Abstract

Neural cells derived from induced pluripotent stem cells (iPSCs) have the potential for autologous cell therapies in treating patients with severe neurological disorders or injury. However, further study of efficacy and safety are needed in large animal preclinical models that have similar neural anatomy and physiology to humans such as the pig. The pig model for pluripotent stem cell therapy has been made possible for the first time with the development of pig iPSCs (piPSCs) capable of in vitro and in vivo differentiation into tissues of all three germ layers. Still, the question remains if piPSCs are capable of undergoing robust neural differentiation using a system similar to those being used with human iPSCs. In this study, we generated a new line of piPSCs from fibroblast cells that expressed pluripotency markers and were capable of embryoid body differentiation into all three germ layers. piPSCs demonstrated robust neural differentiation forming βIII-TUB/MAP2+ neurons, GFAP+ astrocytes, and O4+ oligodendrocytes and demonstrated strong upregulation of neural cell genes representative of all three major neural lineages of the central nervous system. In the presence of motor neuron signaling factors, piPSC-derived neurons showed expression of transcription factors associated with motor neuron differentiation (HB9 and ISLET1). Our findings demonstrate that SSEA4 expression is required for piPSCs to differentiate into neurons, astrocytes, and oligodendrocytes and furthermore develop specific neuronal subtypes. This indicates that the pigs can fill the need for a powerful model to study autologous neural iPSC therapies in a system similar to humans.
Introduction
Induced pluripotent stem cell (iPSC) therapies have the potential to revolutionize modern day medicine, providing regenerative cell therapies for neurodegenerative diseases such as Parkinson's and neural injuries including stroke and spinal cord damage (6, 8, 19, 23, 34). Studies of iPSCs and derived neural cells in rodent systems have shown significant promise and have demonstrated that these cells can differentiate and successfully integrate into neural disease and injury models leading to functional recovery (8, 16, 17). However, further preclinical studies of efficacy and safety are likely to be needed in large animal models, such as the pig, that have similar neural anatomy and physiology to humans before neural iPSC therapies become a viable treatment option for human patients (18, 27, 37, 44, 45). The recent development of bona fide pluripotent pig induced pluripotent stem cells (piPSCs) capable of achieving the highest pluripotency standards, teratomas, and germline competent chimeras has made the pig a strong candidate for neural iPSC research (12, 13, 49–51). However, robust directed in vitro neural lineage differentiation of piPSCs has not been demonstrated and is a critical first step toward utilizing piPSCs as a large animal preclinical model for the development of neural iPSC therapies.
piPSCs have recently been derived by several groups and have demonstrated significant developmental plasticity forming embryoid bodies (EB) in vitro and producing teratomas and contributing to germline chimeras in vivo (12, 13, 49–51). Analyses of EBs and teratomas have shown piPSCs are capable of undergoing spontaneous neural differentiation with gene and protein expression of neural markers for neurons and glia (12, 13, 51). Chimeric pigs generated from piPSCs also showed contribution of piPSCs to brain tissue, indicating that these cells are capable of forming cells of the central nervous system (CNS) in an in vivo setting (49). These lines of evidence suggest that piPSCs have the capacity to undergo neural differentiation both in vitro and in vivo. piPSCs capable of directed in vitro neural differentiation similar to human and mouse iPSCs would open the door for in vitro and in vivo comparative studies between species to determine optimum differentiation conditions for forming functional neural therapeutics. Studies using piPSCs are of considerable biomedical significance, as pigs have gyrencephalic brains with similar gray and white matter composition and size and spinal cords with length more comparable to humans than rodents (27, 38, 45). These are all important aspects to consider when developing neural iPSC therapies. In vivo, piPSCs also enable studies of autologous pig model transplants that overcome the potential confounding variables of species-specific differences found in large animal xenotransplantation models (e.g., human neural cells in pig models). piPSCs show significant promise as a comparative tool to study neural iPSC therapies, yet it must be established that piPSCs can undergo in vitro directed neural differentiation using conditions established for human and mouse iPSCs to be a useful comparative model.
In vitro human and mouse iPSCs are capable of robust neural differentiation into neurons, astrocytes, and oligodendrocytes and specialized neural subtypes such as motor neurons (4, 24, 25, 39, 42, 46, 55). Despite significant variability in iPSCs to neural culture systems, neural differentiation is highly ordered and can be typically induced by either EBs or adherent differentiation giving rise to early neural progenitors (NPs). NPs can then be driven to more restricted neuronal, astrocytic, and oligodendritic cell types by the addition of neural growth supplements such as B27 and N2 (24, 39, 42, 46, 55). Ultimately, specified neurons such as motor neurons can be induced by adding a cocktail of morphogens and signaling factors including retinoic acid (RA), sonic hedgehog (SHH), and brain-derived growth factor (BDNF) (4, 35). Utilizing similar differentiation approaches, we believe that piPSCs will demonstrate similar neural differentiation potential.
In this report, we generated piPSCs from fibroblast cells that display morphology and immunoreactivity indicative of a pluripotent state that were capable of embryoid body differentiation into all three germ layers. piPSCs demonstrated differential expression of the pluripotency marker stage-specific embryonic antigen 4 (SSEA4) with some being SSEA4+ and other being SSEA4-. Isolation and examination of SSEA4+ and SSEA4- piPSCs' neural differentiation potential demonstrated that SSEA4+ piPSCs showed a higher propensity to undergo processive neural development including neurons, astrocytes, and oligodendrocytes. piPSC-derived neurons were then tested for their ability to form motor neuron cells. Understanding the in vitro neural differentiation potential of piPSCs represents a critical first step in understanding the ability of these cells and the pig as a whole to be utilized as a neural iPSC therapy developmental model.
