Abstract
BACKGROUND:
Alberta stroke program early CT score (ASPECTS) is a semi-quantitative evaluation method used to evaluate early ischemic changes in patients with acute ischemic stroke, which can guide physicians in treatment decisions and prognostic judgments.
OBJECTIVE:
We propose a method combining deep learning and radiomics to alleviate the problem of large inter-observer variance in ASPECTS faced by physicians and assist them to improve the accuracy and comprehensiveness of the ASPECTS.
METHODS:
Our study used a brain region segmentation method based on an improved encoding-decoding network. Through the deep convolutional neural network, 10 regions defined for ASPECTS will be obtained. Then, we used Pyradiomics to extract features associated with cerebral infarction and select those significantly associated with stroke to train machine learning classifiers to determine the presence of cerebral infarction in each scored brain region.
RESULTS:
The experimental results show that the Dice coefficient for brain region segmentation reaches 0.79. Three radioactive features are selected to identify cerebral infarction in brain regions, and the 5-fold cross-validation experiment proves that these 3 features are reliable. The classifier trained based on 3 features reaches prediction performance of AUC = 0.95. Moreover, the intraclass correlation coefficient of ASPECTS between those obtained by the automated ASPECTS method and physicians is 0.86 (95% confidence interval, 0.56-0.96).
CONCLUSIONS:
This study demonstrates advantages of using a deep learning network to replace the traditional template registration for brain region segmentation, which can determine the shape and location of each brain region more precisely. In addition, a new brain region classifier based on radiomics features has potential to assist physicians in clinical stroke detection and improve the consistency of ASPECTS.
Keywords
Introduction
Acute ischemic stroke (AIS) is an infarction of brain tissue due to cerebral artery occlusion, which accounts for 60% to 80% of all strokes and is characterized by high incidence, high recurrence, and high disability rates [1]. Nowadays, arterial thrombectomy is being performed in various hospitals [2]. Interventional treatment such as injection of intravenous plasminogen at the early stage of AIS can help improve the impaired neurological function of stroke patients, reduce the degree of disability, and improve their quality of life [3].
It is very important to accurately screen cases suitable for thrombus removal. However, there is a phenomenon of underutilization of CT images by clinicians [4, 5]. Therefore, achieving effective diagnosis and assessment of AIS is of great importance for the treatment and prognosis of patients. The American Heart Association recommends Alberta stroke program early CT score (ASPECTS) as one of the evaluation indicators for AIS in the Guidelines for the Early Management of Patients with Acute Ischemic Stroke [6–8]. The total score of ASPECTS is 10 points, and any brain area with early ischemic features will be subtracted by one point. The ASPECTS is a scoring tool for ischemia in the MCA donor region only, dividing the MCA donor region into a total of 10 sections at 2 levels. As shown in Fig. 1: the nucleus cluster levels accounted for 7 zones (C, L, IC, I, M1, M2, and M3), and 3 zones at the level above the nucleus cluster (M4, M5, M6). Then, if the left and right sides of one region show signs of ischemic stroke, such as hypodensity, gray-white distinction, and focal swelling, one point is deducted from the initial score of 10 points. The lower the score, the greater the extent of cerebral infarction and the worse the patient’s prognosis [9].
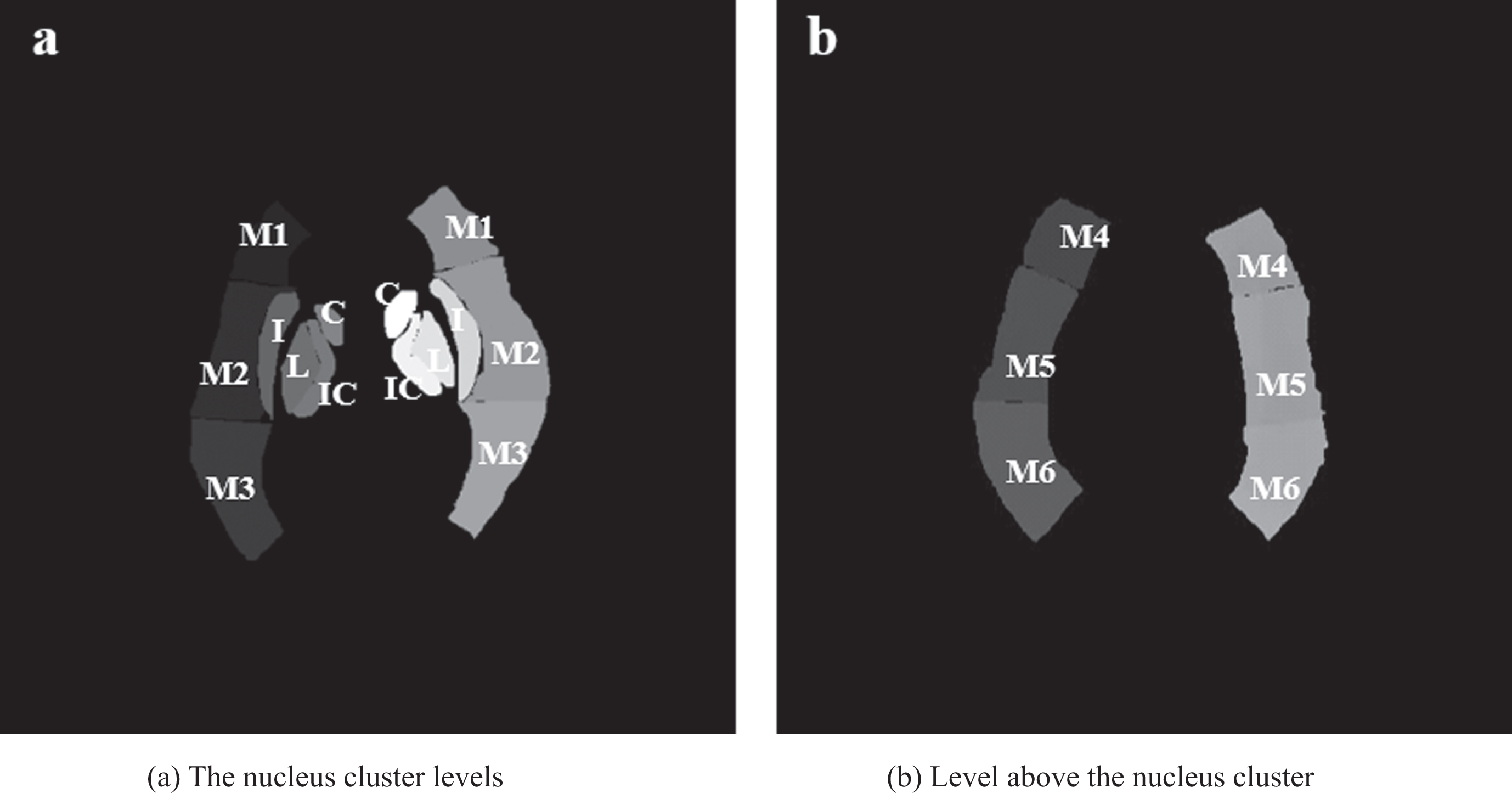
Brain regions template involved in the ASPECTS.
