Abstract
The therapeutic use of stem cells to treat diseases and injuries is a promising tool in regenerative medicine. The umbilical cord provides a rich source of stem cells; we have previously reported a population of stem cells isolated from Wharton's jelly. In this report, we aimed to isolate a novel cell population that was different than those found in Wharton's jelly. We isolated stem cells from the subepithelial layer of the umbilical cord; the cells could be expanded for greater than 90 population doubling and had mesenchymal stem cell characteristics, expressing CD9, SSEA4, CD44, CD90, CD166, CD73, and CD146 but were negative for STRO-1. The cells can be directionally differentiated and undergo osteo-, chondro-, adipo-, and cardiogenesis. In addition, we have identified for the first time that mesenchymal stem cells isolated from umbilical cord can produce microvesicles, termed exosomes. This is the first report describing a stem cell population isolated from the subepithelial layer of the umbilical cord. Given the growth capacity, multilineage potential, and most importantly the low levels of HLA-ABC, we propose that this novel cell isolated from the subepithelial layer of umbilical cord is an ideal candidate for allogeneic cell-based therapy.
Introduction
Mesenchymal stem cells (MSCs) are a valuable source of cells that have the capability of self-renewal and multi lineage potential and have shown promising clinical results in safety and efficacy studies for treating cardio vascular and orthopedic disorders (14,15,21). Bone marrow-derived mesenchymal stem cells (BM-MSCs) have been one of the most studied stem cell types, and their clinical translation has advanced to clinical trials seeking to develop cell-based therapies (3,11,19,20,23,24). The immunosuppressive properties of MSCs (2,13) make the cells a good candidate for potential allogeneic uses and treatment for acute graft-versus-host disease (aGVHD) (18). Recently, extraction and characterization of MSCs derived from pre- and postnatal tissues, such as placenta and umbilical cord, as well as from umbilical cord blood (UCB), have been described (1,5,16).
Inside the umbilical cord lining is a jelly-like matrix material that, in addition to providing cellular composition, protects the cord arteries and the vein. This gelatinous matrix was initially described by Thomas Wharton in 1656 and termed as “Wharton's jelly.” Mesenchymal stem cells derived from the Wharton's jelly of the umbilical cord have the same characteristics of BM-MSCs; they adhere to plastic, express characteristic surface markers, and have multilineage potential by being able to differentiate into cell of mesenchymal origin, including bone, cartilage, and adipose tissue (5,7,10,17,22).
We have previously described a population of mesenchymal stem cells derived from Wharton's jelly (5). In this study, we identify a novel population of MSCs derived from the subepithelial layer of the umbilical cord (Fig. 1), the cells express pluripotent markers octamer-binding transcription factor 4 (Oct4) and sex-determining region Y box 2 (Sox2) and, unlike other MSC populations derived from umbilical cord, do not express STRO-1. The cells can be directionally differentiated and undergo osteo-, chondro-, adipo-, and cardiogenesis. In addition, we have identified for the first time that MSCs isolated from umbilical cord can produce microvesicles, termed as exosomes. This is the first report describing a stem cell population isolated from the subepithelial layer of the umbilical cord.
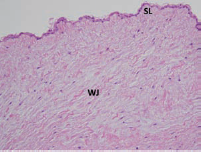
H&E staining of umbilical cord tissue. SL, subepithelial layer; WJ, Wharton's jelly.
Materials and Methods
Umbilical Cord Tissue Procurement
Umbilical cords (n = 5) were obtained from Akron Biotech (Boca Raton, FL). All umbilical cords were obtained from clinically normal pregnancies after signing of informed consent.
Umbilical Cord Tissue Processing
Umbilical cord tissue was washed with Hank's balanced salt solution (HBSS) (Hyclone, Rochester, NY). The cords were dissected; care was taken to remove the artery, veins, and Wharton's jelly. For histological analysis, a 10-mm piece of umbilical cord was placed into 4% paraformaldehyde. The remaining subepithelial layer was then placed into Dulbecco's phosphate-buffered saline (DPBS) containing 10% XcyteMLPL™ supplement (JadiCell, Salt Lake City, UT).
Histological Analysis of Umbilical Cord Tissue
For histological analysis, 5-μm deparaffinized sections were prepared and stained using the haematoxylin/eosin (H&E) technique by Arup Laboratories (Salt Lake City, UT).
Derivation of MSC From Umbilical Cord Tissue
The subepithelial layer was cut into 10 × 10-mm sections and placed into six-well dishes. A 22 × 22-mm sterile coverslip (Fisher Scientific, Pittsburgh, PA) was placed over the subepithelial layer. Explants were cultured in Dulbecco's modified Eagle's medium (DMEM) low glucose, 10% XcyteMLPL™ supplement, 1x penicillin/streptomycin, 1x Glutamax, and 1x minimum essential medium with nonessential amino acids (MEM-NEAA; Life Technologies) and cultured in 5% CO2 incubators at 37°C.