Materials and Methods
Cell Culture and Lentiviral Transduction
piPSCs were generated as previously described (49). Briefly, pig dermal fibroblasts were derived from a skin biopsy of a 2-year-old Hampshire pig (Kewanee Farm, Dudley, GA, USA). Fibroblasts were expanded and maintained in fibroblast growth medium consisting of Dulbecco's modified Eagle's medium (DMEM)/high glucose (Gibco, Grand Island, NY, USA), 10% fetal bovine serum (HyClone, Logan, UT, USA), 2 mM l-glutamine, 50 units/mL penicillin and 50 μg/ml streptomycin (Gibco). Cells were maintained in 5% CO2 and at 37°C. Fibroblasts were plated at a density of 43,000 cells/cm2 and after 24 h cells were transduced at an multiplicity of infection (MOI) of 20 using viPS lentiviral vectors (Thermo Scientific, West Palm Beach, FL, USA) expressing human POU domain class 5, transcription factor 1 [POU5F1 also known as octamer binding transcription factor 4 (OCT4)], NANOG, sex determining region Y box 2 (SOX2), Cell Lineage 28 (LIN28), Krüppel-like factor 4 (KLF4) and v-myc myelocytomatosis viral oncogene homolog (C-MYC) open reading frames under the control of the human elongation factor-1α (EF-1α) promoter in the presence of 1.2% GeneJammer (a polyamine-based transfection reagent; Stratagene, La Jolla, CA, USA). After 24 h the transduced cells were passaged onto mitomycin C (Sigma-Aldrich, St. Louis, MO, USA) mitotically inactivated mouse embryonic fibroblast feeders using 0.05% trypsin (Gibco) and cultured in stem cell medium which is composed of DMEM/F12 (Gibco) supplemented with 20% knockout serum replacement (KSR; Gibco), 2 mM l-glutamine (Gibco), 0.1 mM non-essential amino acids (Gibco), 50 units/ml penicillin and 50 μg/ml streptomycin (Gibco), 0.1 mM β-mercaptoethanol (Sigma-Aldrich), and 10 ng/ml fibroblast growth factor-2 (R&D Systems, Minneapolis, MN, USA). piPSC colonies were manually selected from transduced cultures every 3 days from days 15 to 25 and maintained by manual passage every 4–5 days. piPSCs were then transferred to feeder-free conditions utilizing 0.05% trypsin and plating cells onto growth factor-reduced Matrigel (diluted 1:100 in DMEM/F12; BD Biosciences, San Jose, CA, USA)-coated plates in the mTeSR1 (Stemcell Technologies, Vancouver, BC, Canada) medium. Cells were passaged using 1 mg/ml of dispase (Stemcell Technologies) every 3–4 days upon reaching 80% confluency. Karyotype analysis was performed after 22 passages under feeder-free conditions by a standard high-resolution G-banding method by Cell Line Genetics (Madison, WI, USA).
The human embryonic stem cell line H9 (WiCell, Madison, WI, USA) was grown in mTeSR1 medium (Stemcell Technologies) on Matrigel-coated dishes as a positive control.
Fluorescence-Activated Cell Sorting
piPSCs were harvested enzymatically as a single cell suspension using trypsin, washed with phosphate-buffered saline (PBS; ThermoFisher Scientific), and blocked in 4% horse serum (Gibco). Cells were incubated with primary antibodies directed against SSEA4 (Developmental Studies Hybridoma Bank, Iowa City, IA, USA; 1:200). Primary antibodies were detected using fluorescently conjugated secondary antibody Alexa Fluor 405 (Invitrogen, Carlsbad, CA, USA; 1:500). Cells were sorted with the FACSCalibur (Becton Dickinson, Franklin Lakes, NJ, USA) flow cytometer. Cells were collected in mTeSR1 medium and centrifuged and resuspended in fresh mTeSR1 medium and plated onto Matrigel-coated dishes.
Neural Differentiation of piPSCs
For neural induction, sorted SSEA4+ and SSEA4– piPSCs were seeded on Matrigel-coated slides in mTeSR1 medium and when the cells were 80% confluent, the cells were cultured for 10 and 20 days in neuronal differentiation medium, which consisted of basal neural medium AB2 (Neuromics, Edina, MN, USA), 1 × ANS (Neural medium Supplement; Neuromics), 2 mM l-glutamine, 50 units/ml penicillin, and 50 μg/ml streptomycin, and supplemented with 5% fetal bovine serum (Gibco). All experiments were replicated a minimum of four times.
To induce motor neuron differentiation, an established protocol utilizing human embryonic stem cells was employed (29, 48). SSEA4+ piPSCs were treated with 2 μM RA (Sigma) and 500 ng/ml SHH (R&D Systems) in neuronal differentiation medium for 7 days. At 7 days, the medium was supplemented with BDNF, glial cell-derived neurotrophic factor (GDNF) and insulin-like growth factor 1 (IGF1) (all at 10 ng/ml, PeproTech, Inc., Rocky Hill, NJ, USA) and maintained with BDNF, GDNF, and IGF1 for an additional 14 days with medium changes every other day.
Alkaline Phosphatase Staining and Immunocytochemistry
Alkaline phosphatase (AP) staining was carried out with VECTOR Red Alkaline Phosphatase Substrate Kit (Vector Laboratories, Burlingame, CA, USA) according to the manufacturer's instructions.