Diffusion-weighted imaging (DWI) is more sensitive to the diffusion motion of water molecules and is mainly used for the determination of ultra-early acute stroke, which is the most accurate assessment of early cerebral infarction [10–12]. However, the DWI examination takes a long time, and time means life for patients. CT angiography (CTA) is important for the imaging evaluation of stroke patients, both for collateral circulation assessment and for early prediction of infarct extent, providing clinicians with additional information [13]. CTA can be obtained in a “One-stop” CT examination of the brain, within a short delay after non-contrast CT (NCCT). Consequently, the use of CTA to extract more evaluation information has a high value for clinical ASPECTS scoring.
Accurate brain region segmentation is the key to achieving ASPECTS scoring. To improve the efficiency of clinical scoring, in this paper, we have developed a deep learning-based computer-aided system using CTA images. In the literature, Shieh et al [14]. have designed an automatic ASPECTS scoring system. The system categorizes patients into high and low scores based on the need for thrombolysis, with high scores of ASPECTS≥6 being able to benefit from thrombolysis, and thrombolysis not recommended for patients with an ASPECTS < 6, and the area under the curve of receiver operating characteristic (AUC) is 0.902. The system uses the registration algorithm to find the brain regions participating in the ASPECTS scoring. A serious weakness with this method, however, is that a brain image used as a registration template needs to be generated using many brain atlases. Otherwise, the model will lack generalization ability. Anbumozhi et al [15]. used gray-level co-occurrence matrix (GLCM) to characterize the second-order texture features of AIS lesions. The final AIS recognition accuracy rate reached 96.15%. It is important to note that Anbumozhi’s research can identify whether there is a cerebral infarction, but it is unable to specify the brain area where a cerebral infarction exists, so it cannot directly guide clinicians to perform ASPECTS scores. In addition, Anbumozhi’s study extracted lots of radiomic features, while our study achieved a 95% accuracy rate of cerebral infarction recognition through only three radiomic features.
In view of the limitations of the above research methods, our contribution in this study is to propose an automatic scoring auxiliary model based on deep learning and image omics. Compared with traditional template registration methods, deep learning significantly improves the efficiency of brain region extraction. On this basis, we used the imaging omics method to determine whether the extracted brain area had ischemic lesions. The results of the consistency analysis indicated that this method could alleviate the current situation of large differences in ASPECTS scores between different neuroradiologists and assist doctors in making better clinical decisions.
Experimental data
This research is derived from the joint medical/industrial project conducted by the University of Shanghai for Science and Technology and the Sixth People’s Hospital Affiliated to Shanghai JiaoTong University. We recruited 90 patients affected by AIS within 6 hours after the stroke’s onset who underwent DWI examinations within 3 hours following the CTA scans. Each patient’s CTA image contained an ASPECTS brain region template created by a professional radiologist (Fig. 2) as the ground truth for the segmentation network. The ASPECTS scoring of DWI images obtained by three experienced radiologists were used as the ground truth for the classification experiments. Table 1 shows the pairwise intra-group correlation coefficients for the ASPECTS scores of the three physicians. All relevant information involving human subjects in this study had been approved before the study.
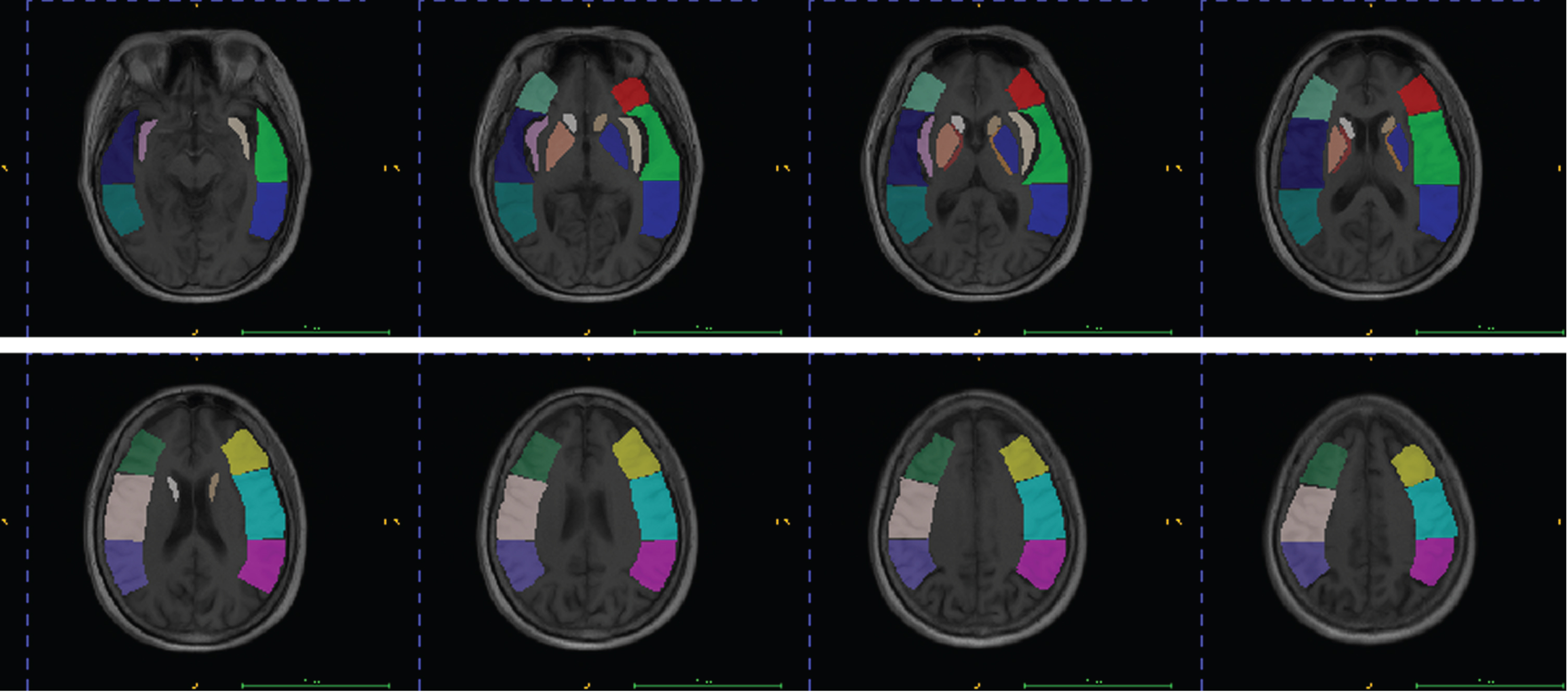
Detailed view of brain region segmentation labels for each slice (same color for same brain region).
