Abstract
BACKGROUND:
Esophageal cancer (EC) is aggressive cancer with a high fatality rate and a rapid rise of the incidence globally. However, early diagnosis of EC remains a challenging task for clinicians.
OBJECTIVE:
To help address and overcome this challenge, this study aims to develop and test a new computer-aided diagnosis (CAD) network that combines several machine learning models and optimization methods to detect EC and classify cancer stages.
METHODS:
The study develops a new deep learning network for the classification of the various stages of EC and the premalignant stage, Barrett’s Esophagus from endoscopic images. The proposed model uses a multi-convolution neural network (CNN) model combined with Xception, Mobilenetv2, GoogLeNet, and Darknet53 for feature extraction. The extracted features are blended and are then applied on to wrapper based Artificial Bee Colony (ABC) optimization technique to grade the most accurate and relevant attributes. A multi-class support vector machine (SVM) classifies the selected feature set into the various stages. A study dataset involving 523 Barrett’s Esophagus images, 217 ESCC images and 288 EAC images is used to train the proposed network and test its classification performance.
RESULTS:
The proposed network combining Xception, mobilenetv2, GoogLeNet, and Darknet53 outperforms all the existing methods with an overall classification accuracy of 97.76% using a 3-fold cross-validation method.
CONCLUSION:
This study demonstrates that a new deep learning network that combines a multi-CNN model with ABC and a multi-SVM is more efficient than those with individual pre-trained networks for the EC analysis and stage classification.
Keywords
Introduction
Globally, EC is the most frequently reported form of malignancy [1, 2]. This disease ranks seventh based on mortality and sixth in terms of prevalence [3]. Among the deaths due to cancer, EC accounts for 5% of the deaths [3]. Even though the early stages are not fatal, the poor prognosis may reduce the survival rate from 95% to 5% [4, 5]. The early detection of EC is vital for decreasing the mortality rate [5]. Hence, the diagnosis of the early stages of EC is clinically significant. Also, the nature of many Early Esophageal Cancer (EEC) lesions is analogous to that of normal tumor cells, which makes the diagnosis more challenging even for expert clinicians [6]. Various imaging modalities like Computed Tomography, Endoscopy, Positron Emission Tomography, and MRI can be used to evaluate the degree of malignancy.
Esophageal cancer is a lethal cancer with a high prevalence and mortality [1, 7]. The condition of chronic gastroesophageal reflux disease (GERD) transforms the indigenous squamous epithelium to the columnar epithelium and, is defined as Barrett’s esophagus (BE) [8]. The development starts from chronic inflammation to non-dysplastic BE, dysplastic BE, and cancer [9, 10]. It is better to undergo an endoscopic investigation for chronic GERD to decrease the mortality rate [11]. The esophagus’s visible mucosal rifts or erosions are the preliminary indicators of Reflux Esophagitis [12]. The diagnosis of Barrett’s esophagus indicates that the patient has chronic GERD and is more prone to malignancy. Barrett’s esophagus investigation involves endoscopic surveillance of the Z-line, lower esophageal sphincter, and top of the gastric folds [13]. Barrett’s esophagus is the premalignant stage which can be diagnosed by employing endoscopic screening [14]. When the precancerous lesions are identified, treatments are initiated, and the neoplastic progression is monitored.
Esophageal adenocarcinoma (EAC) and esophageal squamous cell carcinoma (ESCC) are the two main kinds of EC [15]. The two tumours differ significantly from one another. The EAC is a severe form of BE that develops in the lower oesophagus close to the stomach area [16]. The ESCC develops in the epithelial mucosa that lines the esophageal lining. Since it is located in the middle or upper esophageal tract, it lacks a precursor [16]. Since ESCC affects flat tissues, polypoid masses that block the lumen emerge as a result [17]. ESCC is a highly invasive cancer with distinct patterns and risk factors [17]. Usually, these cancer types are highly aggressive with a meagre prognosis rate. Even though squamous-cell carcinoma accounts for over 90% of all occurrences of EC worldwide, the incidence and mortality rates of esophageal adenocarcinoma are on the rise [13]. They have surpassed those of esophageal squamous cell carcinoma.
