390 | 5 | 5 | 2 | |
| HNRPU_HUMAN | Q00839 | 10 | 5 | 0 | 5 |
| HNRL1 | Q9BUJ2 | 6 | 9 | 0 | 3 |
| PURG_HUMAN | Q9UJV8 | 6 | 2 | 0 | 1 |
| DDX4_HUMAN | Q9NQI0 | 16 | 14 | 0 | |
| THOC4_HUMAN | Q86V81 | 4 | 5 | 0 | 1 |
| NOLA1_HUMAN | Q9NY12 | 3 | 1 | 0 | |
| SFPQ_HUMAN | P23246 | 7 | 4 | 2 | 1 |
| FBRL_HUMAN | P22087 | 5 | 3 | 0 | |
| HABP4_HUMAN | Q5JVS0 | 5 | 8 | 2 | 2 |
| PAIRB_HUMAN | Q8NC51 | 6 | 10 | 0 | 12 |
| FUS_HUMAN | P35637 | 7 | 6 | 1 | |
| EWS_HUMAN | Q01844 | 4 | 5 | 0 | |
| RB56_HUMAN | Q92804 | 7 | 12 | 2 | 1 |
| CIRPB_HUMAN | Q14011 | 4 | 2 | 1 | |
| PP1RA_HUMAN | Q96QC0 | 10 | 12 | 2 | 4 |
| FMR1_HUMAN | Q06787 | 9 | 12 | 1 | 1 f |
| NUCL_HUMAN | P19338 | 8 | 23 | 0 | 14 |
| G3BP1_HUMAN | Q13283 | 2 | 6 | 1 | 5 |
| RGMC_HUMAN | Q6ZVN8 | 11 | 2 | 0 | |
| ZNH14_HUMAN | Q9C086 | 3 | 0 | 0 | |
| K1C9_HUMAN | P35527 | 7 | 14 | 3 | |
| MRE11_HUMAN | P49959 | 16 | 17 | 0 | 4 |
| WBP7_HUMAN | Q9UMN6 | 40 | 36 | 3 | 4 |
| BRWD3_HUMAN | Q6RI45 | 34 | 42 | 4 | 4 |
| CA077_HUMAN | Q9Y3Y2 | 4 | 2 | 0 | 1 |
| FA98A_HUMAN | Q8NCA5 | 5 | 9 | 1 | |
| LS14A_HUMAN | Q8ND56 | 4 | 7 | 1 | 11 |
Searches were conducted using the ScanProsite program available on the ExPASy Proteomics Server of the Swiss Institute of Bioinformatics website http://au.expasy.org/.
Number of protein kinase C phosphorylation sites (PS00005).
Number of casein kinase II phosphorylation sites (PS00006).
Number of tyrosine kinase phosphorylation sites (PS00007).
As annotated in the SwissProt database.
Mazroui, R., Huot, M.E., Tremblay, S., Boilard, N., Labelle, Y. and Khandjian, E.W. (2003) Fragile X Mental Retardation protein determinants required for its association with polyribosomal mRNPs. Hum Mol Genet, 12;3087–96.
