Abstract
Anaplastic astrocytoma (AA) is a grade III glioma that often occurs in middle-aged patients and presents a uniformly poor prognosis. A small subpopulation of cancer stem cells (CSCs) possessing a self-renewing capacity is reported to be responsible for tumor recurrence and therapeutic resistance. An accumulating amount of microRNAs (miRNA) were found aberrantly expressed in human cancers and regulate CSCs. Efforts have been made to couple miRNAs with nonviral gene delivery approaches to target specific genes in cancer cells. However, the efficiency of delivery of miRNAs to AA-derived CSCs is still an applicability hurdle. The present study aimed to investigate the effectiveness and applicability of nonviral vector-mediated delivery of Let-7a with regard to eradication of AA and AA-derived CSC cells. Herein, our miRNA/mRNA microarray and RT-PCR analysis showed that the expression of Let-7a, a tumor-suppressive miRNA, is inversely correlated with the levels of HMGA2 and Sox2 in the AA side population (SP+) cells. Luciferase reporter assay showed that Let-7a directly targets the 3′-UTRs of HMGA2 in AA-SP+ cells. Knockdown of HMGA2 significantly suppressed the protein expression of Sox2 in AA-SP+ cells, whereas overexpression of HMGA2 upregulated Sox2 expression in AA-SP-. Nuclear localization signal (NLS) peptides can facilitate nuclear targeting of DNA and are used to improve gene delivery. Using polyurethane-short branch polyethylenimine (PU-PEI) as a therapeutic delivery vehicle, we conjugated NLS with Let-7 and successfully delivered it to AA-SP+ cells, resulting in significantly suppressed expression of HMGA2 and Sox2, tumorigenicity, and CSC-like abilities. This treatment facilitated the differentiation of AA-SP+ cells into non-SP CSCs. Furthermore, PU-PEI-mediated delivery of NLS-conjugated Let-7a in AA-SP+ cells suppressed the expression of drug-resistant and antiapoptotic genes, and increased cell sensitivity to radiation. Finally, the in vivo delivery of PU-PEI-NLS-Let-7a significantly suppressed the tumorigenesis of AA-SP+ cells and synergistically improved the survival rate of orthotopically AA-SP+-transplanted immunocompromised mice when combined with radiotherapy. Therefore, PU-PEI-NLS-Let-7a is a potential novel therapeutic approach for AA.
Introduction
Glioma is the most common primary tumor in the central nervous system and is classified from grade I to grade IV based on histopathological and clinical criteria established by the World Health Organization (14,36). Anaplastic astrocytoma (AA), grade III glioma, accounts for approximately 7.5% of all glial tumors. Developments in the standard treatment for AA have been stagnant for decades, with only surgery followed by radiotherapy currently used. The effectiveness of chemotherapy in AA is not fully established, although a possible benefit in prolonging survival has been proposed (43). The median survival of patients with AA remains poor and is only around 2 to 3 years, albeit the advancements in treatment. The failure of current approaches in AA treatment has been attributed to the existence of a subpopulation of cancer cells known as cancer stem cells (CSCs). CSCs have the ability to withstand ionizing radiation and chemotherapies because of the high expression levels of antiapoptotic proteins and a remarkable DNA repair capacity (1,13,24). Advanced treatment approaches to overcome the unique CSC abilities of eliminating anticancer drugs and protecting cells from death are urgently needed. Therefore, therapeutic methods combining nanotechnology-based approaches that can cross the protective barriers of CSCs are considered as a prospective direction for malignant glioma and glioma CSCs.
MicroRNAs (miRNAs) are endogenous small non-protein-coding RNA molecules that negatively regulate gene expression through complementarity to the 3′-untranslated region (3′-UTR) of mRNAs (30). Through targeting important regulatory genes, miRNAs play a pivotal role in diverse biological processes, including development, differentiation, and apoptosis and therefore affect a variety of pathological and physiological responses (2,31). Furthermore, each miRNA can potentially regulate hundreds of mRNAs, and potentially more than one third of human genes are miRNA targets (3,33). Recent studies have shown that miRNAs are implicated in tumorigenesis and function as tumor suppressor or oncogenes. Characterization of the miRNA expression patterns in cancer cells is thought to have substantial value for diagnosis and prognosis as well as for eventual therapeutic interventions. Various miRNAs, such as miR-21 (7), miR-221/222 (18), miR-125 (44), and miR-10b (17), have been associated with the initiation, progression, and invasive nature of GBM. In contrast, miR-34a (21) and miR-326 (58) have been implicated as tumor-suppressor miRNAs in these tumors. MiR-106a-5p was found recently to be significantly downregulated in AA; the reduced expression of miR-106a-5p (56) is associated with poor survival, suggesting that miR-106a-5p plays a tumor-suppressive role during astrocytoma development and/or progression. However, the nature of the involvement of miRNAs in astrocytomas remains elusive, and thus further research is required to fully understand their contributions to this malignancy.
The Let-7 family of miRNAs is one of the first identified tumor-suppressive miRNAs and has become a prototype of miRNAs that possess tumor-inhibitory effects (20). Reduced expression of the Let-7 family in lung cancer patients is associated with shortened postoperative survival. Overexpression of the Let-7 family was found to induce cell cycle arrest and apoptosis and further suppress the tumor progession of non-small cell lung cancer (47). Molecular mechanism analysis showed that Let-7a, a member of the let-7 family, suppresses tumor growth in nude mice through directly suppressing K-ras and c-Myc in lung cancer cells (22). Overexpression of Let-7c in relatively highly metastatic non-small cell lung cancer cells markedly suppressed their migration and invasion through degrading integrin β3 and mitogen-activated protein kinase kinase kinase kinase and prevented lung cancer metastasis (55). However, the role of the Let-7 family in cancer stem-like cells of AA and malignant glioma is still unclear.
Recent reports suggest that GBMs possess CSC-like properties and present the characteristics of aggressive tumor. Side population (SP) has been suggested as a potential CSC of GBM (16). Presently, viral vectors are the most commonly used vehicles for efficient gene transfer, including the delivery of miRNAs, into most tissues (5,29). The therapeutic effects of adenovirus-mediated corepression of miR221 and miR222 have been applied to GBM and GBM-CSC-like cells (51). Nonviral gene vectors have been recognized as a genetically safe delivery method for specific gene-mediated therapeutic studies and do not have some of the disadvantages of viral vectors (48). In this article, we isolated SP from AA and used comparative miRNA screening in SP and non-SP AA cells and identified Let-7a as a candidate miRNA that plays a role in the regulation of SP stem-like cancer cells in AA. We further applied microarray analysis to identify its downstream targets, HMGA2 and Sox2, and elucidated their roles as well as downstream signaling in Let-7a-mediated suppression of drug resistance and cell survival. We dissected a Let-7a-centered regulatory pathway in regulating the drug-resistant ability of the stem-like AA cells. In addition, it has been shown that cationic polyurethane-short branch polyethylenimine (PU-PEI) exhibits high transfection efficiency with relatively low cytotoxicity (25,34,54). Therefore, we explored the potential utility of PU-PEI as a nonviral vector in the gene delivery of miRNAs for targeting of AA or AA-CSCs.