Clinical analysis of the endoscopic images by experts is a time-consuming process. Moreover, there is a shortage of qualified doctors in many of the rural areas. How can we address this problem? An efficient computer-aided detection method could be used in hospitals for real clinical diagnosis of esophageal cancer and its variants. Even though there are a few computer-aided diagnosis methods for the detection of oesophageal cancer, more accurate methods are required for efficient clinical trial. Developing a more accurate models can address this problem. Machine Learning’s evolution has enhanced the AI system’s ability to extract information from scratch and learn from experience [18]. The Deep CNN-based networks attained state-of-the-art image classification and segmentation [19]. Deep learning models in medical applications are enriched by their ability to dynamically acquire the edges and corner features and relate them to objects in the images [20]. Deep learning with CNN is a powerful generic approach exploited for performing many tasks such as classification, object identification, and analysis. The deep learning networks require a vast annotated dataset for training, which the medical sectors lack. The convolution neural networks trained on a large natural dataset can be utilized to overcome this difficulty. The paper proposes a method for detecting EEC stages using features extracted by different pre-trained networks.
Related Work
The author [21] evaluated the ability of a computer-based technique for early neoplasia diagnosis in BE. For evaluating the disease, texture features, colour filters, and ML methods like SVM are used. The system performed with a sensitivity and specificity of 83% for individual image analysis and at patient level, with a specificity of 87% and sensitivity of 86%. A combination of White Light Endoscopy (WLE) and Narrow Band Imaging (NBI) images were used with a ResNet18 model proposed by Van der Putten et al. [22] for better localization of early neoplasia in BE and accurate biopsy. Canny edge detection and image registration are combined to improve image pairs. The multi-modality-based model has a 93% accuracy, 92 % sensitivity, and 95% specificity. Ebigbo et al. [23] performed a BE analysis with a tissue-based ResNet network using a leave one out cross-validation (LOOCV). The specificity and sensitivity of the WLE images are higher than the NBI images, at 88% and 97%, respectively. With a precision of 78.06 % and recall of 77.58%, where the author in [24] proposed a DeepLabV3 + model for EEC segmentation and classification. To improve classification, localization, and segmentation of Barrett’s Esophagus (BE) with dysplasia, the author in [25] proposed a new network structure that used a semi-supervised learning technique with resampling of the training data and diversity concept. With ensemble learning, the model attained a sensitivity of 92.5 %, specificity of 82.5 %, and accuracy of 87.5 %. In [26], the authors introduced an automated CNN model for diagnosing esophageal variabilities. By concatenating local features acquired by the Gabor filters and the DenseNet network, the automated system with Faster RCNN enhances the performance. The model had a 90.2 % recall rate and a 92.1 % precision rate.
To diagnose non-dysplastic BE and BE neoplasia, Struyvenberg et al. [27] developed a ResNet-UNet hybrid model. Transfer learning and ensemble learning strategies were utilized to classify the NBI zoom images. The model achieves an accuracy of 84 %, a ROC value of 95%, a sensitivity of 88%, and a specificity of 78%. In [28], the author proposes a real-time CAD system with a ResNet-based DeeplabV3 to classify BE and Esophageal cancer. Here the last layers or the ResNet consist of dilated convolutions with spatial pyramid pooling for better classification. The CAD system performs the classification with a specificity of 100%, a sensitivity of 83.7%, % and overall accuracy of 89.9%. Cia et al. [29] developed a CNN-based CAD system to detect early ESCC using WLE images. The CNN model has a sensitivity of 97.8%, an accuracy of 91.4%, and a specificity of 85.4%. Everson et al. [30] proposed a CNN-based network for the evaluation of the Intrapapillary capillary loops (IPCLs) pattern using magnified NBI images for the detection of the early squamous cell neoplasia (ESCN) pattern and classification of the stages of ESCN. The proposed model has an accuracy of 93.7%, a sensitivity of 89.3%, and a specificity of 98%. de Groof et al. [31] developed a CAD system trained with the color and texture features of the WLE images. The system performs well for the LOOCV for detecting the BE neoplasia with a sensitivity of 95%, an accuracy of 92%, and a specificity of 85%.