Cell Culture
After 2 days, the cells were 70% confluent and trypsinized using trypLE (Life Technologies). Cells were passaged onto T225 flasks at a density of 1,000 cells/cm2.
Exosome Isolation
Supernatant was collected from cells at 70% confluency. The supernatant was spun at 3,000 × g for 30 min to remove cells and cellular debris. The supernatant was carefully removed and incubated with a 1:1 mixture of thromboplastin D. Medium was incubated for 30 min at 37°C and spun at 4,800 × g for 20 min to remove any precipitate. The supernatant was collected and filtered through a 25-mm 0.2-μM syringe filter. The filtered supernatant was then ultracentrifuged at 100,000 × g for 90 min at 4°C to generate an exosome pellet. The pellet was resuspended in 300 μl PBS at 4°C overnight. Protein content was measured using the Qubit protein assay kit and Qubit 2.0 (Invitrogen, Carlsbad, CA) and then stored at −80°C.
Electron Microscopy (EM)
A total of 10 μl of PBS suspended exosomes were fixed with an equal volume of 4% paraformaldehyde. A total of 5 μl was placed onto a sheet of parafilm, and a formvarcoated EM grid was placed on top and allowed to float on the fixed exosomes for 20 min. The grid was washed twice in PBS for 2 min each. The grid was then postfixed with 1% glutaraldehyde for 5 min. The grid was then washed twice in PBS. To stain the exosomes, 50 μl of uranyl acetate, pH 7.0, was added to the grid for 1 min. The uranyl acetate was blotted from the grid on filter paper and allowed to dry for 10 min. Analysis was made with a FEI TECNAI T-12 electron microscope.
Growth Characteristics
Growth kinetics experiments were performed as previously described (5). Briefly, to generate a growth curve and determine population doublings (PDs), cells were cultured in 25-cm2 flasks, harvested, counted, and replated when they reached 80% confluency. Cultures were terminated after the cells failed to double after 1 week in culture. Population doublings was calculated as previously described (4).
Karyotyping
Karyotyping was performed by Cell Line Genetics (Madison, WI) using standard cytogenetic protocols. Briefly, G-banding was performed on karyograms developed from at least 20 metaphases of cells in log phase.
Immunocytochemistry
Passage 2 cells were collected and fixed with 4% paraformaldehyde in 1x phosphate-buffered saline (PBS) (pH 7.4) for 30 min at room temperature. Cells were then rinsed three times with 1x PBS and blocked with 5% normal donkey serum overnight at 4°C. Cells were incubated at 4°C with primary antibody diluted in 5% donkey serum with 1x PBS overnight (1:200). Cells were then washed three times with blocking solution. Cells were incubated with secondary antibody in blocking solution for 2 h at room temperature. Counterstaining of the cell was accomplished with DAPI (Molecular Probes, Carlsbad, CA) and mounted with antifading mounting solution. Primary antibodies used were homing cell adhesion molecule (H-CAM; CD44) clone F10-44-2 (Millipore, Billerica, MA), CD90 clone F15-42-1 (Millipore), CD146 clone P1H12 (Millipore), CD19 clone FMC63 (Millipore), and CD14 clone 2D-15C (Millipore). Secondary antibodies cyanine 3 (Cy3) and fluorescein isothiocyanate (FITC) were used (Millipore). For negative controls, incubation without primary antibody and with corresponding specific nonimmune immunoglobin (Millipore) in blocking buffer was used. Stained cells were observed under a fluorescent microscope (Olympus Optical, Melville, NY). For immunocytochemistry (ICC) of cardiogenic differentiated cells, a cardiomyocyte characterization kit (Millipore) was used, while for osteogenesis and adipogenesis ICC experiments, a mesenchymal stem cell identification kit (R&D Systems) was used.
Flow Cytometry
Passage 2 cells were collected, resuspended in MEM + HEPES (Life Technologies) with 2% bovine serum albumin (BSA; Sigma Aldrich), and counted. The directly conjugated antibodies used were human leukocyte antigen (HLA)-DP DQ DR (BD Biosciences), CD90, LIN, CD166, STRO-1, stage-specific embryonic antigen 4 (SSEA-4), CD44, CD106, CD73, CD117, CD105, HLA-ABC, CD86, CD63, CD9, CD80 (Biolegend), CD45, CD133, and CD34 (Miltenyi Biotech). After staining, the cells were fixed with 4% paraformaldehyde (PFA) washed three times with DPBS and analyzed using a FACSCanto II analyzer (BD Biosciences).
RNA Extraction and RT-PCR Analysis
Total RNA was extracted using RNeasy Plus Kit (Qiagen) according to the manufacturer's instructions. RNA was reverse-transcribed using LongRange Reverse Transcriptase using standard procedures to synthesize first-strand cDNA. RT-PCR products were resolved on agarose gels and analyzed on a Bio-Rad Molecular Imager ChemiDoc XRS+ System. Primer sequences can be found in Table 1.
Primer Sequences Used in This Study