Materials and Methods
Cell Culture, Isolation of Primary Cells From AA Tissues
All procedures for tissue acquisition followed the guidelines of the Declaration of Helsinki, and all samples were obtained after patients had given informed consent. This protocol was approved by the Institutional Review Board of Taichung Veterans General Hospital (Taichung, Taiwan, CF 13239). Tumor specimens from patients with the preliminary diagnosis of malignant gliomas, determined by frozen section during operation, were handled in accordance with the following procedure after acquiring the formal histological diagnosis. In brief, the five patients included a 43-year-old right-handed female presented to our hospital with intermittent headache, nausea, and vomiting for 2 months; axial contrast enhanced T1-weighted MR revealed a 3.2 χ 3.5-cm tumor lesion over the right frontal area. A 47-year-old right-handed male presented to our hospital with progressive dizziness and unsteady gait for 3 weeks; axial contrast-enhanced T1-weighted MR revealed a 2.5 χ 2.8 cm tumor lesion in the right cerebellum with the fourth ventricle compromise. A 48-year-old right-handed female presented to our emergency room because of seizure attack and loss of consciousness; axial contrast enhanced T1-weighted MR revealed an irregular-shaped tumor lesion about 4.5 cm in diameter over the left frontal-temple area. A 52-year-old right-handed male presented to our hospital with intermittent headache and dizziness for 2 months; axial contrast enhanced T1-weighted MR revealed a 4 χ 4.5-cm tumor lesion over the right frontal-temple area. A 62-year-old right-handed female presented to our hospital with progressive weakness of left limbs for 2 months; axial contrast-enhanced T1-weighted MR revealed an irregular-shaped tumor lesion about 5 cm in diameter over the right frontalparietal area. After surgical removal, the AA tissues were washed three times in glucose containing Hank's balanced salt solution (HBSS; Invitrogen, Carlsbad, CA, USA). The samples were sliced to a 300-mm thickness, and the sliced tissues were immersed in 0.1% (w/w) collagenase (Sevapharma, Hostivar, Czech Republic) containing glucose-containing HBSS for 15 min at 37°C and shaken at 125 rpm in a rotation shaker. AA primary cells were cultured in MEM (Invitrogen) with 10% fetal bovine serum (FBS; Invitrogen), supplemented with 1 mM sodium pyruvate (Sigma-Aldrich, St. Louis, MO, USA), nonessential amino acids, 2 mM l-glutamine (Invitrogen), 100 units/ml penicillin, and 100 μg/ml streptomycin (Invitrogen) under standard culture conditions (37°C, 95% humidified air, 5% CO2). AA primary cells from different patients were cultured, preserved, and evaluated separately.
Detection and Sorting of Side Population (SP) Cell Subsets From AA Cells
Cells were digested with 0.25% trypsin-EDTA (Invitrogen) and then centrifuged for 5 min at 1,000 rpm (115 χ g). The cells were subsequently suspended at 1 χ 106/ml in prewarmed Dulbecco's modified Eagle's medium (DMEM; Invitrogen) with 2% FBS (Invitrogen). Hoechst 33342 dye (Invitrogen) was added at a final concentration of 5 μg/ml in the presence or absence of verapamil (50 μM; Sigma-Aldrich) and was incubated at 37°C for 90 min with intermittent shaking. At the end of the incubation, the cells were washed with ice-cold HBSS with 2% FBS and centrifuged down at 4°C; then they were resuspended in ice-cold HBSS containing 2% FBS. Propidium iodide (Sigma-Aldrich) at a final concentration of 2 μg/ml was added to the cells to gate viable cells. The cells were filtered through a 40-μm cell strainer (BD Biosciences, Franklin Lakes, NJ, USA) to obtain single-cell suspension and then stored in the dark at 4°C before analysis. The Hoechst 33342 dye was excited at 357 nm, and its fluorescence was dual-wavelength analyzed (blue, 402-446 nm; red, 650-670 nm). Analyses were done on flow cytometer (FACS Aria II; BD Biosciences).
Radiation Treatment and Clonogenic Assay
Ionizing irradiation (IR) was delivered from a Theratronic T-1000 cobalt unit (Theratronic International, Inc., Ottawa, Canada) at a dose rate of 1.1 Gy/min (SSD = 57.5 cm). In brief, cells in the control and irradiated groups were exposed to different radiation dosages (0, 2, 4, 6, 8, and 10 Gy). After incubating for 10 days, colonies (>50 cells per colony) were fixed and stained for 20 min with a solution containing crystal violet (Sigma-Aldrich) and methanol (Sigma-Aldrich). Cell survival was determined using a colony formation assay. Plating efficiency (PE) and survival fraction (SF) were calculated as follows: PE = (colony number/number of inoculated cells) χ 100%; SF = colonies counted/[cells seeded × (PE/100)].
Constructions of Nuclear Localization Signal-Conjugated Expression Vector
The experiment procedure for constructions of the nuclear localization signal (NLS) link was as described with modifications (35). In brief, synthesized 5′ protruding GGCC sequences of spontaneous hairpin structure formation single-strand DNA with modification at loop region was synthesized. [5′-d (GGCCTGTCCG CGTTGGCTTATGCC AACGCGGACA), 5′-d (CCGGCTACCTTGCGAGCTT TTGCTCGCAAGG TAG)] The active group binding to the modified base was covalently linked to 4-(N-maleimidomethyl) cyclohexane-1-carboxylic acid N-hydroxysuccinimide ester (Molecular Probes, Eugene, OR, USA). The U6 promoter combined with Let-7a pre-miRNA sequences (5′-TGGGATGAGGT AGTAGGTTGTATAGTTTTAGGGTCACACC CACCACTGGGAGATAACTATACAATCTACTGTCTTTCCTA-3′) and CMV promoter-driven eGFP expression sequences were amplified by polymerase chain reaction with specific primers with additional ApaI restriction enzyme cutting sites in pLV-miRNA plasmid (miR-Scr.: scrambled sequence control; BioSettia, San Diego, CA, USA). The PCR-amplified fragments were digested with ApaI (Promega, Madison, WI, USA), and the digested DNA fragments were recovered from 1.2% low melting gel with TBE electrophoresis buffer (Invitrogen). Spontaneous hairpin loop formation single-strand DNA incubated with the ApaI digested PCR fragments in the ligation buffer, T4 DNA ligase (Invitrogen) was added, and the ligated DNAs were extracted and then incubated with NLS polypeptides (PKKKRKV) with reaction buffer for the formation of NLS-preconjugated DNA sequences. The NLS-linked DNA vectors were examined with 10% polyacrylamide gel (Invitrogen) and were also extracted for further transfection experiments.