Nakagawa et al. [32] proposed an AI-based Single Shot multi-box Detector (SSD) to detect ESCC and perform the classification based on the depth of invasion. The classification among the various classes, mucosal, submucosal microinvasive (SM1), and submucosal deep invasive cancers, was performed with a sensitivity of 89.8%, the accuracy of 89.6%, and specificity of 88.3%. de Groof et al. [33] proposed a CNN-based model to detect BE neoplasia. A ResNet-UNet-based Gastronet performed the classification with an accuracy of 88.8%, 90% sensitivity, 87.5% specificity, and an F1 score of 88.7%. Hashimoto et al. [34] proposed a 2-stage analysis for the classification and the localisation of the early esophageal neoplasia in BE.
The Xception architecture performs the binary classification followed by a YOLO V2 network for the localisation of the lesion. The proposed models are 95.4% accurate with a specificity of 94.2% and sensitivity of 96.4% for the NBI image and 95.4% accuracy,88.88% specificity, and 98.6% sensitivity for the WLE images. Ohmori et al. [35] proposed a CNN model SSD to develop a model for the diagnosis of ESCC and classification among the superficial esophageal SCC, non-cancerous lesions, and normal esophageal mucosa. The NBI, BLI, and WLI modalities were used in magnified and non-magnified formats. The AI system detected and classified ESCC with a sensitivity of 98%, accuracy 77%, and a specificity of 56% in the magnified format. In the non-magnified format, for WLI, the system performed with an accuracy of 81%, sensitivity and specificity of 90% and 76%, respectively.
For NBI/BLI, the accuracy is 77%, sensitivity is 100%, and specificity is 63%. Tokai et al. [36] proposed a deep learning model for assessing the infiltration depth and classification of ESCC as mucosal, submucosal microinvasive (SM1) cancers and submucosal deep invasive cancers using a GoogLeNet and the detection using SSD. The model performed the classification process with an accuracy of 80.9%, sensitivity of 84.1% and, specificity of 73.3%. An automatic evaluation and localization of EC tissues is proposed by Zhan Wu et al. [37]. The Dual stream network performs the classification of the lesions into normal, inflammation, Barrett and Cancer. The classification is done using the global and local features of the lesion images with a sensitivity of 90.34%, specificity of 97.18%, and accuracy of 96.28%.
Liu et al. [38] introduced the Inception-ResNet model, classifying precancerous tissue and cancer tissues with 85.83% accuracy. Ishihara et al. [39] evaluate the diagnostic performance of an ensembled EUS and the conventional endoscopic images to diagnose ESCC. Initially, non-magnifying endoscopy (non-ME) and magnifying endoscopy (ME) were used to diagnose cancer invasion depth. The model’s performance with the conventional endoscopic images (with the non-ME and ME) has a sensitivity was 50.4%, specificity was 89.6%, and accuracy was 72.9%. When the endoscopic images were ensembled with the EUS, the model’s performance improved with a sensitivity of 64.3%, specificity of 81.2%, and accuracy of 74%.
Du et al. [40] proposed a deep learning model for classifying four major EC stages starting from BE to EAC. The model achieved a classification performance with an accuracy of 90.63%, precision of 89.71%, recall of 89.26%, and F1score of 89.45%. Wang et al. [41] developed a deep learning model to diagnose the EEC and classify them into low- and high-grade dysplasia and ESCC with a sensitivity, specificity, and diagnostic accuracy of 96.2%, 70.4%, and 90.9%, respectively. Mohammad et al. [42] proposed a fuzzy C Means-based method with morphological analysis with three fully connected hidden layers to detect esophageal cancer. The model uses a feed-forward network for the analysis with an accuracy of 95%. Yu et al. [43] proposed an AI-based model for diagnosing different esophageal lesions. The sophisticated model performs both classification and detection of the different lesions with an accuracy of 96.76%.
Contribution of the proposed work
The methods mentioned above have used a single pre-trained CNN to classify and segment various Esophageal abnormalities from the available dataset. Combining the unique features obtained by each pre-trained network can enhance the performance of diagnosis systems. The proposed study integrates features from a more significant number of CNNs for classification. The proposed work has the following contributions. In the proposed work, the features are extracted by more than one pre-trained CNNs. Almost all the prevailing approaches use only one pre-trained network for feature extraction. The proposed study uses a composition of multiple deep learning networks, a wrapper-based feature selection with the Artificial Bee Colony optimization method (ABC), and a multi-class SVM classifier for classification. No existing methods have employed such a combination to classify stages of EC. In this approach, we are performing a multi-class classification with BE, ESCC and EAC using the SVM classifier.