In Vitro Cell Differentiation
Adipogenic Differentiation
Fifty thousand cells per well were plated into six-well dishes and cultured until the cells reached 90% confluency. StemPro® Adipogenesis Differentiation medium (Life Technologies) was added. The medium was changed every 3 days, and after 21 days, the cells were fixed with 4% PFA. Control cells were maintained in 10% culture medium. Oil Red O (0.3%; Sigma Aldrich) was used for staining to detect intracellular lipid accumulation.
Osteogenic Differentiation
Fifty thousand cells perwell were plated into six-well dishes and cultured until the cells reached 90% confluency. StemPro® osteogenesis differentiation medium (Life Technologies) was added. The medium was changed every 3 days, and after 21 days, the cells were fixed with 4% PFA. Control cells were maintained in 10% culture medium. Alizarin Red S (2%; Sigma Aldrich) was used for staining to detect de novo formation of bone matrix.
Chondrogenic Differentiation
Cells were harvested, and 700,000 cells/15 ml tube were pelleted and induced with StemPro® chondrogenesis differentiation medium (Life Technologies). Control cells were maintained in 10% culture medium. Cell pellets were stained with 1% Alcian Blue (Sigma Aldrich) to detect the secretion of sulfated glycosaminoglycans.
Cardiogenic Differentiation
Fifty thousand cells per well were plated into six-well dishes and cultured until the cells reached 90% confluency. Cells were treated with 10 μM 5-azacytidine. The medium was changed every 3 days, and after 21 days, the cells were fixed with 4% PFA. Control cells were maintained in 10% culture medium. Cells were analyzed for cardiac-specific ICC.
Results
Derivation and Characterization of MSCs Isolated From the Subepithelial Layer of Umbilical Cord Tissue
In order to identify a stem cell population within the subepithelial layer of umbilical cord tissue, we carefully dissected and separated out the veins, artery, and Wharton's jelly components of the umbilical cord tissue. Explants were generated and plated into tissue culture plates. Fibroblastic-like cells were successfully derived from all five umbilical cords, representing 100% derivation efficiency. After 2 days, the cells reached 70% confluency and could be expanded to greater than 90 population doublings within 70 days of culture (Fig. 2), without changes in either growth patterns or morphology. RT-PCR analysis of the cells at passage 2 demonstrated that the cells expressed OCT4 and SOX2 but were negative for nanog (Fig. 3). To further characterize these cells, we performed flow cytometry and immunocytochemistry. The cells exhibited characteristics similar of MSCs (6,8,13) but not identical. The cells were positive for CD9, SSEA4, CD44, CD90, CD166, CD73, and CD146 (Fig. 4), but negative for hematopoietic markers CD14, CD34, and CD45. Interestingly, the cells did not express STRO-1, which we have previously found expressed in Wharton's jelly cells (5,9). ICC demonstrated that the cells were positive for CD44, SSEA4, and CD146 (Fig. 3).