Synthesis of PU-PEI
PU-PEI was synthesized as described in our previously published papers (11,35,42). l-Lysine-diisocyanate (LDI; Kyowa Hakko Co. Ltd., Tokyo, Japan) 0.145 g (a) and N,N′-bis-(2-hydrayethyl)-piperazine (Sigma-Aldrich) 0.1024 g (b) were respectively dissolved in 1 mL N,N-dimethylformamide (anhydrous DMF; J. T. Baker, Phillipsburg, NJ, USA) solvent and mixed in a three-neck reaction flask (Corning Inc., Corning, NY, USA) under a dry nitrogen purge, heated at 60°C, and allowed to react for 12 h using a 0.5 wt% dibutyltin dilaurate (Sigma-Aldrich) catalyst. Then an excess amount of methanol (4 ml) was slowly added into the reaction mixture until no unreacted isocyanate was detected. The polyurethane was precipitated and purified in ethyl ether (Sigma-Aldrich) and dried at 40°C under vacuum. The polymers were characterized by Fourier transform infrared spectroscopy (FTIR) and proton nuclear magnetic resonance (1H-NMR). 1H-NMR (400 MHz, DMSOd6, ppm) δ: 2.50-2.71 [-N2(CH2CH2)2], 2.99, 3.9 (-NCH2cH2O-), 3.12 (-NHCH(COOCH3)cH2-), 1,21-1.81 [6H, -CH(COOCH3)CH2CH2CH2CH2-], 2.90 (-CH2CH2NH-), 3.67 (-NHCOOCH3), 3.4 (-COOCH3), 8.01 [-NHCH(COOCH3)CH2-], 3.51 (-CH2NHCOOCH3). PU-sbPEI was synthesized using the aminolysis reaction of polyurethane (c) and small branch PEI (sbPEI; MΩ = 600) in scheme 1. First, 0.1 g polyurethane was dissolved in 1 ml of anhydrous DMF, and 0.6 g sbPEI (d) was dissolved in 0.5 ml MeOH with 1 ml triethylamine (Et3N; Sigma-Aldrich). Two solutions were mixed slowly and allowed to react at 45°C for at least 48 h. The polymer was precipitated in an excess amount of anhydrous ethyl ether. Purification was performed by redissolving the above polymer in 3 ml MeOH and precipitating in 4 ml three times before vacuum drying at 40°C. The polymer (PU-sbPEI) (e) was further characterized by FTIR and 1H-NMR. PU-sbPEI: 1H-NMR (400 MHz, D2O, ppm) δ: 2.48 [-N2(-CH2-CH2)2], 2.91 (-NCH2CH2O-), 3.99 (-NCH2CH2O-), 4.35 [-NHCH (CO-)CH2-], 2.84, 1.32, 1.53 (-CHCH2CH2CH2-), 2.91 (-CH2CH2NH-), 3.56 (-O-CH3), 3.8 (-CONHCH2-), 2.48, 3.14 (PEI:-CH2-CH2-), 3.51 (-CH2NHCOCH-), 8.01 [-NHCH(CONH-)CH2-], 0.95 (PEI:-NH-). The molecular weights of PU and PU-sbPEI were 15 kDa and 36 kDa, respectively, measured by gel permeation chromatography. The percent of grafted PEI to PU was then calculated and showed 85% of the methyl ester from LDI blocks was conjugated.
Microarray Analysis and Bioinformatics
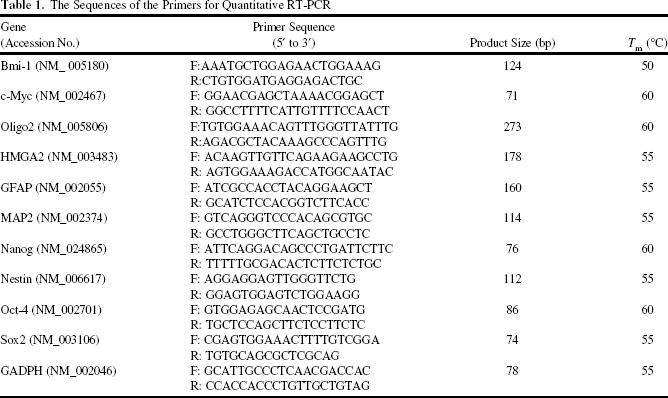
Total RNA was extracted from cells using Trizol reagent (Life Technologies, Bethesda, MD, USA) and the Qiagen RNAeasy (Qiagen, Valencia, CA, USA) column for purification. Total RNA was reverse-transcribed with Superscript II RNase H-reverse transcriptase (Gibco BRL, Grand Island, NY, USA) to generate Cy3- and Cy5-labeled (Amersham Biosciences Co., Piscataway, NJ, USA) cDNA probes for the control and treated samples, respectively. The labeled probes were hybridized to a cDNA microarray containing 10,000 gene clone immobilized cDNA fragments. Fluorescence intensities of Cy3 and Cy5 targets were measured and scanned separately using a GenePix 4000B Array Scanner (Axon Instruments, Burlingame, CA, USA). Data analysis was performed using GenePix Pro 3.0.5.56 (Axon Instruments, Union City, CA, USA) and GeneSpring GX 7.3.1 software (Agilent, Palo Alto, CA, USA). The average linkage distance was used to assess the similarity between two groups of gene expression profiles as described below. The difference in distance between two groups of sample expression profiles to a third was assessed by comparing the corresponding average linkage distances [the mean of all pairwise distances (linkages) between members of the two groups concerned]. The error of such a comparison was estimated by combining the standard errors (the standard deviation of pairwise linkages divided by the square root of the number of linkages) of the average linkage distances involved. Classical multidimensional scaling was performed using the standard function of the R program to provide a visual impression of how the various sample groups are related. Validation of the microarray dataset was performed with quantitative RT-PCR in order to compare the mRNA levels of Let-7a, HMGA2, and Sox2 between the SP+ and SP- AA cells and between the sphere-forming and parental cells. The primer pairs used to validate Let-7a are forward primer 5′-TCTTATCACTCACACAGGAAACCAG-3′ and reverse primer 5′-CTGCACTACATCTCTTTAAGACA AG-3′. The primers used to validate HMGA2 and Sox2 are shown in Table 1.
The Sequences of the Primers for Quantitative RT-PCR
Soft Agar Clonogenicity Assay
Each well (35 mm) of a six-well culture dish (Corning Inc.) was coated with 2 ml of a bottom agar (Sigma-Aldrich) mixture [DMEM, 10% (v/v) FBS, 0.6% (w/v) agar]. After the bottom layer was solidified, 2 ml of the top agar medium mixture [DMEM, 10% (v/v) FBS, 0.3% (w/v) agar] containing 104 cells was added, and the dishes were incubated at 37°C for 4 weeks. Plates were stained with 0.005% crystal violet (Sigma-Aldrich), and then the colonies were counted. The number of total colonies with a diameter >100 μm was counted over five fields per well for a total of 15 fields in triplicate experiments.
Sphere Formation After Serial Passage Assay
AA cells (1 χ 104) were suspended and cultivated in defined serum-free medium composed of DMEM/F12 medium (Gibco), N2 supplement (Gibco), 10 ng/ml human recombinant basic fibroblast growth factor (bFGF), and 10 ng/ml epidermal growth factor (EGF) (R&D Systems, Minneapolis, MN, USA). The spheres were resuspended to generate secondary and tertiary spheres. The number of spheres was counted by microscopy after 14 days.
Let-7a Overexpression by PU-PEI
PU-PEIs were used, following previously established methods (11,35,42). The miRNA Let-7a expression plasmid (pLet-7a) was purchased from Addgene (Cambridge, MA, USA). pLet-7a was dissolved in opti-MEM (Invitrogen) with a final concentration of 800 ng/μl. PU-PEI was dissolved in opti-MEM with final concentrations of 2.4 μg/μl. pLet-7a and PU-PEI were mixed at a 1:1 ratio to reach the final concentration of 400 ng/μl of plasmid DNA and 1.2 μg/μl of PU-PEI, to form the DNA-PU-PEI complexes. The complexes were then incubated at room temperature for 30 min. The pLet-7a expression levels were further confirmed by Taqman microRNA amplification systems with premade pLet-7a-specific primers (forward primer 5′-TCTTATCACTCACACAGGAAACCAG-3′, reverse primer 5′-CTGCACTACATCTCTTTAAGACAAG-3′). The TaqMan® MicroRNA Assays Kit was purchased from Applied Biosystems (Foster City, CA, USA). Cells were grown to about 70% confluency prior to transfection. The complexes were added directly to cells and were removed at 6 h posttransfection. Forty-eight hours later, cells were harvested and subjected to further experiments. The oligonucleotide sequence of the microRNA scramble was shown as 5′-CATTAATTGTCGGACAACTCAAT-3′ and of Let-7a as 5′-UGAGGUAG UAGGUUGUAUAGUU-3′.