Pairwise intra-group correlation coefficients for the results of the three physicians’ scores
After the elimination of abnormal data such as incomplete brain and severe tail shadow from the CTA images obtained from the AIS patients, 271 slices were selected for follow-up research. In the segmentation experiment, 80 patients (200 slices) were randomly selected as the training set, enhanced by rotation, scaling, and left-right flipping, and then trained using the deep learning model. The remaining 10 patients (71 slices) were used as a test set to evaluate the trained model. In the classification experiment, 200 training set slices were utilized for feature screening and classifier training. All experimental CTA images were 512 x 512 pixels in size, and the slice thicknesses were 0.7 mm, 0.8 mm, and 1 mm.
As shown in Fig. 3, The workflow of this study included the following steps: (1) data preprocessing: The preprocessing steps for CTA data include rotation correction, skull stripping and window width/window level adjustment, (2) brain region segmentation: We utilized an improved U-Net architecture for brain region extraction, (3) brain region classification: The obtained brain regions were processed using radiomics to derive a determination of the presence or absence of ischemic changes in brain regions, and finally the total ASPECTS score was calculated. A more detailed description of the experimental scheme is provided in the following subsections.
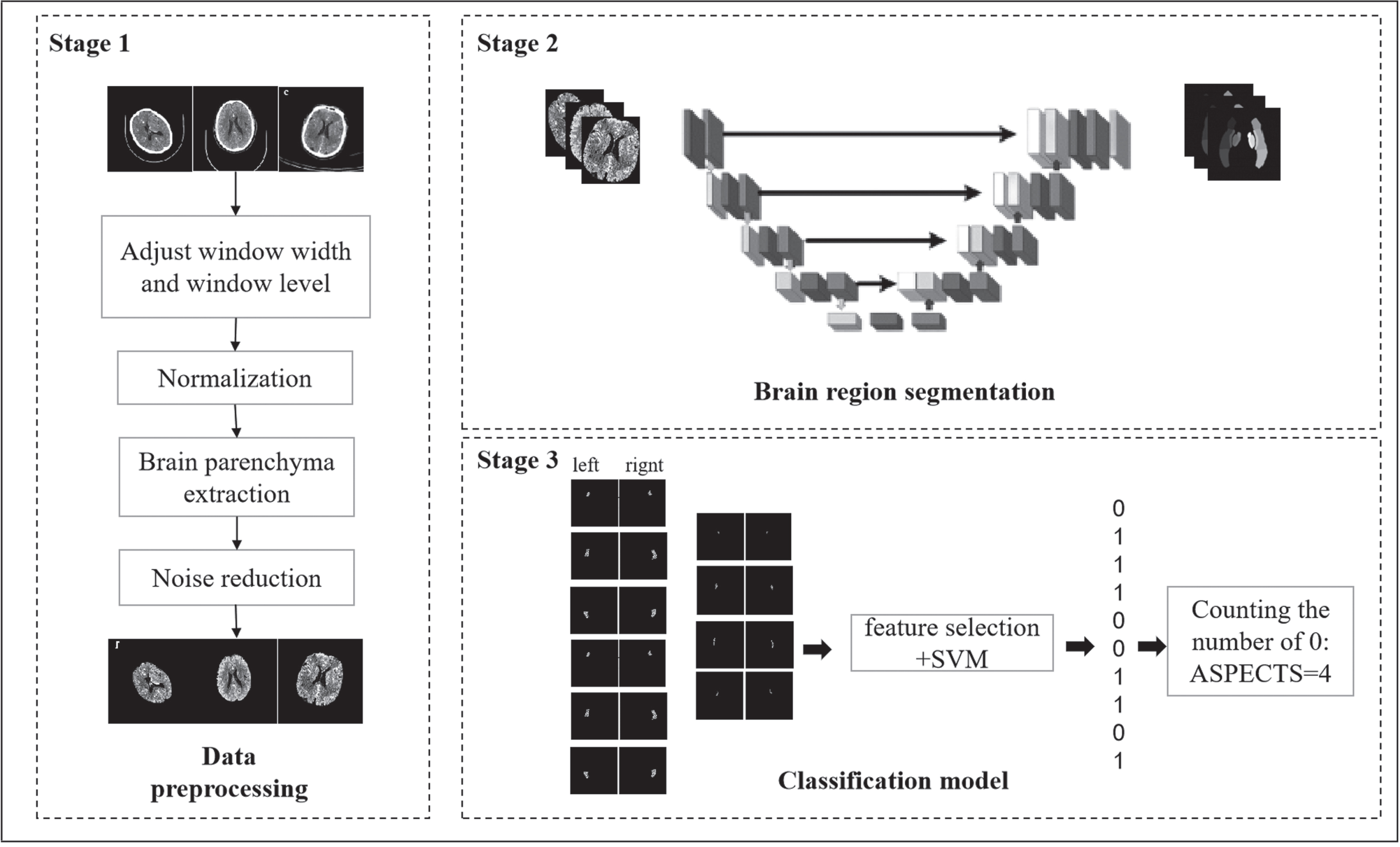
Flow chart of processes in all parts of our proposed system.