For immunostaining, cells were cultured on Matrigel-coated glass slides and then fixed in 4% paraformaldehyde (PFA; Electron Microscopy Sciences, Hatfield, PA, USA). For intracellular staining, cells were permeabilized with Triton X-100 (Sigma-Aldrich), 1% polyvinylpyrrolidone (PVP; Sigma-Aldrich) in a PBS blocking solution containing 4% normal goat serum (Gibco). For extracellular staining, cells were blocked in a PBS containing 4% normal goat serum. Primary antibodies used were POU5F1 (R&D Systems, 1:200), NANOG (Millipore, Billerica, MA, USA; 1:200), SOX1 (R&D Systems, 1:20), SOX2 (R&D Systems, 1:200), SSEA4 (Developmental Studies Hybridoma Bank, 1:200), TRA 1-60 (Chemicon, Temecula, CA, USA; 1:200), TRA 1-81 (Chemicon, 1:200), human neuronal protein C/D (HU C/D; Invitrogen, 1:200), βIII-TUBULIN (βIII-TUB, Neuromics, 1:200), microtubule-associated protein 2 (MAP2; Millipore, 1:200), glial fibrillary acidic protein (GFAP; Chemicon, 1:200), oligodendrocyte marker 4 (O4; Chemicon, 1:100), motor neuron and pancreas homeobox protein 1 (homeobox 9 or HB9; Santa Cruz Biotechnologies, Dallas, TX, USA; 1:100) and insulin gene enhancer protein 1 (ISLET1, 40.2D6; Developmental Studies Hybridoma Bank, 1:50). Primary antibodies were detected using a fluorescently conjugated secondary antibody, Alexa Fluor 405 (Molecular Probes, Eugene, OR, USA; 1:500) and 594 (Molecular Probes, 1:500). Cells were washed three times with PBS and then treated with Prolong Gold and DAPI (Invitrogen). Cells plated on slides were imaged on IX81 with Disc-Spinning Unit (Olympus, Inc., Center Valley, PA, USA) using SlideBook Software (Intelligent Imaging Innovations, Inc., Denver, CO, USA). Cells expressing or not expressing proteins of interest were quantified using Image-Pro Plus Version 5.1 Software (Media Cybernetics, Rockville, MD, USA). Each experiment was replicated a minimum of three times.
Embryoid Body Formation and Differentiation
Embryoid bodies (EBs) were formed by plating 2.0 × 106 cells in mTeSR1 medium and Y-27632 ROCK inhibitor (10 μM, Calbiochem, Torrey Pines, CA, USA) into a single well of an AggreWell plate (Stemcell Technologies) and centrifuged at 100 × g for 3 min per manufacturer's instruction. After 24 h, EBs were maintained in Petri dish (BD Falcon) suspension culture in 20% KSR medium without fibroblast growth factor-2 and allowed to differentiate for 10 days. Each experiment was replicated a minimum of four times.
RNA Isolation and RT-PCR
Total RNA was extracted using the Qiashredder and RNeasy mini kits (Qiagen, Germantown, MD, USA) according to the manufacturer's protocol. The RNA quality and quantity was determined using the NanoDrop 8000 (Thermo Scientific). Total RNA (500 ng) was reverse-transcribed using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) following manufacturer's instructions.
Primer Sequences

See text for definitions of gene abbreviations.
For quantitative RT-PCR, each reaction contained 10 μl Taqman Universal Master Mix II (Applied Biosystems), 1 μl Taqman Assay (20x), and 20 ng cDNA as a template in a final volume of 20 μl. Assays were performed using 384-well clear optical reaction plates on the ABI PRISM 7900 Sequence Detection System (Applied Biosystems). All of the oligonucleotide primer and fluorogenic probe sets [cluster of differentiation 4 (CD4), γ-aminobutyric acid type B receptor subunit 1 (GABBR1), neuronal differentiation 1 (NEURO D1), GFAP, secreted protein, acidic, cysteine-rich (osteonectin; SPARC), and glutamate-ammonia ligase (GLUL), myelin oligodendrocyte glycoprotein (MOG), proteolipid protein 1 (PLP1), and transferrin (TF)] were commercially designed by Applied Biosystems, Inc., for Taqman™ real-time PCR. The cycle conditions were 95°C for 10 min, followed by 40 cycles of 95°C for 15 s, and 60°C for 1 min. mRNA levels were normalized by using glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as an endogenous control. Relative gene expression was shown as fold changes (ratio) in gene expression using the comparative Ct method. For statistical analysis, the relative quantification (RQ) values were compared between groups for each gene. These experiments were performed with three technical replicates for each sample and three biological replicates.
Statistical Analysis
Immunocytochemistry and quantitative RT-PCR data were analyzed using a one-way ANOVA (GLM procedure, Statistica, version 7.0; StatSoft, Tulsa, OK, USA) followed by Tukey's HSD post hoc test to determine significant differences between groups. Statistically significant differences are defined at the 95% confidence index (p < 0.05). Data shown are means±standard error of the mean.