HMGA2 Expression Plasmid Constructions
For HMGA2 3′-UTR reporter plasmids, HMGA2 3′-UTR was amplified by PCR with high-fidelity DNA polymerase (Takara Bio Inc., Tokyo, Japan) from extracted human DNAs with the following primer pairs: forward primer 5′-GGGCGCCAACGTTCGATTTCTACCT-3′, reverse primer 5′-CTGTTTTGACCAAATTTATTACTCATTT-3′. The amplified DNA fragments were subcloned into pMIR-REPORT plasmids (Applied Biosystems). For Mut-HMGA2 3′-UTR reporter plasmid construction, four-point mutations were induced into the HMGA2 3′-UTR region by mutagenesis primers: 5′-GCCAACGTTCGATTTTTCGCTAAGC-3′ and 5′-GCTTAGCGAAAAATCGAACGTTGGC-3′. All plasmid sequences were validated by sequencing. The HMGA2 expression plasmid and plasmid vector encoding EGFP and shHMGA2 (Origene Technologies, Rockville, MD, USA) were delivered by PU-PEI into SP+ AA cells following the established methods (described as above) to generate stable HMGA2-overexpressing or HMGA2-knockdown AA cells.
Luciferase Reporter Assay
AA cells were grown in six-well tissue culture dishes to 30% confluence and then cotransfected with firefly luciferase expression vector (pGL3; 20 ng/well; Promega) and the internal control vector: renilla luciferase expression vector (pRL; 2 ng/well; Promega). The transfection procedures were performed using the liposome-mediated transfection method with Lipofectamine 2000 (Invitrogen) as a transfection reagent at a final concentration of 0.38 μl/ml. Sixteen hours after transfection, the cells were lysed, and all cell extracts were subjected to firefly and Renilla luciferase activity detection using a dual luciferase assay (Promega). Luminance signals were detected by luminance recorder (Berthold detection system Sirius single tube luminometer; Pforzheim, Germany). Relative expression levels were measured with the firefly luciferase activities normalized to Renilla luciferase activities.
Real-Time Reverse Transcription-Polymerase Chain Reaction (RT-PCR)
For real-time RT-PCR analysis, the total RNA of cells was extracted using the RNAeasy kit (Qiagen). Briefly, the total RNA (1 μg) from each sample was reversely transcribed using 0.5 μg of oligo dT and 200 U Superscript II RT (Invitrogen). The amplification was carried out in a total volume of 20 μl containing 0.5 μM of each primer, 4 mM MgCl2, 2 ml LightCycler FastStart DNA Master SYBR green I (Roche Diagnostics, Pleasanton, CA, USA), and 2 ml of 1:10 diluted cDNA. The quantification of the unknown samples was performed by LightCycler Relative Quantification Software, version 3.3 (Roche Diagnostics). In each experiment, the GAPDH housekeeping gene was amplified as a reference standard. PCR reactions were prepared and performed in duplicate and heated to 95°C for 10 min followed by 40 cycles of denaturation at 95°C for 10 s, annealing at 55°C for 5 s, and 40 cycles of extension at 72°C for 20 s. Standard curves (cycle threshold values vs. template concentration) were prepared for each target gene and for the endogenous reference (GAPDH) in each sample. The primers and cycling conditions for RT-PCR are shown in Table 1.
In Vivo Orthotopic AA-Derived SP+-Transplanted Nude Mice
All procedures involving animals were in accordance with the institutional animal welfare guidelines and the approved animal protocol of Taipei Veterans General Hospital. Female BALB/c nude mice, 6-8 weeks of age, were obtained from the National Laboratory Animal Center (Taipei, Taiwan). Nude mice were subcutaneously (SC) or intracranially (IC) injected with AA-derived SP+ cells (suspended in physiological saline; Nang Kuang Pharmaceutical, Tainan, Taiwan). For SC injection, the mice were anesthetized with ketamine/xylazine (Pfizer, Dusseldorf, Germany) and then 1 χ 105 AA-derived SP+ cells in a volume of 20 μl per mouse were implanted SC in the lower left flank. The size of tumors was monitored every 5 days up to 30 days and was measured with calipers, and the tumor volume was calculated as (length χ width2)/2. For IC injection, the mouse was anesthetized with ketamine/xylazine and then positioned in a stereotactic frame (David Kopf, Instruments, Tujunga, CA, USA) with small animal ear bars. A burr hole was made using an electronic drill (Dremel, Racine, WI, USA) approximately 3 mm lateral and 1 mm anterior to the intersection of the coronal and sagittal sutures (bregma). Luciferase expression AA cells were established by transfecting pGL4 luciferase expression vector (Promega) with neomycin resistance gene. To establish luciferase expression stable AA cells, AA cells at 30% confluence were transfected by liposome-based gene delivery method. The procedures of Lipofectamine 2000 (Invitrogen) and pGL4/Neo+-forming liposome/DNA complexes were according to manufacturer's suggestions. In brief, 3 μg of pGL4/Neo+ was used to transfect AA cells cultured in 10-cm culture plate. Forty-eight hours after transfection, culture medium was replaced with neomycin (1 mg/ml; Sigma-Aldrich) containing culture medium for 3 weeks. Neomycin-containing culture medium was replaced every 3 days. Individual colonies of AA cells were picked up, and the luciferase activities were determined by luciferase activity assay. AA cells with highest luciferase activities were selected and used for in vivo animal experiment. Cells (1000) were injected using a Hamilton syringe at a depth of 2.5 mm in a volume of 2 μl. The size of tumors was monitored every 7 days up to 6 weeks with bioluminescent imaging. To obtain bioluminescent imaging, the mice were anesthetized with isoflurane (Forawick Vaporizer; Muraco Medical Co., Tokyo, Japan). Seven minutes after intraperitoneal injection of D-luciferin (30 mg/ml; Xenogen Corp., Alameda, CA, USA), the mice were imaged in a Xenogen IVIS50 imaging system (Xenogen Corp.) coupled to a CCD camera. The mice were imaged for 15 min with an acquisition time of 8-10 s. Analysis of bioluminescent imaging data was completed with Living Image software (Xenogen Corp.). Regions of interest were drawn around discrete anatomical areas and used to calculate bioluminescent signals. This signal was expressed and graphed as total flux (photos/second).
Statistical Analysis
Statistical Package for the Social Sciences software version 13.0 (SPSS, Inc., Chicago, IL, USA) was used for all statistical analyses. Results are reported as mean ± SD. Statistical analysis was performed using Student's t-test or a one- or two-way analysis of variance (ANOVA) followed by Tukey's test, as appropriate. The survival rate analysis was performed using log rank test. Results were considered statistically significant at p <0.05.
Results
Isolation of SP Cells From Patient Tissue with AA
Recent studies have demonstrated that tumors contain a small subpopulation of CSCs that possess self-renewing capacities and are responsible for tumor metastasis, recurrence, and therapeutic resistance (59). The use of the Hoechst 33342 dye to identify and isolate SP provides a practical method of CSC isolation and overcomes the pheno-typical marker barrier (23). In the present study, we enriched the AA-CSCs by isolating SP from primary AA cells derived from five patient specimens (patients No. 1 -5). SP cells represent 0.1-2.2% of the viable tumor cells (median SP: 0.4%; Fig. 1A, left). Verapamil, an ATP-binding cassette transporter inhibitor, was able to block Hoechst dye efflux from SP cells and was used to define the SP population (Fig. 1A, right). To further analyze the function of SP cells in these AA cell lines, SP and non-SP (SP+ and SP-) cells were sorted out and subjected to functional assays. As shown in Figure 1B, SP+ AA cells showed significantly more proliferation than the corresponding SP- AA cells. The ability of SP+ and SP- cells to generate spheres and colonies were evaluated by sphere formation and soft agar assays (Fig. 1C, upper). It was difficult to observe floating spheres from SP- cells, while SP+ cells started to form floating spheres after 2 to 3 days of seeding and became the primary spheres of more than 30 cells after 14 days. The colony formation ability of SP+ cells was also significantly better than that of SP- cells. Furthermore, TransWell cell migration and invasion assay showed that the motility of SP+ cells was significantly better than that of SP- cells (Fig. 1C, lower). To assess the chemosensitivity and radio-sensitivity of the SP+ and SP- cells, cells were treated with cisplatin and paclitaxel for 48 h and with radiation up to 10 Gy to evaluate cell viability, which was measured by a cell viability assay (MTT assay kit; Cell Biolabs, Inc., San Diego, CA, USA). Our data show that SP+ cells were more resistant to chemo- and radiotherapy than SP- cells (Fig. 1D, E). Taken together, these results suggest that the SP cancer cells have some intrinsic stem cell properties.