(A) Subepithelial layer (SL) explant 2 days in culture. (B) Culture of mesenchymal stem cells after 5 days in culture of the SL explant. (C) Growth kinetics of subepithelial layer-derived mesenchymal stem cells. (D) Cytogenetic analysis of cells demonstrating normal karyotype.
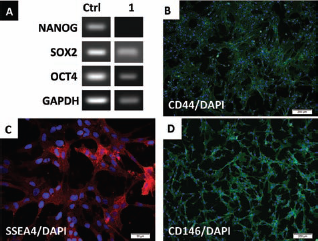
(A) RT-PCR analysis of pluripotent markers. (B) Immunocytochemistry of CD44 expression. (C) Stage-specific embryonic antigen-4 (SSEA4) expression of cultured cells. (D) CD146 staining of cultured cells. SOX2, sex-determining region Y box 2; OCT4, octamer-binding transcription factor 4; GAPDH, glyceraldehyde 3-phosphate dehydrogenase.

Flow cytometry of subepithelial-derived mesenchymal stem cells. Cells express CD90 and CD44 and are negative for STRO-1.
Multilineage Differentiation of MSCs Isolated From the Subepithelial Layer of Umbilical Cord Tissue
Passage 2 cells were induced to differentiate into osteo-, chondro-, adipo-, and cardiogenic cell lineages to determine multilineage potential. After 3 weeks of induction, the cells demonstrated the ability to differentiate into osteoblasts, chondrocytes, adipocytes, and cardiomyocytes. When induced to differentiate under osteogenic promoting conditions, the cells formed a mineralized matrix; this was confirmed by alizarin red staining (Fig. 5), ICC staining of osteocalcin (OC), and RT-PCR analysis for osteopontin (OP), osteonectin (ON), and alkaline-phosphatase (AP) (Fig. 5). Chondrogenic differentiation was confirmed by alcian blue staining for sulfated proteoglycans on induced cell pellets (Fig. 6); RT-PCR confirmed expression of collagen 2A, biglycan (BG), and annexin A6 (A6) (Fig. 6). Adipogenic differentiation was confirmed by Oil Red O staining of lipid droplets (Fig. 7). ICC confirmed expression of fatty acid binding protein 4 (FABP4) (Fig. 7); RT-PCR confirmed expression of FABP4, lipoprotein lipase (LPL), and peroxisome proliferator-activated receptor γ coactivator 1α (PPARγC1A) (Fig. 7). Cardiogenic differentiation was confirmed by ICC staining of induced cells with troponin I and tropomyosin (Fig. 8); RT-PCR analysis of RNA isolated from induced cells demonstrated expression of myogenic factor 5 (myf5), gap junction protein, α1 (connexin 43, cx43), and actin (Fig. 8). Our findings demonstrate that the subepithelial layer of umbilical cord tissue is a source of stem cells that have unique properties different from those found in previous reports (5,7,10,17,22) and have the ability to differentiate into multiple cell types.

(A) RT-PCR of osteogenic markers. (B) Osteocalcin immunocytochemistry staining of differentiated cells. (C) Undifferentiated subepithelial-derived mesenchymal stem cells. (D) Alizarian Red staining of differentiated subepithelial-derived mesenchymal stem cells. OP, osteopontin; ON, osteonectin; AP, alkaline phosphatase.
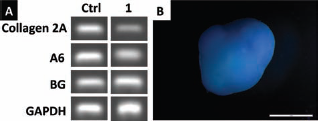
(A) RT-PCR of chondrogenic markers. (B) Alcian blue staining of chondrocyte differentiation subepithelial-derived mesenchymal stem cells. A6, annexin A6; BG, biglycan.

(A) RT-PCR of adipogenic markers. (B) Fatty acid binding protein 4 (FABP4) immunocytochemistry staining of differentiated cells. (C) Phase contrast image of adipogenic differentiation. (D) Oil Red O staining of differentiated cells. LPL, lipoprotein lipase; PPARγC1A, peroxisome proliferating activating receptor γ coactivator 1α.

(A) RT-PCR of cardiogenic specific markers. Cells expressed myogenic factor 5 (myf5) and connexin 43 (cx43). (D) Merged image demonstrating dual staining of troponin I and tropomyosin.
Exosome Purification of MSCs Isolated From the Subepithelial Layer of Umbilical Cord Tissue
Supernatants were collected from cultured cells and ultracentrifuged. Confirmation of the presence of exosomes was confirmed by electron microscopy (Fig. 9).

Electron microscopy (17,000x) of subepithelial layer-derived mesenchymal stem cell exosomes.
Discussion
We have previously described a population of stem cells by explanting the gelatinous portion of the umbilical cord to derive a cell that expresses markers characteristic of mesenchymal stem cells (5). In this report, we describe a novel population of stem cells isolated from the subepithelial layer of the umbilical cord, we have characterized this population of cells and demonstrate their growth potential, marker expression, and differentiation potential. The cells can be cultured for greater than 90 population doublings with no indication of senescence, changes in morphology, or changes in their multilineage potential. The cells were capable of differentiating into chondrocytes, osteocytes, adipocytes, and cardiomyocytes; this was confirmed by RT-PCR and immunocytochemistry. In addition, we have further characterized this cell population by demonstrating that they secrete exosomes. Previous reports have identified exosome production in mesenchymal stem cells derived from bone marrow and demonstrated that these microvesicles play a role in cell-to-cell communication (12).
Various groups have isolated MSC-like cells from the umbilical cord (7,10,17); compared with our previous results, cells isolated from the subepithelial layer have a higher proliferation capacity (Fig. 2) and lower levels of Lin and HLA-ABC. In addition, they do not express STRO-1 (Fig. 4). HLA-ABC expression can be problematic for allogeneic cell-based therapies; the lower expression of HLA-ABC in our cell population allows for allogeneic cell therapy. The use of cell-based therapy in regenerative medicine is expanding; novel cells, methods of culture, and delivery methods are now being optimized for therapeutic use. Given the growth capacity, multilineage potential, and most importantly the low levels of HLA-ABC, we propose that this novel cell isolated from the subepithelial layer of umbilical cord is an ideal candidate for allogeneic cell-based therapy.
Footnotes
Acknowledgments
Dr. Patel is a member of the SAB for JadiCell LLC. The authors declare no conflict of interest.