Materials and methods
Materials and dataset
Here, the proposed model performs the classification of Barrett’s esophagus, ESCC, and EAC’s cancerous stages. The publicly available Kvasir dataset [44], with folders labelled as Barrett’s esophagus and esophagitis, is used in this work. It contains images and videos from various endoscopic procedures captured using different endoscope models and manufacturers, including Olympus and Fuji. The dataset contains 94 images of Barrett’s esophagus with a constancy of 612 x 521 to 1278 x 1022. Another publicly available repository [45] contains high-quality Narrow band imaging gastroscopic videos. The dataset comprises of endoscopic videos from Olympus 100,140,160 series systems and Pentax gastroscopic equipment. From these videos 219 BE images, 110 images of ESCC, and 150 images of EAC were obtained from the 44 high-quality gastroscopic videos. A large dataset is a key for deep learning analysis. The data augmentation process is applied to the training dataset, generating more images to improve performance. Additional training images are generated by rotating the existing image at multiple angles (45°, 135°, 330°), pixel shifting, flipping, and pixel elongation both vertically and horizontally. By applying the rotation process we obtained a total of 1028 images with an additional of 210 BE images, 107 ESCC images, and 138 images of EAC were obtained by rotation. A total of 523 BE images, 217 ESCC images and 288 EAC images is used for the analysis and classification purpose. For the proposed study, we obtained about 1028 images in three classes. Figure 1 displays the gastroscopic images of EAC, ESCC, and BE.
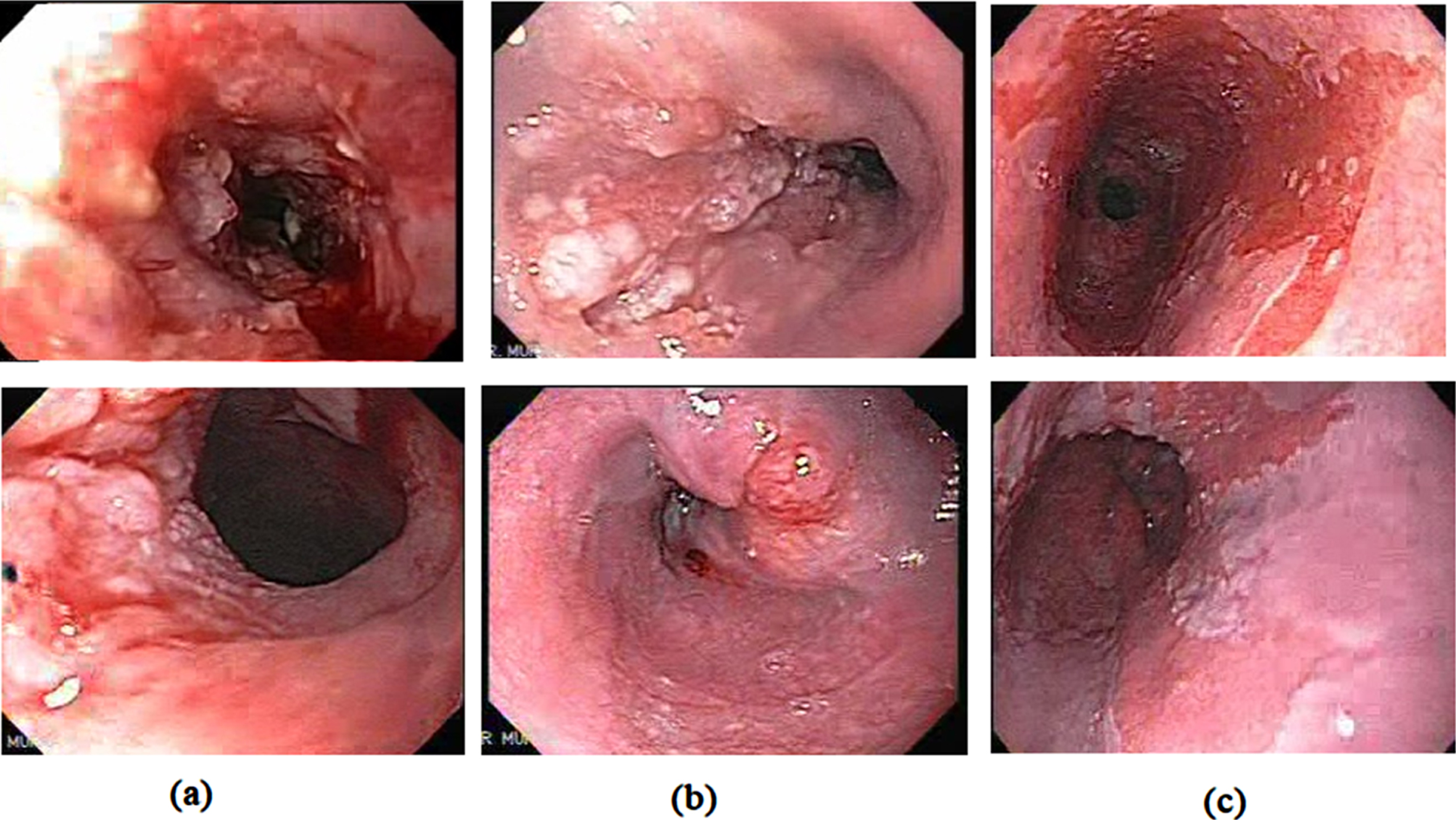
(a) EAC (b) ESCC (c) BE.