Enrichment of the cancer stem cells (CSCs) by isolating side population (SP) from primary AA cells. (A) SP of primary AA cells, derived from patient specimens, was isolated by FACS in the presence or absence of verapamil. (B) Representative photomicrographs and quantification of cell numbers of SP+ and SP- AA cells. Scale bar: 100 μM; *p <0.05. (C) SP+ and SP- cells were subjected to spheroid formation (top left), soft agar colony formation (top right), migration (bottom left), and invasion (bottom right) assays. The quantitative data are demonstrated in the charts. *p <0.05. (D) SP+ and SP- cells were subjected to treatment with different concentrations of cisplatin and paclitaxel. (E) Radioresistance was then evaluated by the survival fraction. Cell viability was determined by MTT assay. *p <0.05.
Let-7a Negatively Correlates with Expression Levels of HMGA2 and Sox2 in AA Cells
miRNAs appear to target oncogenes, cell cycle regulators, and transcription factors and are involved in many aspects of brain tumor progression (7,17,18,21,44,56,58). In this study, bioinformatics and microarray analyses were used to distinguish miRNAs and their associated candidates of target genes that may play important roles in putative AA-derived CSCs. The three pairs of SP+ and SP- AA cells derived from three patient specimens were subjected to cDNA array analysis, which focused on ESC-related genes, and a miRNA expression array analysis. The genes and miRNAs that were either upregulated (more than twofold) or downregulated (<0.5-fold) in all specimen-derived SP+ and AA cells compared to their counterparts were considered for further analysis. The data revealed 18 miRNAs and 20 genes that are consistently up- or downregulated in SP+ AA cells (data not shown). Using a software screening strategy (the Target Scan program, www.targetscan.org) based on the published literature (53), we performed a comparative analysis of both microarray data sets, from which Let-7a and its targets, human high-mobility group A2 (HMGA2) and sex-determining region Y (SRY)-Box2 (Sox2), were selected for further investigation (Fig. 2A). Validation of the microarray dataset was performed with quantitative RT-PCR in order to compare the mRNA levels of Let-7a, HMGA2, and Sox2 between the SP+ and SP- AA cells and between the sphere-forming and parental cells (Fig. 2B, C). Consistent with the microarray results, statistical analysis indicated a negative correlation between Let-7a and HMGA2/Sox2 expression levels. Let-7a levels were low in SP+ and sphere-forming AA cells but high in SP- and parental AA cells, whereas HMGA2 and Sox2 had inverse mRNA expression patterns. The statistical analysis of the mRNA expression of these genes predicted a strong negative correlation between Let-7a and downstream HMGA2 or Sox2 in the AA cells.
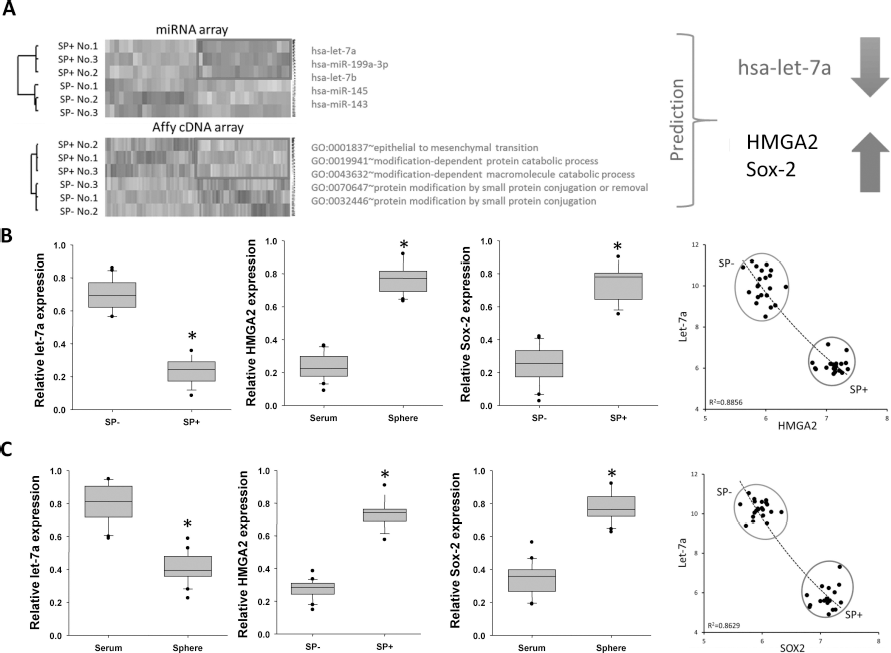
Expression levels of Let-7a negatively correlate with those of HMGA2 and Sox2 in AA. (A) SP+ and SP- cells were isolated from AA patient specimens and subjected to a genome-wide miRNA array and an ESC-related gene expression array. The miRNAs and genes with consistent expression patterns in SP+ cells compared to their respective SP- counterparts are shown in the hierarchical heat map. A literature-based comparison of the two microarrays and a software-based (Targetscan program, www.targetscan.org) comparative analysis of the two datasets indicated that the Let-7a/HMGA2/Sox2 regulatory axis is consistently present in both microarray datasets. (B) Quantitative RT-PCR analysis of Let-7a, HMGA2, and Sox2 expression levels in SP+ and SP- cells. Data shown here are the mean±SD of six independent experiments. *p <0.05 versus SP-. Statistical correlation analysis indicated a highly negative correlation between Let-7a and Sox2 expression as well as between Let-7a and HMGA2 expression. (C) Quantitative RT-PCR analysis of Let-7a, HMGA2, and Sox2 expression levels in parental and sphere cells. Data shown here are the mean±SD of six independent experiments. *p <0.05 versus parental. Two groups of AA patient sample-derived cells (parental and sphere cells) were analyzed for their expression of Let-7a, SOX2, and HMGA2. Statistical correlation analysis indicated a highly negative correlation between Let-7a and Sox2 expression as well as between Let-7a and HMGA2 expression.