The image preprocessing part includes the adjustment of the window width, window position, and extraction of the brain parenchyma. The detailed processing steps are as follows:
First, adjust the window width and level, choose an ultranarrow window that is conducive to the observation of the cerebral infarction focus; specifically, the window width is chosen to be equal to 80, the window level is 40, and the image-gray value is normalized to the range of 0–255.
Subsequently, extract the brain parenchyma from the normalized image. Given that there is a connected domain between the brain parenchyma and the skull in the CTA image, we can easily eliminate interferences, such as the skull and bed board, with the help of morphological operations. The specific extraction step is to open and close the image first to remove noise and fill the cavity, then perform connected domain extraction to remove the bedplate and skull to obtain the brain parenchymal mask. Finally, the brain parenchyma is extracted by multiplying the original image on a pixel-by-pixel basis with the brain parenchymal mask. We randomly selected several preprocessed images as shown in Fig. 4.
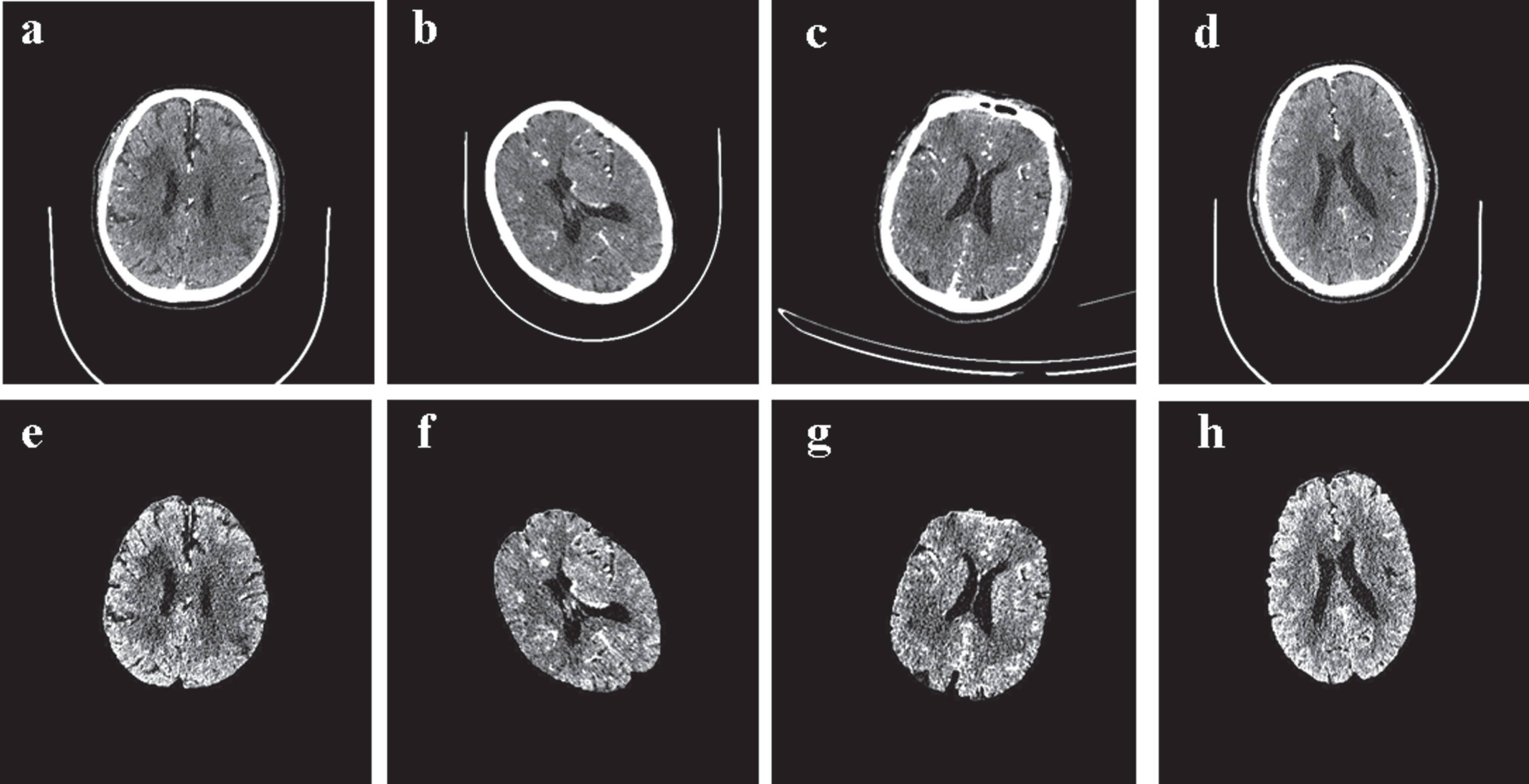
(a)–(d) Original CTA images, (e)–(h) images after processing.
Traditional brain region segmentation methods build energy functions based on image features to extract the contour edges of images. For example, automatic diffusion segmentation is usually based on a fixed Apparent Diffusion Coefficient (ADC) [12]. Deep learning based on labeled training data can avoid the influence caused by ADC threshold delays and improve the segmentation accuracy.
Fully Convolutional Network (FCN) [14] is the first deep learning model to use neural networks for image segmentation. The classic deep learning model U-Net [13] is an improved FCN structure that consists of a compression channel on the left and an expansion channel on the right, which ensures that the sizes of the input and output images through the convolution and deconvolution layers are maintained the same. Relying on the feature extraction of the convolutional layer and the scale transformation of the pooling layer, the original medical image data can provide semantic segmentation information for the subsequent segmentation after three to four down-samplings, as well as the information mapping from the encoder to the decoder of the same level can be realized after the concatenation operation in the network, which improves the details of the segmentation task and achieves more accurate image segmentation. U-Net has been shown to yield excellent performance in the segmentation of small medical datasets. Therefore, in this research we consider using the U-Net as the baseline. However, in this segmentation task, it can be seen from Fig. 1 that the labels of the ten brain regions involved in the ASPECTS scoring in the CTA image have different shapes and locations. While U-Net can only learn the deep-level features through the convolutional network, ignoring the low-level features which are vital for segmentation and is unable to accurately focus on the locations where different brain regions exist. Although the Dense-Block structure [15] proposed in 2017 was claimed to be able to overcome the problem of shallow feature-wasting, but it was also unable to solve the problem that the scoring brain regions that we need to segment are small of the entire brain in this study. To solve the above problems, we proposed a brain region segmentation method for the ASPECTS combined with feature dimensionality reduction and remapping based on deep learning. The feature dimensionality reduction and remapping module is illustrated in Fig. 5.