In the initial phase, the performance of the pre-trained networks is analysed by applying the transfer learning concept and fine-tuning the hyperparameters. These CNN models can also extract the feature maps from the dataset images. From the FC layers of the CNN models, we get 1000 features so that the feature matrix dimension becomes n×1000, where ‘n’ is the number of images in the dataset. Here in our work, we utilize the combination of networks for obtaining a larger feature map for a better analysis. A multi-CNN network with ‘m’ networks results in n×1000×m features. The proposed work’s best performing multi-CNN model consists of four networks, Xception, MobilenetV2, GoogLeNet, and Darknet53. The feature map of each of these CNN models is concatenated to generate the feature map of the multi-CNN model. The combined feature set consists of 4000 features for each image. Since we have many images and each has 4000 image features, we need to reduce the feature set for a more accurate analysis. For this purpose, we apply the wrapper-based ABC Optimization feature selection method. The image feature set gets reduced from 4000×1028 to 1972×1028 features. The optimal feature set is then fed as input to a multi-class SVM classifier with a polynomial kernel. Figure 2 shows the proposed model used in this study.
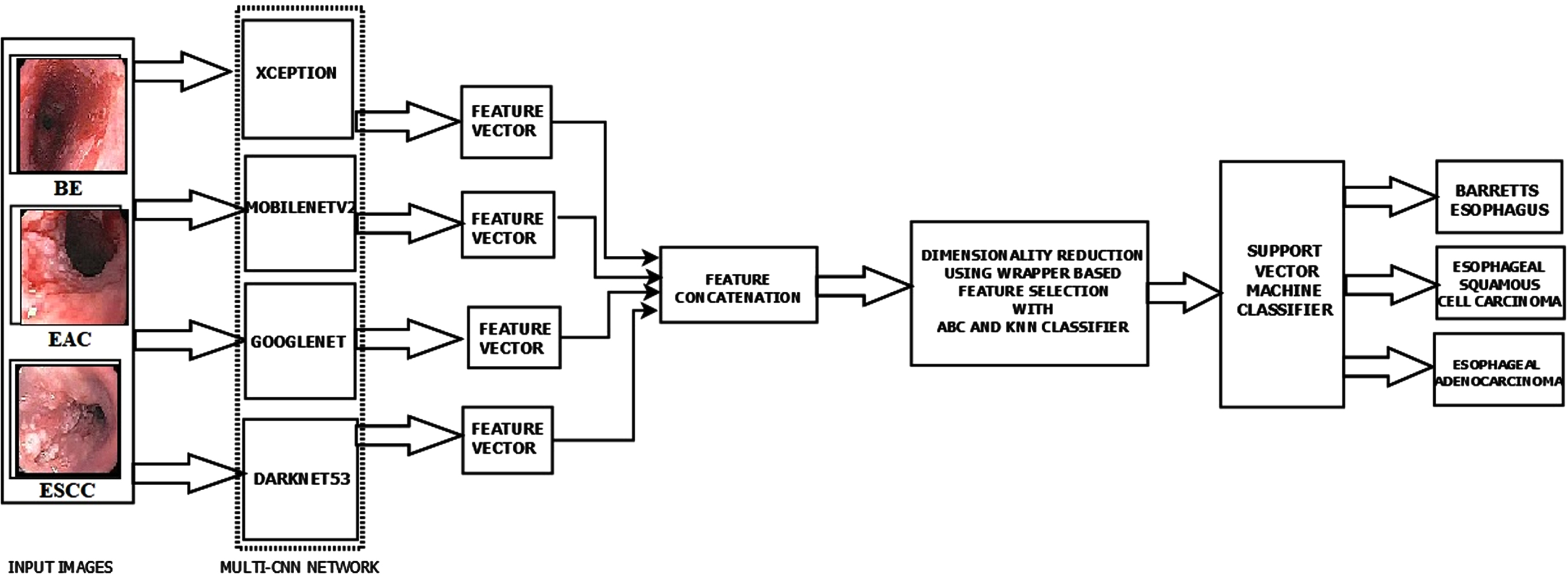
The proposed model architecture.
The images are resized to the input size of the corresponding pretrained network used for the analysis. Table 1 Illustrates the input size of the pretrained network used in this study.
Input size of the pretrained networks
Input size of the pretrained networks
The machine learning algorithms simulate the cognitive behaviour of a humanoid brain. The CNN networks extract the feature using a set of CNNs layers. More the number of layers deeper the network. The layers extract various attributes according to the dimension of the kernels used. These feature vectors can solve many problems by