Let-7a Directly Targets HMGA2 AA Cells
Previous studies demonstrated that the 3′-UTR of HMGA2 contains seven sequences complementary to the Let-7-family of miRNAs (32). Binding of the complementary sequences by Let-7 miRNAs posttranscriptionally and negatively regulates HMGA2 mRNA and protein expression. However, the regulation of Let-7a and HMGA2/ Sox2 has not been investigated in AA cells. Using the Target Scan program, we found potential Let-7a targeting sites within the 3′-UTR of HMGA2 (Fig. 3A). We then constructed a series of reporter plasmids containing either wild-type (WT) or mutated (Mut) forms of the HMGA2 3′-UTR (Fig. 3B). The reporter assays with WT and Mut Let-7a target sequences showed that Let-7a-mediated inhibition of HMGA2 specifically depends on the Let-7a target sequence, as Let-7a was incapable of inhibiting the luciferase activities of reporter plasmids containing mutated Let-7a target sequences (Fig. 3B). These data demonstrated that Let-7a directly targets HMGA2 through their 3′-UTR regions. In line with these data, overexpression of Let-7a in SP+ and sphere AA cells decreased the protein levels of HMGA2 and SOX2 (Fig. 3C). Functional analysis of these cell lines showed that Let-7a-overexpressed SP+ AA cells had reduced sphere and soft agar colony-formation capabilities as well as inhibited migration and invasion abilities, compared with their parental AA and control AA/pLV [control plasmid (BioSettia)] cells. Moreover, concomitant overexpression of Let-7a and HMGA2 increased the ability of sphere formation, colony formation, migration, and invasion (Fig. 3D). Knockdown of HMGA2 had similar effects to overexpression of Let-7a in SP+ AA cells. Both reduced in vivo tumorigenicity of SP+ AA cells in immunocompromised mice (Fig. 3E). Taken together, these data suggested that Let-7a negatively regulated the expression of HMGA2 in AA-isolated SP+ cells through direct targeting of the 3′-UTR of HMGA2. On the other hand, overexpression of HMGA2 in Let-7a-overexpressing SP+ cells increased the tumorigenicity in vitro as well as in vivo, indicating that HMGA2 is an important target of Let-7a to suppress tumorigenesis.

Let-7a directly targets HMGA2 in AA-isolated SP+ cells. (A) Let-7a target sites were predicted in the 3′-UTR regions of HMGA2 using Target Scan. The nucleotides designed to be mutated are displayed in gray. Both WT and mutated HMGA2 3′-UTR were constructed in Luciferase reporter plasmid. (B) Schematic representation of the constructed 3′-UTR reporter plasmids. The effects of Let-7a on WT or Mut HMGA2 3′-UTRs were assessed by a luciferase activity assay. The data shown are the mean ± SD of three independent experiments. (C) The protein expression levels of Sox2 and HMGA2 in Let-7a-transfected SP+ and sphere cells were analyzed by Western blot. Data shown here are the mean ± SD of six independent experiments. *p <0.05 versus SP+/plv. #p <0.05 versus treated groups. (D) SP+ AA cells transfected with pLV or pLV-Let-7a as well as concomitant overexpression of HMGA2 were subjected to sphere formation, soft agar colony formation, migration and invasion assays. (E) NOD/SCID (BALB/c strain) mice were injected in the right flank with 10,000 cells as indicated, and the growth of the tumor xenografts was monitored by caliper measurements for up to 6 weeks.
The Let-7a/HMGA2 Axis Leads to the Regulation of Sox2
Sox2, a high-mobility group DNA-binding protein, is a critical marker of neural stem cells and a putative marker of brain tumor-associated CSCs (19,45). Sox2 has been shown to play a key role in regulating self-renewal properties in glioma stem cells and pediatric brain tumors, and its expression level negatively correlates with medulloblastoma patient survival (46). In our previous study, we demonstrated that Sox2 is a downstream effector of HMGA2 (12). We further explored the potential roles of HMGA2 and Sox2 in regulating the CSC properties of AA. First, knockdown of HMGA2 in SP+ AA cells did not affect mRNA expression of GFAP and MAP2, but suppressed nestin, Oligo2-, and Sox2 mRNA levels (Fig. 4A, Table 1). Second, overexpression of HMGA2 in SP- AA cells increased the protein levels of Sox2, and knockdown of HMGA2 in SP+ AA cells suppressed Sox2 (Fig. 4B). Functional analysis showed that HMGA2-overexpressed SP- AA cells increased the number of spheres (Fig. 4C) and soft agar colonies (Fig. 4D), as well as enhanced in vivo tumorigenicity (Fig. 4E), compared with their parental and control pLV SP- AA cells. Moreover, overexpression of HMGA2 with simultaneous knockdown of Sox2 using shRNA reduced sphere and colony formation ability and in vivo tumorigenicity (Fig. 4C-E). These data suggested that downregulation of Let-7a leads to upregulation of HMGA2 and Sox2, thus conferring AA cells the ability to retain cancer stem-like properties.

Let-7a/HMGA2 axis leads to the regulation of Sox2. (A) Quantitative RT-PCR analysis of HMGA2 and its associated downstream mediators, including GFAP, MAP2, nestin, Oligo2, and Sox2, in SP- AA cells transfected with shHMGA2 method (Table 2; National RNAi Core Platform, Academia Sinica, Taiwan). (B) The protein expression levels of HMGA2 and SOX2 in shHMGA2-transfected SP+ cells and HMGA2-transfected SP- cells were analyzed by Western blot. (C) SP+ AA cells transfected with pLV or pLV-HMGA2 and concomitant knockdown of Sox2 were subjected to sphere formation. #p < 0.05 versus treated groups (D) SP+ AA cells transfected with pLV or pLV-HMGA2 as well as concomitant knockdown of Sox2 were subjected to a soft agar assay for clonogenic ability. (E) NOD/SCID (BALB/c strain) mice were injected in the right flank with 10,000 cells as indicated, and the growth of the tumor xenografts was monitored by caliper measurements for up to 6 weeks.
Target Sequence and Plasmid Construction Primers

PU-PEI-Mediated Delivery of Let-7a in SP+ AA Cells Suppressed Cell Survival with the Use of Chemotherapeutic Drugs and IR Treatments
PU-PEI was demonstrated to present low cytotoxicity and high transfection efficiency and was therefore used as an ideal delivery vehicle (25,34). Our previous work demonstrated that anticancer efficacy was enhanced due to elevated nuclear delivery by preconjugation of NLS to siRNA-expressing dsDNA (35,54). To further achieve the ultimate goal of designing synthetic gene delivery vehicles to improve efficiency enough to be used in human therapy, we therefore used PU-PEI as a nonviral delivery system to transfer Let-7a conjugated with NLS into AA-SP+ CSC-like cells to elucidate the consequence of Let-7a overexpression in such lineage. In the preparation of expression vectors and the PU-PEI/Let-7a complex, both plasmids and PU-PEI complexes were dissolved in opti-MEM and mixed as described in Materials and Methods. Cocktails of PU-PEI and plasmid complexes were applied to 70% confluence-attached culture cells for 24 h, and the culture medium was replaced with fresh culture medium (Fig. 5A, B). In the following experiments to investigate Let-7a-associated signaling, plasmid vectors encoding EGFP and Let-7a-NLS or shHMGA2-NLS were delivered by PU-PEI in SP+ AA cells to generate stable Let-7a-overexpressing or HMGA2-knockdown SP+ AA cells. After successful delivery of Let-7a and shHMGA2, these Let-7a-overexpressing (SP+/PU-PEI-Let7a) (Table 2 and HMGA2-knockdown (SP+/PU-PEI-shHMGA2) SP+ AA cells, along with PU-PEI-only-delivered control cells (SP+/PU-PEI), were assigned for the in vitro and in vivo analysis to assess the therapeutic effects of PU-PEI-delivery of Let-7a in AA-SP+ CSC-like cells.

PU-PEI-mediated delivery of microRNA-expressing Let-7a preconjugated with NLS peptides. (A) Preconjugation of microRNA sequences of Let-7a with NLS peptides. (B) Delivery of PU-PEI vehicle NLS-preconjugated Let-7a with eGFP plasmids and subsequent gene silencing.