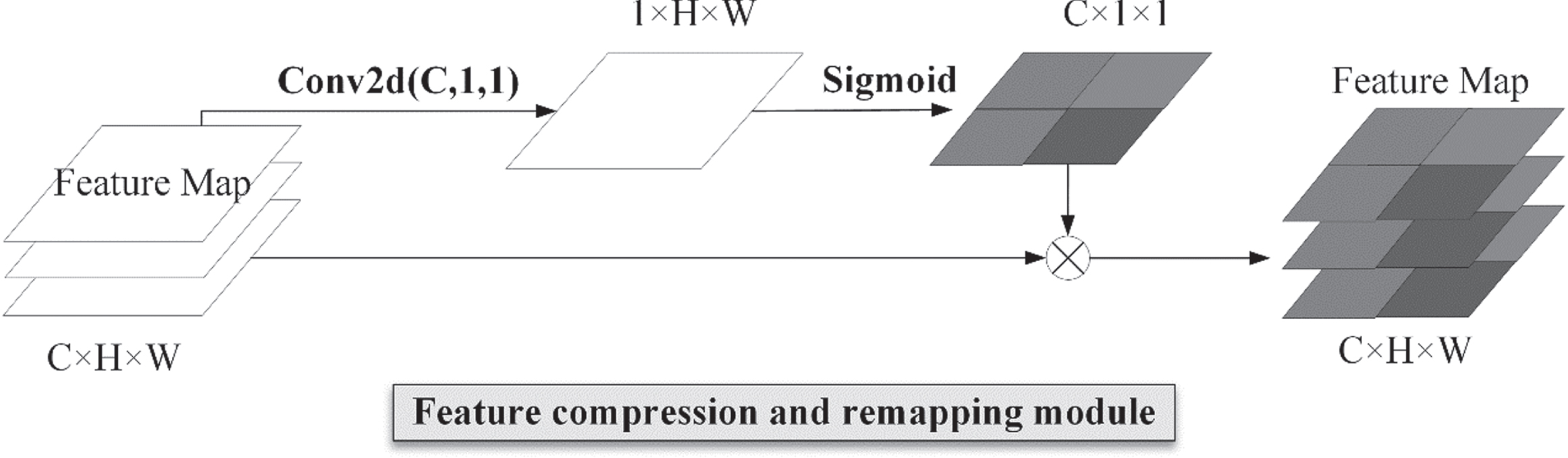
Feature compression and remapping module.
The feature compression and remapping module performs a 1×1 convolution on the input feature map, and the 1×1 convolution kernel is able to perform channel dimensionality reduction on the input image without changing the width and height of the output, reducing the number of feature channels from a certain C to 1; that is, from C×512×512 to 1×512×512, and then uses the sigmoid activation function to activate the feature map to obtain the excitation feature map after feature remapping. Finally, the broadcast mechanism is used to multiply the spatial excitation feature map with the original feature map and assign different weights to different pixel values in the feature map to complete the spatial information calibration.
The network (Fig. 6) extracted 512×512 images from the CTA image dataset of stroke patients obtained in the preprocessing stage as input, and the input channel was set to one. The segmentation model included four layers of down-sampling and four layers of up-sampling. In this study, the 3×3 convolution was used for feature extraction, and the 1×1 convolution was used to replace the fully connected layer of the original U-Net network to achieve multichannel dimensionality reduction while reducing the amount of calculation.
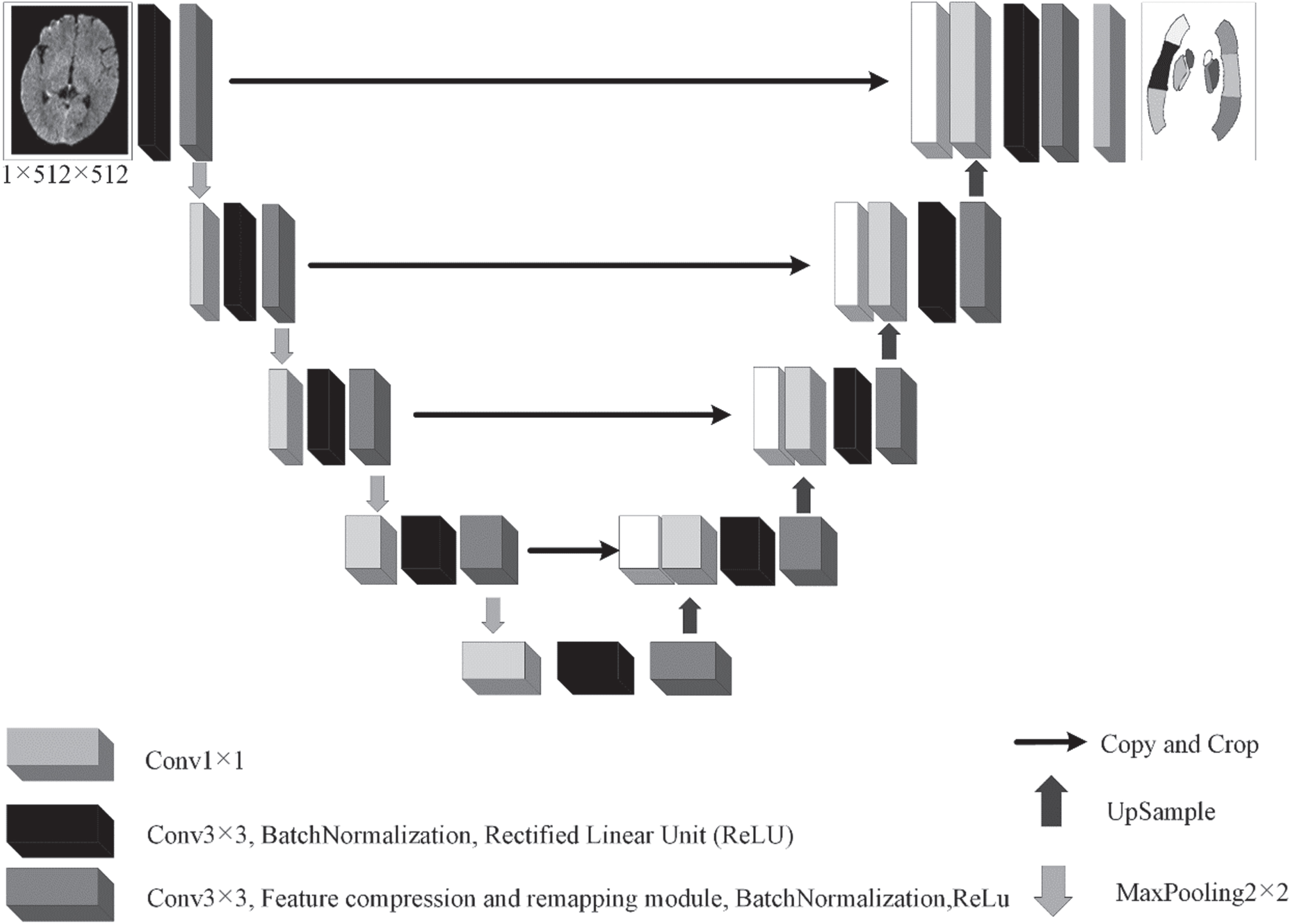
Segmentation model combining feature compression and remapping mechanisms.