varying the kernel size accordingly. The various deep Learning models are generated based on the number of convolutional layers, pooling methods, and activation functions. The network starts with an input layer and ends with an output layer. We have the hidden layers between these layers that perform the feature extraction process. The Fully Connected (FC) layer of a CNN model contains all the attributes of the input image. The FC layer is tailed by a classification layer called Softmax classifier. The various DL models, ResNet50, DenseNet201, Nasnetmobile, Xception, Darknet53, Darknet19, MobilenetV2 and GoogLeNet are trained for detecting and classifying the precancerous stage BE and the malignant stages ESCC & EAC.
Usually, a large data set is required to train a CNN from scratch and extract the appropriate characteristics. To overcome this limitation, we apply the principle of transfer learning. Integrating information from an earlier occurrence that applies to a current activity is known as transfer learning. The acquired attributes are easily transferable to the new task by using fewer training datasets [46]. In this study, initially, we analysed the performance of the various pre-trained networks, ResNet50, DenseNet201, Nasnetmobile, Xception, Darknet53, Darknet19, MobilenetV2, and GoogLeNet, with the transfer learning technique for the classification. The pretrained models are not over or underfitted since we are using the 3-fold stratified cross validation. All the models are trained and then evaluated using the performance metrics in the initial stage. Based on its performance these models were used for feature extraction process. Further analysis of feature map fusion and feature selection process were performed. Finally, the classification is performed using machine learning models for obtaining better classification accuracy.