PU-PEI-Mediated Delivery of Let-7a Reduced CSC-Like Properties
Compared to the parental and SP+/PU-PEI control AA cells, PU-PEI-mediated Let-7a-NLS delivery inhibited the number of soft agar colonies (Fig. 6A) in the three patient-derived SP+ AA cells. Overexpression of Let-7a-NLS via PU-PEI delivery had effects similar to knockdown of HMGA2 in SP+ AA cells: both of them decreased the protein levels of HMGA2, and SOX2 (Fig. 6B). PU-PEI-mediated Let-7a-NLS delivery inhibited the proliferation rate (Fig. 6C) and reduced the number of spheres (Fig. 6D). Moreover, the number of secondary and tertiary spheres was also reduced by Let-7a delivery (Fig. 6D). The size of the SP in SP+/PU-PEI-shHMGA2 and SP+/PU-PEI-Let7a cells was smaller than that of the parental and control cells (Fig. 7A). We used a gene expression microarray to analyze the genomic traits of the SP-, SP+, SP+/PU-PEI, and SP+/PU-PEI-Let-7a-NLS cell lines, along with ESCs. The results were entered into multidimensional scaling analysis and revealed that the gene expression patterns of the SP+ AA cells resembled the expression patterns of ESCs and mesenchymal stem cells (MSCs; Fig. 7B). PU-PEI-mediated Let-7a-NLS or shHMGA2-NLS delivery changed the gene expression pattern of the SP+ AA cells to one closer to that of the SP- AA cells. Furthermore, quantitative RT-PCR analysis confirmed that, in addition to HMGA2 and Sox2, the expressions of other stemness genes, including Nanog, c-Myc, and the oncogene Bmi-1, were substantially attenuated by PU-PEI-mediated Let-7a-NLS delivery (Fig. 7C).

PU-PEI-mediated delivery of Let-7a reduces the AA-CSC population. (A) AA-SP+ cells isolated from two patient-derived specimens (patient Nos. 1 and 2) were cultured in serum-free media. Using a GFP-conjugated PU-PEI-based system, Let-7a was delivered into the SP+ AA cells. Scale bar: 100 μM. (B) The protein expression levels of Sox2 and HMGA2 in PU-PEI-mediated Let-7a-transfected SP+ AA cells were analyzed by Western blot. Data shown here are the mean ± SD of six independent experiments. *p <0.05 versus parental. These PU-PEI-mediated Let-7a-transfected AA-CSCs were then assigned to the (C) proliferation assay and (D) sphere formation assay. Data shown here are the mean ± SD of six independent experiments. *p <0.05 versus parental. #p <0.05 versus treated groups.

PU-PEI-mediated Let-7a delivery represses the stem-like gene expression signature of AA-CSCs. (A) The effect of PU-PEI-mediated delivery of NLS-preconjugated dsDNA encoding left-7a and shRNA-HMGA2 on the percentage of AA-derived SP cells. (B) Multidimensional scaling analysis illustrates the average lineage transcriptome distances among ESCs, MSCs, NSCs, AA-SP+, AA-SP+/PU-PEI, AA-SP+/PU-PEI-Let-7a, AA-SP+/PU-PEI-shHMGA2, and AA-SP- cells. The PU-PEI-Let-7a and PU-PEI-shHMGA2 complexes shifted the transcriptome distance of the AA-SP+ cells from their parental cells toward the AA-SP- cells. (C) Quantitative RT-PCR analysis of the mRNA expression levels of the stemness genes and the oncogene Bmi-1 after receiving PU-PEI-mediated Let-7a delivery. Data shown here are the mean ± SD of six independent experiments. *p <0.05 versus parental.
Therapeutic Delivery of PU-PEl-Let-7a Complex Enhances Chemoradiosensitivity In Vivo
The observation of Let-7a- and Sox2/HMGA2-mediated regulation of the CSC gene expression pattern and its properties suggested their potential involvements in modulating the chemo- and radioresistance of AA-CSCs. Cell viability was measured to evaluate the sensitivity of the SP+ AA cells to radiation or chemotherapeutic drugs. We found that SP+/PU-PEI-Let-7a cells had a lower survival fraction at all given doses of cisplatin (Fig. 8A), paclitaxel (Fig. 8B), and radiation (Fig. 8C) than the SP+/PU-PEI control cells, indicating that PU-PEI-mediated Let-7a-NLS delivery enhanced chemoradiosensitivity in SP+ AA cells. These data indicated that PU-PEI-mediated Let-7a-NLS delivery is able to reduce the CSC property of SP+ AA cells and enhance the therapeutic effects of chemotherapy and radiotherapy in SP+ AA cells. Based on the in vitro findings, we further investigated the therapeutic potential of the PU-PEI-Let-7a-NLS complex against AAs in vivo. Nude mice were subcutaneously transplanted with 105 patient-derived AA-SP+ cells, and the size of the tumors was monitored every 5 days up to 30 days. Five days after xenotransplantation of tumor cells, PU-PEI or PU-PEI-Let-7a-NLS was delivered into the xenograft, which was then subjected to a course of treatment in the presence of IR (Fig. 8D). PU-PEI-mediated Let-7a delivery alone moderately restricted tumor growth. The xenograft was not responsive to IR during the 30-day experimental course, but Let-7a delivery combined with IR led to the maximal inhibition of tumor growth (Fig. 8D). Quantification of bioluminescence imaging at 6 weeks posttumor transplantation (1,000 patient-derived AA-SP+ cells, IC) confirmed that maximal tumor repression was observed with a combination of PU-PEI-mediated Let-7a-NLS delivery and chemoradiotherapy (Cisplatin + IR) (Fig. 8E). The combination of Let-7a delivery and radiotherapy almost eliminated tumorigenesis in all of the recipients (AA-CSCs from No. 1 and No. 2 patients; Fig. 8F), which supported the remarkable efficacy of Let-7a delivery to suppress tumorigenesis. Taken together, our findings demonstrated that PU-PEI-mediated Let-7a-NLS delivery is a useful approach to prevent AA tumorigenesis in vivo, and the combination of Let-7a delivery and chemoradiotherapy is potentially an effective strategy for the treatment of AA and AA-CSCs.