Results
piPSCs Generated by Viral Transduction Express Pluripotency Markers
iPSCs were generated from pig dermal fibroblasts using lentiviral vectors encoding the six reprogramming factors POU5F1, SOX2, NANOG, LIN28, KLF4, and c-MYC with expression driven by the EF-1α promoter. After 24 h, transduced porcine fibroblasts were plated onto feeder cells in stem cell medium (Fig. 1). After 7 days, putative piPSCs began to emerge. piPSC colonies possessed pluripotent cell characteristics of well-defined borders and cells displayed a high nuclear to cytoplasmic ratio with prominent nucleoli (Fig. 1A). piPSCs were directly passaged into feeder free conditions on Matrigel in mTeSR1 medium without any subsequent passages on feeders. To determine if putative piPSCs demonstrated immunoreactivity consistent with pluripotent stem cells, cells were initially stained for alkaline phosphatase (AP) and were found to be positive (Fig. 1B). Immunostaining of piPSCs for additional pluripotency markers expression revealed that piPSCs were positive for the intracellular stem cell markers POU5F1 (Fig. 1C, E, F, H), SOX2 (Fig. 1D, E), and NANOG (Fig. 1G, H) and the extracellular stem cell surface markers SSEA4, TRA 1-60, and TRA 1-81 (Fig. 1I–K).
piPSCs display morphological characteristic and immunoreactivity indicative of a pluripotent state. Pig fibroblast cells transduced with POU domain class 5, transcription factor 1 (POU5F1 also known as octamer binding transcription factor 4 [OCT4]), NANOG, sex-determining region Y box 2 (SOX2), cell lineage 28 (LIN28), Krüppel-like factor 4 (KLF4), and v-myc myelocytomatosis viral oncogene homolog (C-MYC) reprogramming genes were plated on feeder cells in 20% knockout serum replacement (KSR) stem cell medium and displayed morphology indicative of an induced pluripotent stem cell (iPSC) state including colonial growth, cells with well-defined borders and high nuclear-to-cytoplasmic ratio (A). piPSCs expressed pluripotency markers alkaline phosphatase (AP) (B), POU5F1 (C, F), SOX2 (D), POU5F1/SOX2 Merge (E), Nanog (G), POU5F1/NANOG Merge (H), SSEA4 (I), TRA1-60 (J), and TRA1-81 (K). Cell localization was indicated by DAPI (blue) nuclear staining. Scale bars: 10 μm.
To determine if the human exogenous (hExo.) reprogramming genes had successfully integrated into the pig genome and activated the expression of pig endogenous (pEndo.) gene homologs, RT-PCR was performed with pig and human specific primers (see Table 1 for primer sequences). RT-PCR analysis showed that piPSCs expressed the hExo.POU5F1, SOX2, NANOG, LIN28, C-MYC, and KLF4 as did the human embryonic stem cell (hESC)-positive control (Fig. 2A). The pig fibroblast (pF) parent population from which the piPSCs were originally derived was negative for all human genes (Fig. 2A). Examination of piPSCs for gene expression of endogenous pig homologs demonstrated that all six genes (pEndo.POU5F1, SOX2, NANOG, LIN28, C-MYC, and KLF4) were expressed in piPSCs but were negative in pF and hESC populations (Fig. 2B). piPSCs also showed positive gene expression for the pluripotency markers epithelial cadherin 1 (CDH1), dosage-sensitive sex reversal, adrenal hypoplasia critical region, on chromosome X, gene 1(DAX1), growth factor receptor-bound protein 7 (GRBP7), interferon-induced transmembrane protein 1 (IFITM1), teratocarcinoma-derived growth factor 1 (TDGF1), telomerase reverse transcriptase (TERT), and RNA exonuclease 1 homolog (REX1), while pF cells were negative (Fig. 2C). After 22 passages, cytogenetic analysis showed that piPSCs had a normal karyotype (Fig. 2D). These data indicate the piPSCs are karyotypically stable pluripotent cell line in both gene and protein expression.
Overexpression of exogenous human reprogramming pluripotent genes leads to expression of endogenous pig homologs in piPSCs. RT-PCR analysis of piPSCs demonstrated that these cells expressed exogenous human (hExo.) reprogramming genes POU5F1, SOX2, NANOG, LIN28, C-MYC, and KLF4 similar to human embryonic stem cells (hESCs), while pig fibroblast (pF) parent cells were negative for all genes (A). piPSCs expressed endogenous pig (pEndo.) homologs POU5F1, SOX2, NANOG, LIN28, C-MYC, and KLF4, while pF and hESC populations were negative (B). piPSCs also expressed the pluripotency genes epithelial cadherin (CDH1), dosage-sensitive sex reversal, adrenal hypoplasia critical region, on chromosome X, gene 1 (DAX1), growth factor receptor-bound protein 7 (GRBP7), interferon-induced transmembrane protein 1 (IFITM1), teratocarcinoma-derived growth factor 1 (TDGF1), telomerase reverse transcriptase (TERT), and RNA exonuclease 1 homolog (REX1), while pF cells were negative (C). Passage 22 piPSCs showed normal karyotype (D). GAPDH, glyceraldehyde 3-phosphate dehydrogenase.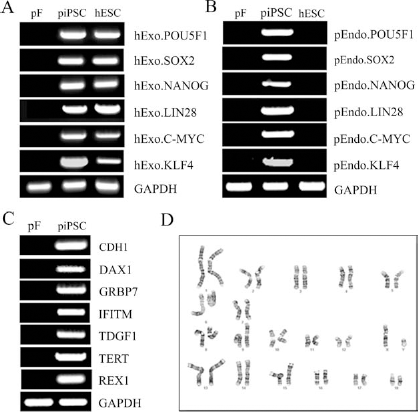
piPSCs Form All Three Germ Layers During EB Differentiation
To determine if piPSCs were capable of producing cell types representing all three germ layers, piPSCs underwent embryoid body (EB) differentiation for 10 days. EBs were formed in AggreWell plates and after 24 h were cultured in suspension in EB differentiation medium. piPSC-derived EBs were round and highly compact (Fig. 3A). RNA was isolated from day 10 EBs, and gene expression analysis showed strong upregulation of ectoderm [NESTIN, neuronal differentiation 1 (NEUROD1), cellular retinoic acid binding protein 2 (CRABP2)], endoderm [α-fetoprotein (AFP), hepatocyte nuclear factor (HNF), KERATIN18], and mesoderm [mutS homolog homeobox 1 (MSX1), actin, alpha, cardiac muscle 1 (ACTC1), cluster of differentiation 34 (CD34)] genes, whereas piPSCs were negative for these markers (Fig. 3B). These results indicated that piPSCs could differentiate into various cell types from all three germ layers.