In image segmentation tasks, the cross-entropy loss is generally selected as the cost function. The cross-entropy loss function predicts independently the category of each pixel and then averages over all the predicted results. However, there is a class imbalance problem in the brain regions evaluated by the ASPECTS. Therefore, this study uses a composite loss function that combines cross-entropy loss and the Dice coefficient to improve sample imbalance. The expression of the loss is shown in Equation 1.
Specifically, pn,c ∈ P and yn,c ∈ Y. pn,c and yn,c represent the target label and predicted probability of the nth pixels of the class c in the batch, respectively. Y and P are the true values and prediction results of the retinal image, respectively. C and N represent the number of dataset classes and pixels in the batch, respectively.
In the model training stage, this study used the gold standard data marked by physicians for supervised learning to realize brain image segmentation. In the network training process, we used the RMSprop optimizer and L2 regularization to optimize the weight parameters. Through experimental verification, the coefficient of L2 regularization was set to 1e-5, the momentum was set to 0.9, and the training batch-size of training was set to one. The initial learning rate was set to 1e-5.
In clinical practice, stroke usually occurs in one cerebral hemisphere with significant heterogeneity between ischemic and healthy tissues, so we can identify the lesioned area by comparing the diseased brain region with the contralateral healthy brain region [16]. Contralateral analyses can also avoid the influence of scanning equipment and parameters and improve the accuracy of cerebral infarction assessments.
In this study, we trained the network proposed in Section 2.2.2. We then used the trained segmentation model to segment the CTA slices of AIS patients and obtained ten brain regions in each of the left and right hemispheres of the patient’s brain. Using the cerebral infarction region in the sequentially acquired DWI images as the ground truth, we used the numbers “0” and “1” to label the brain area for the presence of cerebral infarction. The number “0” represents normal brain area, and the number “1” represents cerebral infarction. We then used Pyradiomics to extract a total of 105 features about shape, histogram, and texture from each brain region to discriminate between healthy and infarcted brain regions. However, there is multicollinearity in the original features. Therefore, a two-sample t-test and a Lasso feature dimension reduction method were used for feature selection. The disadvantage of Lasso is that when there is a set of highly correlated features, it tends to select one of them and ignore all the others, which can lead to unreliable results. Therefore, we first used a two-sample t-test to screen out 53 features with significant differences, and then combined with Lasso for feature screening to finally determine three features. The features selected after dimensional reduction are listed in Table 2. The “Mean” represents the average grayscale intensity in the region of interest. “RootMeanSquared” is the square root of the mean of all squared intensity values, which can be used to measure the intensity of image’s gray value. “HighGrayLevelRunEmphasis” is used to measure the distribution of higher gray values. The larger the value, the more concentrated the gray scale of the image.
Three selected features after dimensional reduction
Three selected features after dimensional reduction
These three selected features were used to train a Support Vector Machine (SVM) model [17], which could distinguish the presence or absence of cerebral infarction in brain regions. During the experimental training, the hyperparameters of the SVM classifiers were optimized using GridSearchCV and cross-validation methods. Then, we selected the SVM classifier with optimal performance to classify each brain region in the test set obtained from the segmentation network. If there was cerebral infarction, one point was subtracted from the total ASPECTS value.
Based on the two-step approach of brain region segmentation and brain infarct identification proposed in this paper, the clinician can obtain the final ASPECTS value by a simple summary.
Evaluation of ASPECTS brain segmentation model
We used the preprocessed data for training and enhanced the data through morphological operations such as rotation, zooming, and deformation. Besides, we used the callback function to observe the internal state of the network during the training process and save the best network model in real-time. To evaluate the generalization ability and segmentation accuracy of the segmentation model and prove that the segmentation model proposed in this study is effective and feasible, this study selected the Dice coefficient as the evaluation index.
To test the effectiveness of the segmentation model method proposed in this study, this section compares our model with SE-UNet, scSE-UNet, Attention-UNet, SMU-UNet, and basic U-Net. Jie Hu et al. [18] proposed the SE module to improve the attention between channels. The SE module implements an attention mechanism through feature compression and spatial incentive assignment, which can effectively improve the accuracy of segmentation. Roy et al. [19] improved the SE module based on the semantic segmentation problem and proposed the scSE module. The improved SE module can further improve the segmentation accuracy and make the segmentation boundary smoother. Oktay et al. [20] proposed an attention gate for the image segmentation problem of abdominal pancreas, which is used to improve the model’s attention to different target shapes. Koushik et al. [21] tried to propose a new activation function, referred to as the smooth maximum unit (SMU), to replace the commonly used rectified linear unit activation function.