The Xception model is a pre-trained network with 170 layers, and the feature extractions are performed by the 36 convolutional layers. The Xception architecture comprises a sequence of depth-wise separable convolution layers with skip connections. So, the network becomes more flexible and easily adaptable. The input image size for an Xception network is 299×299×3, which generates about 22.8 million learnable parameters. MobileNet V2 model is a 154-layer pre-trained network with 53 convolution layers and one average pooling layer. MobileNetV2 pre-trained model has a residual block with stride one and a bottleneck block stride of 2 for downsizing. Both blocks comprise three layers: Initial layer consists of a 1×1 kernel that strides over the image and performs convolution with the pixel values followed by ReLu activation, then a depth-wise convolution layer. The third layer consists of a 1×1 convolution layer with no non-linearity. The input size compatible is 224×224×3. The Mobilenetv2 has 4.258 million learnable parameters. The GoogLeNet is 27 layers deep with max-pooling layers. The basic building block of GoogLeNet is an inception module. The GoogLeNet has 22 CNN layers, with 9 inception units. Inception module incorporates all possible filter size. The module has a CNN block with multiple filter size. The network introduces the 1×1 kernel layer, which reduces computation—every inception module act as a multi-level feature extractor. The GoogLeNet consists of auxiliary classifiers, which avoids the gradient vanishing problem. The network has an input size of 224×224×3 and has 4 million learnable parameters. DarkNet-53 is a 184-layer model with 53 convolutional neural network layers. The network-compatible input size of 256×256×3. DarkNet-53 forms the basis of all the object detection models, especially the YOLO network. The model has 41.025 million learnable parameters.
Feature selection
2.2.3.1. Wrapper-based method of feature selection The wrapper system is a supervised system of feature selection. Figure 3 shows the block level description of a wrapper system where the search algorithms find the relevant features and generate a feature subset [47]. The feature subset is then applied to the ML models that check the most significant attribute. This process is repeated for every iteration. The attribute with the lowest error value is added to the feature subset. For better results, simple algorithms such as sequential search or evolutionary algorithms like the nature-inspired algorithm, which offer optimal local outcomes and are computationally practical, are used to obtain good results. One of the significant limitations of wrapper techniques is the vast computational time required to produce the feature subset.
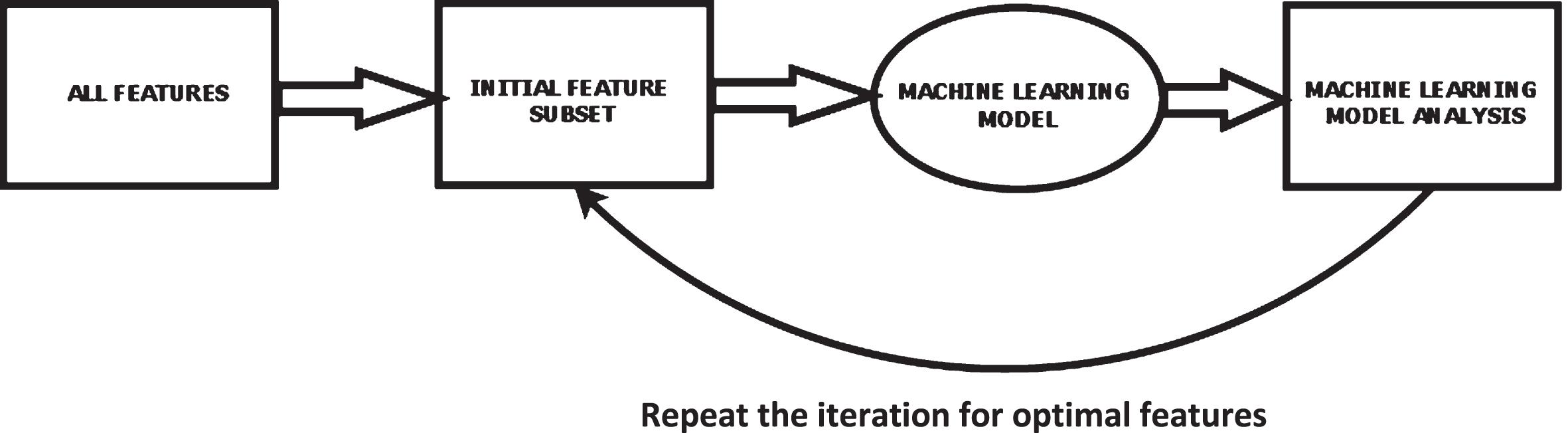
Block-level description of wrapper system.