EBs derived from piPSC express markers of all three germ layers. piPSCs underwent 10 days of embryoid body (EB) differentiation (A) with analysis of gene expression demonstrating the presence of ectoderm, mesoderm, and endoderm genes within the cells (B). NEUROD1, neuronal differentiation 1; CRABP2, cellular retinoic acid binding protein 2; MSX1, mutS homolog homeobox 1; ACTC1, actin, alpha, cardiac muscle 1; CD34, cluster of differentiation 34; AFP, α-fetoprotein; HNF, hepatocyte nuclear factor.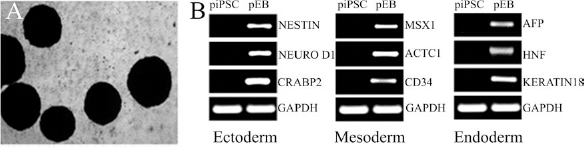
SSEA4-Positive piPSCs Differentiate Into Neurons, Astrocytes, and Oligodendrocytes
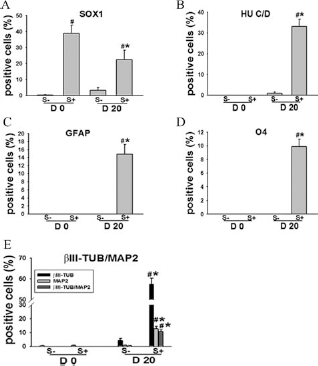
A subpopulation of piPSCs were positive for the stem cell marker SSEA4, while a significant population was SSEA4 negative. This suggests a heterogeneous population of cells that may have divergent differentiation potentials with cells positive for SSEA4 likely to have a higher level of plasticity and be more capable of neural differentiation. To directly compare the ability of SSEA4+ and SSEA4-cells to form cells of the neural lineage, piPSCs underwent fluorescent-activated cell sorting (FACS) and were stained for neural markers at day 0 and after 20 days of adherent neural differentiation (Figs. 4 and 5). At day 0 a subpopulation of the SSEA4+ cells showed spontaneous differentiation into NP-like cells with 38.8% of cells being SOX1+ (Figs. 4A, B and 5A), while < 0.5% of SSEA4- cells were SOX1+ (Fig. 5A). Both SSEA4+ and SSEA4- populations were negative for more mature neural markers at day 0 (Fig. 5). piPSCs were then cultured in neural differentiation medium for 20 days. Immunostaining results showed that the differentiated SSEA4+ population was positive for the post mitotic neural marker HU C/D (33.2%) (Figs. 4C, D and 5B), the mature neural markers βIII-TUB and MAP2 57.2%, 12.6%, and 10.5% of cells were positive for βIII-TUB, MAP2, and both, respectively) (Figs. 4E–G and 5E), whereas SSEA4- cells were predominantly negative for the neural proteins after 20 days of differentiation. βIII-TUB/MAP2+ neuron-like cells possessed significant neurite outgrowths. Furthermore, differentiated SSEA4+ cells produced both GFAP+ astrocytes (14.8%) (Figs. 4H, I and 5C) that displayed stellate morphologies and O4+ oligodendrocytes (9.9%) (Figs. 4J, K and 5D) with a typical multipolar oligodendroglial morphology, while no obvious differentiation towards the astrocytic and oligodendritic lineages were observed in differentiated SSEA4-. These immunostaining results indicated that SSEA4+ piPSCs are capable of differentiating into neurons, astrocytes, and oligodendrocytes with neuronal marker-positive cells being the most abundant.
SSEA4+ piPSCs differentiate into early and later stage neuronal cell types. Immunofluorescence staining of day 20 neural differentiation cultures showed that stage-specific embryonic antigen 4 (SSEA4) cells were positive for the SOX1 neural progenitor marker (A, B Merge), postmitotic neuron marker human neuronal protein C/D (HU C/D) (C, D Merge), and βIII-TUBULIN/microtubule-associated protein 2 (βIII-TUB/MAP2) neuronal proteins (E, F, G Merge). In addition, cells in differentiation cultures were positive for the astrocyte marker glial fibrillary acidic marker (GFAP) (H, I Merge) and oligodendrocyte specific marker O4 (J, K Merge). Scale bars: 10 μm. SSEA4+ piPSCs show robust differentiation of into all three neural lineages. piPSCs were fluorescence activated cell sorted (FACS) into SSEA4+ (S+) and SSEA4- (S-) populations. At day 0 (D 0), expanded SSEA4+ cultures showed spontaneous differentiation with 38.8% of cells being SOX1+ (A), while all other neuronal markers were negative (B–E). Day 20 (D 20) neural differentiation cultures from SSEA4+ piPSCs resulted in βIII-TUB (57.2%) (E), HU C/D (33.2%) (B), and MAP2 (12.6%) (E) positive neuron populations, while also forming GFAP+ (14.8%) (C) astrocyte and O4+ (9.9%) (D) oligodendrocyte populations. SSEA4- piPSCs were negative for neural markers at day 0 and <5% cells were positive for neural markers in day 20 differentiation cultures (A–E). 