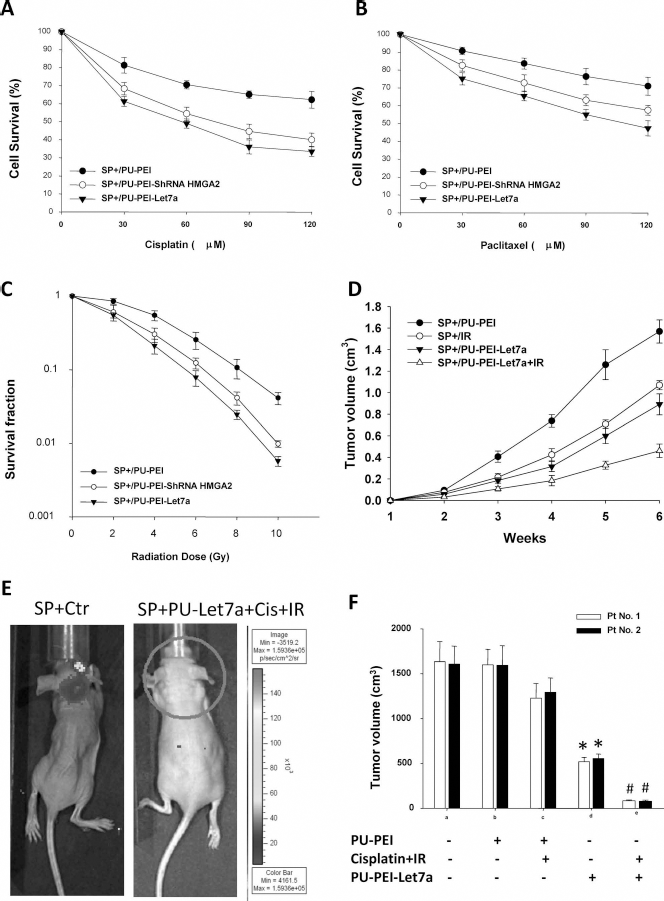
PU-PEI-mediated Let-7a delivery enhances the efficacy of irradiation and TMZ on AA tumorigenesis in orthotopic human AA-SP+-transplanted immunocompromised mice. The effect of PU-PEI-mediated Let-7a delivery on the size of the AA-SP+ cells. After exposure to different doses of (A) cisplatin and (B) pactitaxel, as indicated, the cell viability of the parental AA-SP+ cells, the AA-SP+ cells transfected with PU-PEI only, and the AA-SP+ cells transfected with PU-PEI-Let-7a were evaluated by MTT assay (Cell Biolabs, Inc., San Diego, CA, USA). (C) After exposure to 2-10 Gy of irradiation, the surviving cell fractions of the parental AA-SP+ cells, the AA-SP+ cells transfected with PU-PEI only, and the AA-SP+ cells transfected with PU-PEI-Let-7a were evaluated. (D) AA-SP+ (1 χ 105 cells) were SC implanted in the lower left flank of nude mice and allowed to develop tumors to around 200 mm3 in size (12 days). PU-PEI-modified scrambled sequence control (BioSettia, Inc.; PU-PEI/miR-Scr.) or Let-7a (PU-PEI/Let-7a) were injected in the left- or right-side tumor locus, respectively, on day 12. The tumor volume was measured by caliper, and the tumor growth curves are shown in the graph. (E) Immunosuppressive (nude) mice were IC transplanted with 103 patient-derived AA-SP+ cells. Five days after the xenotransplantation of the tumor cells, PU-PEI, or PU-PEI-Let-7a-NLS was IC delivered to the mice, which were then subjected to a course of treatment in the presence of cisplatin and IR. The size of the tumors was monitored by bioluminescence imaging every 7 days, up to 6 weeks. (F) The tumor volume was measured by caliper, and the tumor growth curves are shown in the graph at 6 weeks. #p <0.05 versus treated groups. Each group n = 6.
Discussion
Malignant glioma is a heterogeneous neoplasm driven by complex signaling pathways and is one of the most aggressive and challenging cancers to treat. The development of treatment for AA patients has been stagnant for decades; patients are still treated with surgery followed by radiation therapy (RT) only. Surgery aids in the reduction of tumor size and adjuvant RT used in the postoperative period have been demonstrated to extend the survival rates of the patients and postpone recurrence. It is important to develop strategies to overcome the resistance of AA cells to current therapies. Nonviral gene delivery, including liposome-, polymer-, and peptide-based gene delivery systems, has therefore been considered potentially safer than viral-mediated delivery (48). However, its efficiency is still in need of improvement (15,48). Recent nanotechnology-based approaches have demonstrated remarkable advances in terms of targeting therapy (49,57), the efficiency or stability of RNAi delivery (26,27), and the ability to overcome chemotherapeutic resistance (41). NLS peptides have been utilized to improve the entry of plasmid DNA into the nucleus (40,54). By conjugating NLS, transfection efficiency could be enhanced 10- to 1,000-fold compared to constructs without NLS (54). NLS peptides have been used to improve gene delivery (8,9,38) and have also been implicated in cancer therapy (28,50). However, an effective strategy using a nonviral delivery method to transport NLS-conjugated miRNAs to cancer cells is lacking. In this study, we showed that Let-7a was significantly down-expressed in isolated AA-derived SP+ cells, which represent a CSC-like model with tumor-initiating and radioresistant properties. Preconjugating Let-7a with a NLS significantly enhanced their tumor-suppressive effects, synergistically potentiated the effects of radiotherapy, and prolonged the survival of xenograft tumor-bearing mice (data not shown). The size of the AA-derived SP+ population and its CSC properties were also substantially blocked by transfection of NLS-conjugated Let-7a plasmid. Taken together, we demonstrated the critical role of NLS in assisting nonviral PU-PEI-mediated Let-7a delivery in AA-derived SP+ cells and in blocking CSC-like properties and chemoradioresistance of AA and AA-derived CSCs.
The CSC hypothesis proposes that tumors contain a small subset of cancer cells that constitute a reservoir of self-sustaining cells with the exclusive ability to self-renew and maintain the tumor. Hirschmann-Jax et al. (23) showed that “SP” CSC-like cells isolated from patients with neuroblastomas express high levels of ABCG2 and ABCG3 transporter genes and have a greater capacity to expel cytotoxic drugs. Herein, we found that AA-derived SP+ cells show stem-like properties (Fig. 1A-C) and that these AA-derived SP+ cells were significantly more resistant to conventional treatment methods than SP- cells (Fig. 1D). Let-7a-overexpressed SP+ AA cells had reduced sphere formation, soft agar colonies, migration, and invasion abilities. Furthermore, bioinformatics and microarray analyses were used to distinguish miRNAs and their associated candidates of target genes (Fig. 2A) that may play important roles in putative AA-SP+-derived CSCs. Based on our data, expression of Let-7a was decreased in SP+ CSC-like tumor cells compared to SP- non-CSC AA cells (Fig. 2A). We also consistently demonstrated a strong negative correlation between Let-7a and HMGA2 or Sox2 in AA cells. HMGA2 is a chromatin-binding protein that contains three AT-hook domains that enable its binding to the minor groove of DNA (6). HMGA2 is highly expressed in ES cells but is generally low or lacking in normal somatic cells (10). HMGA2 protein is overexpressed in many types of cancer, such as lung cancer (37), breast cancer (4), oral squamous cell carcinoma (39), and colorectal cancer (52). In the present study, luciferase reporter assay showed that Let-7a directly targets the 3′-UTRs of HMGA2 in AA-SP+. Moreover, overexpression of Let-7a negatively regulated the expression of HMGA2 in AA-SP+ cells, leading to suppressed tumorigenicity in vitro and in vivo. SOX2 is a member of the SOX family of transcription factors responsible for coordinating disparate functions such as maintaining stem cell properties and differentiation restriction. In this study, our data supported the role of Sox2 as a possible downstream effector of HMGA2. In the SP+ AA cells, the Let-7a level was low, but HMGA2 and Sox2 expression was high; an inverse pattern of Let-7a, HMGA2, and Sox2 gene expression was observed in SP- AA cells.
The miRNA could be a drug that induces apoptosis and/or cell cycle arrest in cancer cells. In this study, we used cationic lipids to condense Let-7a (PU-PEI-Let-7a-NLS) in an attempt to enhance cellular uptake and pharmacological effectiveness in vitro and in vivo. PU-PEI-Let-7a-NLS treatment showed significantly inhibited sphere formation, tumorigenicity, and migration and dramatically enhanced chemoradiosensitivity in SP+ AA cells. Of more importance, the orthotropic injection of the PU-PEI-Let-7a-NLS into immunocompromised mice reduced the size of the AA-SP+-transplanted tumors and inhibited the CSC-like properties. We suggested that the cellular levels of Let-7a act as a switch that modulates the CSC properties in AA. The promising utility of PU-PEI-mediated Let-7a-NLS delivery indicated that it might improve anti-AA treatment, especially in brain tumors that are resistant to chemoradiotherapy. These results demonstrated the potential for the development of miRNA-based strategies to inhibit the malignancy of CSC and improve conventional therapeutics.
Footnotes
Acknowledgment
This study was funded by the National Science Council (NSC 98-2314-B-075A-004-MY3 and NSC 102-2314-B-016 -031 -MY3). The authors declare no conflicts of interest.