The experimental parameters and average segmentation results for different experimental methods are shown in Table 3. And the specific segmentation DICE coefficients for each brain region are shown in Table 4. Compared with the original U-Net, the dice value of the segmentation model proposed in this study is improved by 0.2 and can reach 0.79. The closer the verification set Dice is to one, the better is the brain segmentation effect of the model. Figure 7 shows the four original CTA images and the corresponding brain regions after segmentation of them using the proposed segmentation network. The results in the figure show very intuitively that our proposed model can accurately segment the shape and location of brain regions.
Experimental parameters and segmentation results corresponding to different experimental methods used herein and in the literature
Experimental parameters and segmentation results corresponding to different experimental methods used herein and in the literature
The specific segmentation DICE coefficients for each brain regions
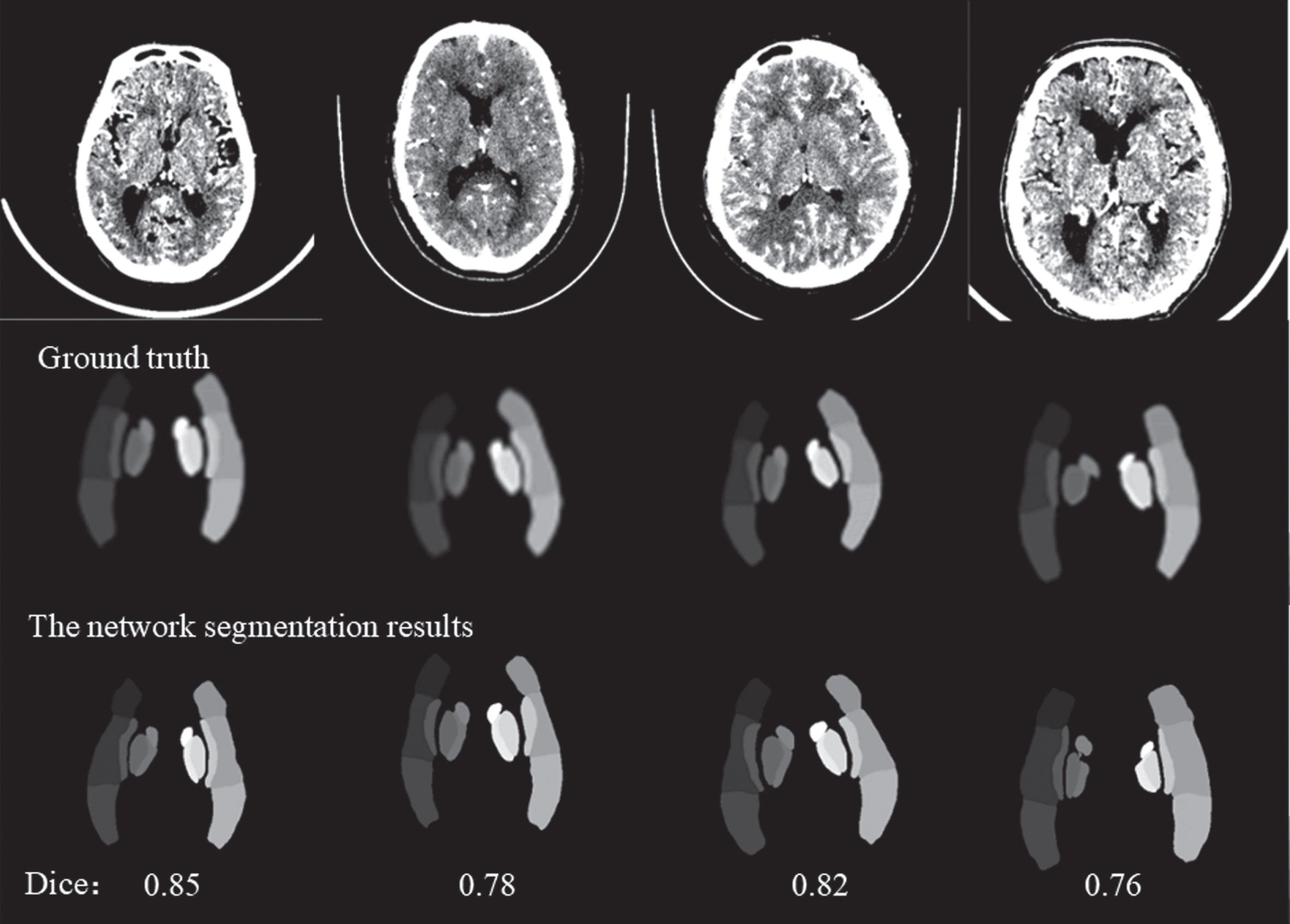
Examples of segmentation network output.
We used the brain regions obtained by the segmentation network to train a SVM classification model based on the three features screened (“Mean”, “RootMeanSquared”, “HighGrayLevelRunEmphasis”), which was used to determine whether there was cerebral infarction in the brain region. Figure 8 shows the comparison of receiver operating characteristic (ROC) between the five-fold cross-validation experiment, and the average AUC is 0.95. In addition, we chose the SVM classifier with optimal performance to classify each brain region in the test set obtained from the segmentation network. The performance of the classifiers for categorization evaluation for each brain region is shown in Table 5. Clinicians could summarize the results of ten brain regions to get the ASPECTS scores based on the judgment of the model (the results are shown in Table 6), compare the obtained scoring results with the manual scoring results of the same batch of data by another physician, and analyze the consistency of the two scores, and the obtained intraclass correlation coefficient was 0.86 (95% confidence interval, 0.56-0.96). The comparison results are shown in Table 7, indicating that our proposed automatic scoring method can help physicians to score and improve the consistency of scoring among different physicians.