POU5F1 is a key transcription factor commonly used for the reprogramming somatic cells into iPSCs. However, POU5F1 is a known oncogene that is also highly expressed in various types of cancer and has been previously demonstrated to be reactivated in mice derived from iPSCs leading to high morbidity (21, 40, 41, 43). Therefore, it is critical to determine if POU5F1 expression is downregulated upon differentiation of piPSCs into neural cell types (Fig. 6). SSEA4+ piPSCs were stained for βIII-TUB and POU5F1 at day 0 and after 20 days of differentiation. At day 0, SSEA4+ piPSCs were highly positive for the POU5F1 (95.2%) (Fig. 6D), while almost all cells were negative for βIII-TUB. However, day 20 differentiated cultures were predominantly positive for the neural marker βIII-TUB (57.8%) (Fig. 6A, C, D) and fewer POU5F1+ cells were present (10.5%) (Fig. 6B–D). Interestingly, differentiated βIII-TUB+ cells were mostly negative for POU5F1 with only 5.6% being βIII-TUB/POU5F1+. These results suggest that differentiated βIII-TUB+ cells may have reduced potential for tumorigenicity; however, additional studies are needed to further determine the tumorigenicity of these cells.
Differentiated βIII-TUB+ cells are predominantly negative for POU5F1. Day 20 SSEA4+ piPSC neural differentiation cultures were coimmunostained for βIII-TUB (A) and POU5F1 (B, C Merge). Quantification of day 0 (D 0) and day 20 (D 20) immunostaining results demonstrated that POU5F1+ cells showed a significant (p < 0.05) decrease after differentiation, while βIII-TUB+ cells showed a significant (p < 0.05) increase (D). In day 20 cultures, 5.6% of cells were positive for both βIII-TUB and POU5F1. Scale bars: 10 μm. *Statistically significant difference p < 0.05 when comparing D 0 and D 20 for each protein.
To gain further insight into the ability of piPSCs to differentiate into neuronal cell types, both SSEA4+ and SSEA4- piPSCs were examined for neuronal gene expression at days 0, 10, and 20. Quantitative RT-PCR revealed that expression levels of the neuronal markers CD4, γ-aminobutyric acid type B receptor subunit 1 (GABBR1) and NEURO D1 in differentiated SSEA4+ piPSCs were significantly (p < 0.05) increased at day 20 relative to day 0 and relative to differentiated SSEA4- piPSCs (Fig. 7A–C). SSEA4- cultures only showed a significant increase in GABBR1 at days 10 and 20. Astrocytic differentiation was examined by analyzing the gene expression levels of GFAP, secreted protein, acidic, cysteine-rich (osteonectin; SPARC), and glutamate–ammonia ligase (GLUL) genes. Significant (p < 0.05) upregulation of astrocyte genes GFAP and SPARC were observed in SSEA4+ differentiation cultures at day 20 relative to day 0 and relative to SSEA4- cultures at day 20 (Fig. 7D–F). Differentiated SSEA4- piPSCs showed a significant increase in only the SPARC astrocyte markers at day 10. Furthermore, the gene expression levels of oligodendrocyte-specific genes myelin oligodendrocyte glycoprotein (MOG), proteolipid protein 1 (PLP1), and transferrin (TF) were significantly (p < 0.05) higher in differentiated SSEA4+ piPSCs at day 20 compared to day 0 populations and differentiated SSEA4- piPSCs. No significant upregulation of MOG, PLP1, and TF expression levels were found in differentiated SSEA4- populations at day 10 or day 20 (Fig. 7G–I). These RT-PCR results further support our previous immunostaining findings that SSEA4+ piPSCs are capable of robust differentiation into cell types of all three neural lineages of the central nervous system.
Neural differentiated SSEA4+ piPSC show high levels of neuron, astrocyte and oligodendrocyte gene expression. SSEA4+ (S+) and SSEA4- (S-) piPSCs underwent neural differentiation for 0, 10, and 20 days and were examined for neuron, astorycte, and oligodendrocyte markers. SSEA4+ piPSCs demonstrated significant (p < 0.05) upregulation at day 20 (D20) for neuron (CD4, GABBR1 and NEURO D1), astrocyte (GFAP, SPARC and GLUL), and oligodendrocyte (MOG, PLP1 and TF) relative to day 0 (D 0). However, SSEA4- day 20 neural differentiation cultures only demonstrated significant (p < 0.05) upregulation of the neural marker GABBR1 relative to day 0. Day 20 SSEA4+ neural differentiation cultures also showed significantly higher gene expression than SSEA4- neural differentiation cultures for all genes examined. #Statistically significant difference with p < 0.05 comparing S- and S+ at each time point; *statistically significant difference with p < 0.05 when comparing D 10 and D 20 to the D 0 time point within each SSEA4 population. GABBR1, g-aminobutyric acid type B receptor subunit 1; SPARC, secreted protein, acidic, cysteine-rich (osteonectin); GLUL, glutamate-ammonia ligase; MOG, myelin oligodendrocyte glycoprotein, PLP1, proteolipid protein 1; TG, transferrin.
SSEA4-Positive piPSC Differentiated Into Motor Neuron
Treating diseases such as amyotrophic lateral sclerosis (ALS), spinal muscular atrophy (SMA), and spinal cord injury that involve loss or severe damage to motor neurons would likely benefit from the establishment of a large animal model with the axons of motor neurons extending for more than a meter in length in humans (52). To examine the capacity of piPSCs to generate motor neurons using an established human embryonic stem cell protocol (29, 48), SSEA4
+
piPSCs were differentiated on a Matrigel substrate in the presence of RA and SHH for 7 days and then with BDNF, GDNF, and IGF1 for an additional 14 days (Fig. 8). Immunostaining analysis indicated that the majority of differentiated cells were positive for βIII-TUB (85.3%) (Fig. 8B, C, F). In addition, >10% of cells were positive for the motor neuron marker ISLET1 (Fig. 8D–F) and >6% were positive for HB9 (Fig. 8A, F), both transcription factors are either enriched or specific to motor neurons respectively (2, 20). Additional staining showed that ISLET1 and HB9 staining typically colocalized in all samples (Fig. 8G–I). Control cells differentiated for 21 days in basal differentiation media without additional growth factors were consistently found to be negative for HB9 and ISLET1 motor neuron markers. These results indicated that piPSCs are capable of in vitro-directed differentiation into specialized neural subtypes such as motor neurons.
Motor neuron differentiation of SSEA4+ piPSCs. SSEA4+ piPSCs underwent 21 days of motor neuron differentiation first being exposed to retinoic acid (RA) and sonic hedgehog (SHH) for 7 days and then brain-derived neurotrophic factor (BDNF), glial-derived neurotrophic factor (GDNF), and insulin-like growth factor 1 (IGF1) for 14 days. Immunocytochemistry results showed that cells were positive for the motor neuron markers motor neuron and pancreas homeobox protein 1 (homeobox 9 or HB9) (A, C Merge) and ISLET1 (D, E Merge) and the neuron marker βIII-TUB (B, C Merge). Quantification of these markers demonstrated that 85%, 10%, and 6% of cells were positive for βIII-TUB, ISLET1, and HB9, respectively (F). ISLET1 and HB9 showed consistent co-localization within analyzed samples (G–I). *Statistically significant difference p < 0.05 when comparing D 0 and D 21 for each protein.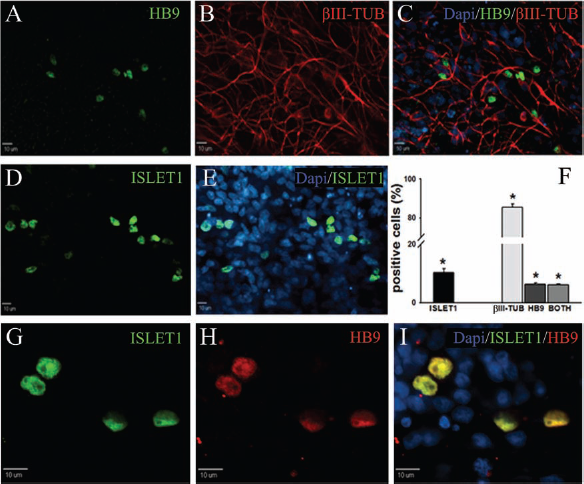
Discussion
In this study, we quantitatively demonstrated for the first time that pig induced pluripotent stem cells are capable of undergoing directed neural differentiation in an adherent culture system forming neurons, astrocytes, and oligodendrocytes. SSEA4 + piPSCs under neural differentiation conditions were capable of forming relatively pure neuron populations where >85% of cells became βIII-TUB+ and produced motor neurons. The ability to derive motor neurons and other specialized neural subtypes are of particular interest in developing treatments for diseases affecting these cells. These cells can be used in vitro to perform comparative studies for diseases such as ALS or in vivo to perform autologous transplant studies to test efficacy and safety of neural iPSC therapies in pig large animal models for spinal cord injury or SMA (30, 31). In this study, SSEA4 was demonstrated to be a robust marker for the isolation of neural competent piPSCs at the gene and protein expression levels. This was a critical finding as significant confusion remains as to what markers are best used to isolate cells of desired developmental potential from piPSCs (12, 13, 49–51). The development of neural competent piPSCs makes it possible for the first time to answer the call for a robust large animal translational model similar to humans in neural anatomy and physiology for research into neural iPSC therapies.
SSEA4+ piPSCs were found to be highly capable of differentiating into a continuum of neural cell types from early progenitors to developmentally restricted cells of all three major neural lineages of the CNS, similar to in vivo-derived pig neural precursor cells (15, 33, 53, 54). The establishment of piPSCs capable of generating cells comparable to in vivo pig neural precursors is an important indicator of the potential functionality of these cells. piPSC-derived neural cells demonstrated neural precursor marker expression with differentiated cells expressing SOX1 similar to examined in vivo-derived counterparts (15, 33, 53, 54). After 10–20 days of differentiation, piPSCs displayed morphological characteristics indicative of a neuron cell fate with the formation of extensive networks of neurite outgrowths. In addition, these cells expressed and showed identical intracytoplasmic localization of neuron proteins βIII-TUB and MAP2, astrocyte protein GFAP, and oligodendrocyte protein O4 to that of differentiated in vivo-derived pig neural precursors under similar in vitro differentiation conditions. These results indicate that piPSC-derived neural cells are analogous to in vivo-derived pig neural precursors, suggesting that these cells should be functional equivalents. However, to confirm the ability of piPSC-derived neural cells to successfully function, additional tests are needed to determine their capacity to form synapses and produce action potentials; additionally, piPSC-derived glia will need to demonstrate that glia are capable of neurotransmitter uptake and can secrete growth factors comparable to endogenous porcine glial cells. Recent studies of human pluripotent stem cell-derived neural cells with similar gene and protein expression phenotypes have proven to be capable of forming synapses and producing action potential in vitro as well as integrating and differentiating into functional cells resulting in notable recovery in rodent CNS injury models. (5, 11, 22, 28). These results suggest that piPSC-derived neural cells are likely to be functional, but further studies will be important to fully characterize their functional potential.