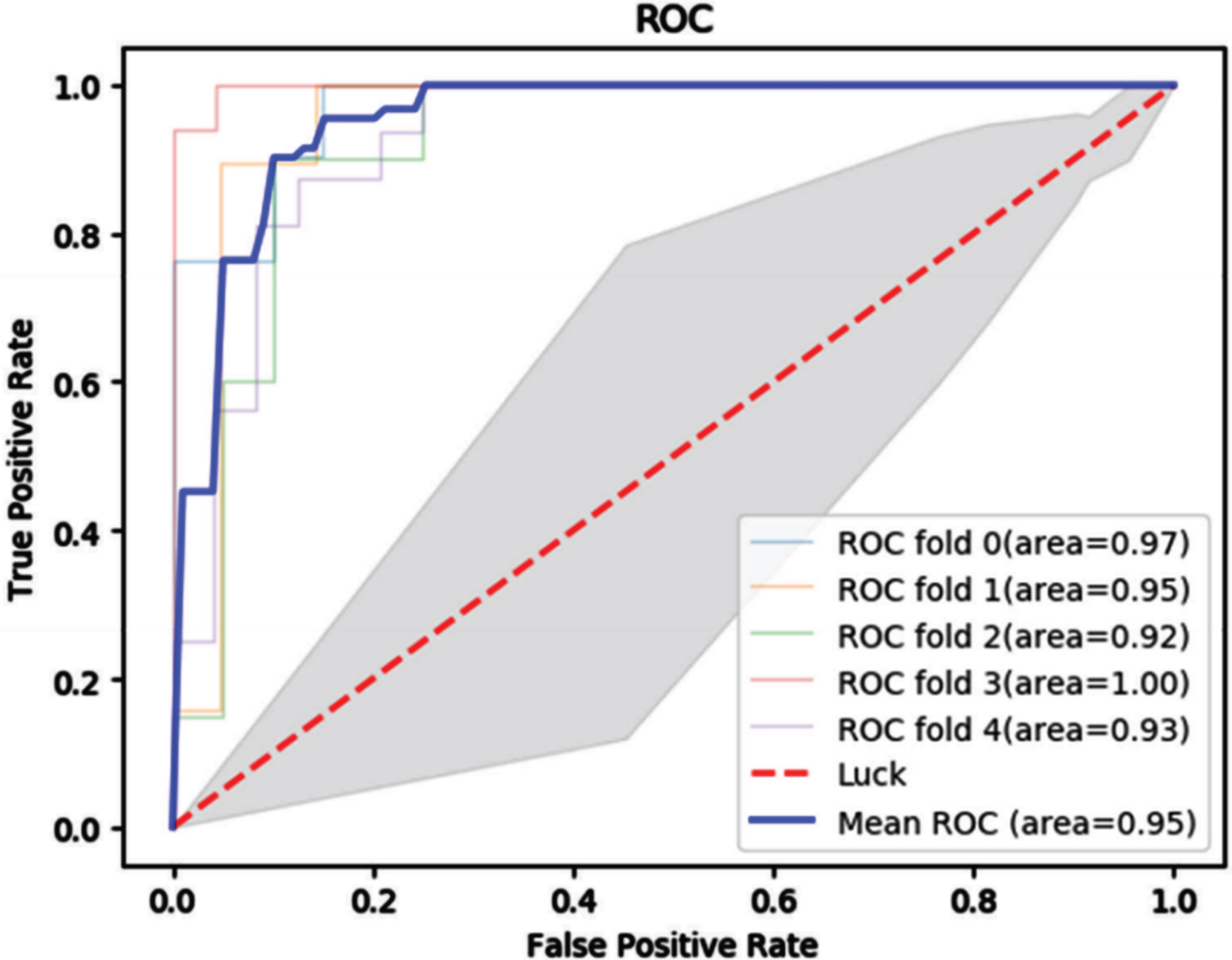
ROC curves used to assess whether there is cerebral infarction in the brain region.
Classification evaluation performance of SVM classifier for each brain region
Results of the overall ASPECTS score based on the model proposed in this paper
Comparison of the consistency of ASPECTS scores based on our model with manual scores
Using 90 patient CTA images for analysis, the ICC coefficients from Table 7 show that the computer-aided ASPECTS method proposed in this article agrees well with expert-read DWI-ASPECTS for the total ASPECTS. This is not the first application of deep learning in the field of stroke, but it is one of the few methods that uses deep learning to replace traditional template segmentation. Shieh et al. [10] designed an automatic end-to-end ASPECTS system using registration algorithm which imposes high accuracy requirements on the standard template, and the model lacks adaptive capabilities. Anbumozhi et al. [11] achieved cerebral infarct identification by radiomics, but failed to achieve automatic segmentation of brain regions and therefore could not locate the brain region where the infarct was located. Our proposed method uses deep learning to effectively overcome the lack of generalization ability caused by template alignment and to solve the problem that traditional radiomics methods are unable to localize brain regions where brain infarcts are located.
Based on the experimental results obtained, we believe that our method has the following advantages compared to other previous state-of-the-art studies in this field: In the brain region extraction stage, we propose a feature reduction and remapping convolutional network that can efficiently learn brain structural features in datasets with a small number of samples. Compared with the traditional template alignment method, the deep learning model significantly improves the efficiency of brain region extraction. In this study, we propose an innovative method for automatic scoring of ASPECTS. The method is based on CTA images, combines deep learning with radiomics, and reported that the three radiomics features of ‘Mean’, ‘HighGrayLevelRun’ and ‘EmphasisRootMeanSquared’ can be used to identify cerebral infarction. The final consistency analysis showed that this method mitigates the wide variation in ASPECTS scores among neuroradiologists and assists clinicians in more quickly and accurately planning treatment for their patients.
This study has some limitations. First, this study is a single-center study with a small sample, so the generalization ability of the model needs to be verified. Second, the number of slices in the test set was small, thus creating some valid concerns about the stability of the ASPECTS analysis results. Validation on a larger data set is needed to demonstrate the robustness of these results.
Conclusions
Rapid ASPECTS scoring is of great significance in the clinical treatment of AIS patients. In this study, we proposed a method that combined radiomics and deep learning to assist clinicians in ASPECTS. This study further validates the reliability of deep learning in the field of stroke and can effectively assist physicians in ASPECTS.
Declaration of competing interest
The authors have no relevant conflicts of interest to disclose.
Footnotes
Acknowledgments
The work was supported by research grants from The National Natural Science Foundation of China (Grant No. 81830052, No. 82271989, No. 82071990, No. 81571629) and Shanghai Key Laboratory of Molecular Imaging(18DZ2260400). We would like to thank Editage (www.editage.cn) for English language editing.