For piPSC transplantation models to be truly indicative of what will occur in humans if iPSC-derived cells are transplanted into patients, it is important that piPSC-derived neural cells demonstrate comparable phenotypic characteristics after exposure to human iPSC neural differentiation conditions. piPSCs demonstrated a high predisposition to differentiate into neural cells with subpopulations of cells even in nonneural differentiation cultures displaying spontaneous differentiation into SOX1+ NPs. To induce neural differentiation, piPSCs were plated on Matrigel, a lamininrich extracellular matrix used to facilitate increased neural differentiation in human pluripotent stem cells and derived neural cell types (9, 32, 47). The piPSCs were allowed to become confluence, and medium was changed to human neural medium for up to 20 days. Similar to human iPSC cultures, piPSC neural differentiation cultures showed high levels of cells expressing later stage neuron, astrocyte, and oligodendrocyte proteins (7, 26, 55). However, it is important to note that differentiation of human iPSCs to later stage neural cells in a short time period such as 20 days typically requires the addition of significantly more growth factors (e.g., noggin or BDNF) to increase neural differentiation progression. The increased speed at which piPSCs are capable of forming later stage neural cell types may perhaps be related to species-specific difference or it may possibly be the result of SOX1+ cells found in the piPSC starting populations. In this study, motor neurons were derived from piPSCs using a human pluripotent stem cell protocol. piPSCs were exposed to RA and SHH for 7 days and then BDNF, GDNF, and IGF1 for an additional 14 days (29, 48). Again there appears to be a difference in temporal development; however, the fact that piPSCs respond in a similar manner as their human counterparts to differentiation signaling demonstrates that neural developmental pathways are highly conserved between pigs and humans. Developmental conservation and response to signaling factors between human and pig iPSCs can also be seen in a recent study where piPSCs are differentiated into rod photoreceptors using a protocol developed for human pluripotent stem cells (56). piPSCs demonstrate robust generation of rod cells with morphological features, gene and protein expression indicating that these cells were comparable to rod cells derived from hiPSCs. This study showed robust differentiation of piPSCs into rod photoreceptors; however, the neural differentiation potential of these piPSCs were not extensively examined beyond their ability to form rod cells. The ability of piPSCs to respond to human neural differentiation conditions in a similar fashion as human iPSCs makes a strong case that findings in a pig neural iPSC therapy model would likely translate to humans.
piPSCs also present a unique opportunity to study another key hurdle blocking iPSC therapy, which is the potential of transplanted iPSCs to cause tumor formation (10, 14, 23, 40, 41, 50). iPSCs are derived utilizing known oncogenic factors including POU5F1 and are often generated using viral approaches. The reactivation of reprogramming genes has been closely linked to tumorigenicity in mice, often leading to death (40, 41). However, a previous in vivo study of chimeric pigs by our group demonstrated that chimeric animals and specifically chimeric tissues did not show any evidence of tumor formation. This raises the question of species-specific differences with respect to iPSC tumorigenicity. In this study, we demonstrated that differentiated βIII-TUB+ cells were predominantly negative for POU5F1. The downregulation of POU5F1 in differentiated neural cells again suggests that these cells will not be tumorigenic. This working hypothesis is further supported by previous findings of Zhou et al. where pig iPSC-derived rod photoreceptors showed downregulation of POU5F1 and other tumorigenic genes (56). After transplant of piPSC-derived rod photoreceptors into pig retina, no tumors were found. Recent studies with human and mouse pluripotent cells have demonstrated that 1) tumorigenicity can be decreased by using purification methods to eliminate undifferentiated cells, 2) the immune system may also decrease tumorgenecity, and 3) the careful selection of less tumorgenic iPSC lines may allow for the safe use of iPSCs and derived cell lines (10, 14, 23, 36). The piPSC model can be used for the development of strategies to decrease tumorigenicity and to study the causes of iPSC tumor formation in a phylogenetically diverse non-rodent system. In addition to safety, there are still a number of efficacy questions that remain to be addressed. For example, transplant of human and mouse pluripotent stem cells, NPs, and early neurons and glia have all resulted in significant improvement in animal models for multiple sclerosis, stroke, and spinal cord injury (1, 3, 8, 10, 46). This raises the important question of which cell stage to transplant will lead to the fastest and most complete recovery. In this study, we demonstrated that distinct neural populations can be derived from piPSCs, which have the potential to study this critical transplantation question.
SSEA4+ piPSCs demonstrated a significantly higher potential to form neural subtypes relative to SSEA4-piPSCs. This increased potential to form neural subtypes may not only suggest that SSEA4 is a marker for neural competent piPSCs but more likely indicates that SSEA4+ piPSCs have an overall higher level of plasticity due to being fully reprogrammed. However, other reports suggest that pluripotent piPSCs should be SSEA1+ and SSEA4- (13) (11). This has left the piPSC field with significant questions to which markers are best to define truly pluripotent stem cells and further highlights the need for more in depth study of markers that maybe linked to specific levels of plasticity and differentiation potentials in the pig.
Conclusion
In this study, we demonstrated SSEA4+ piPSCs undergo robust and developmentally ordered differentiation with early cultures possessing a higher percentage of SOX1+ NPs and later cultures having significantly large populations of neurons (up to 85%), astrocytes, and oligodendrocytes as indicated by gene and protein expression levels. With the addition of neural signaling factors, piPSCs were ultimately found to be capable of directed motor neuron differentiation. These findings demonstrate that piPSCs are proficient at in vitro neural differentiation utilizing a system similar to that used for human iPSCs, thus supporting their potential to be used in biomedically relevant large animal models for the development of neural iPSC therapies.
Footnotes
Acknowledgments
We would like to thank Julie Nelson at the Center for Tropical and Emerging Global Diseases Flow Cytometry, University of Georgia. The authors declare no conflicts of interest.