Researchers have developed several attribute selection algorithms to generate the most relevant feature subsets to speed up the feature selection procedure. Search methods include greedy algorithms, floating search, step forward feature selection (SFS), heuristic search, step backward feature selection (SBS), duplex search, and random search [48]. SFS provides an improved balance between processing time and the eminence of the chosen feature subset than the other search algorithms. The SFS algorithm starts with a null set. The most relevant attribute is selected and fed to the classifier, which chooses the candidate feature. Again, the search for the next appropriate feature is performed, and the feature that improves classification accuracy is selected as the candidate feature. This process repeats if there is no further increase in the classification accuracy or no additional features are available [49]. Another limitation is the overfitting problem of the classifier used to improve the performance of the wrapper technique [50]. A suboptimal attribute subset with high accuracy and low power can arise when classification accuracy is used in subset selection. To overcome the biasing problem of the classifier, we use a hold-out validation method for the classification [50].
2.2.3.2. Artificial bee colony algorithm Many optimization algorithms are highly efficient and are designed to solve highly challenging optimization issues. Nature-inspired algorithms are a collection of innovative problem-solving methodologies and approaches inspired by natural phenomena. The Artificial Bee Colony (ABC) algorithm is a population-based meta-heuristic algorithm optimizing numerical problems [51]. It was inspired by the intelligent foraging behavior of honey bees. The bee’s intelligence to change its food position in accordance with the amount of nectar produced is conceptualized in ABC. The final position is the one with the most nectar. Artificial bees fly around in a multidimensional search space in the ABC system. Some (employed and observer bees) choose food sources based on their nest mates’ experience and modify their positions. Some (scouts) fly and seek food sources based on chance rather than experience. If the nectar amount of a new source is higher than the previous one in their memory, they memorize the new position and forget the previous one. As a result, the ABC system tries to balance hired and observer bees’ local search methods and onlookers’ and scouts’ global search methods. The position of the food source represents a feasible solution, and the amount of nectar refers to the quality or fitness of the solution for a problem. Similarly, the number of bees employed matches the number of solutions.
The general scheme of the ABC [52] algorithm is as follows steps: Initial random search for the food source within the bounded space. Search for the initial food position. Employed Bees finds a food source. Onlooker Bees checks the food source quality. Scout Bees go in search of new food resource. Memorize the best solution achieved so far.
UNTIL (Cycle = Maximum Cycle Number or a Maximum CPU time)
The general algorithm [53] of a basic ABC optimization method is: Define the random distributed initial population with SN solutions (food resource positions), where SN denotes the population size. Define a D dimensional vector for each random solution xi (i = 1, 2,..., SN). Repeated cycles of iteration take place in search of the solutions (the process of employed, onlooker bees) until the maximum CPU time. The solution is modified every time a better value is analyzed based on the nectar amount, which denotes the fitness of the solution given by;
If the fitness value is better than the previous one, the new value is stored by replacing the old one. The better fitness value is shared with the onlooker bees for the quality assessment. The quality of the nectar is based on the likelihood value correlated with that food resource, pi, The equation for updating of the memory using the new candidate position (best solution) is, The scout bees will check for the new food resources once it abandons the old food source. All the best solutions are kept in the D dimensional vector, and the iteration is repeated. On further iteration, if no better solutions are obtained over a predetermined number of iterations termed as ‘limit’, then that food resource is discarded.
Artificial bee colony algorithm was previously used successfully for a number of disease diagnosis. Sharma et al. [54] proposed a hybrid algorithm with PCA and ABC for the detection and classification of the leukaemia. Here ABC is used for selecting the relevant subset of attributes. There is a significant improvement in the classification accuracy when ABC is used along with reduced computation time. The classification accuracy of the back propagation neural network with PCA- ABC is 98.72% which is higher some of the other methods used. Zhang et al. [55] use a hybrid model with discrete wavelet transform, PCA, modified ABC with forward neural network for the classification of MRI brain images. An improved ABC with fitness scaling and chaotic theory was applied for the optimization process.
From among the various optimization methods used the improved ABC had a performance with reduced MSE and 100% classification accuracy. In Agarwal et al. [56] the classification performance of the various machine learning models is compared for the analysis of cervical cancer CT images. The ABC and SVM with gaussian kernel had a classification accuracy of 99% which is higher than the accuracy values with KNN and SVM linear kernel. Reddy et al. [57] proposed a hybrid model with DWT, ELM and ABC for the brain disorder classification. In this work the ELM-ABC performed better than the latest technology across various datasets used. In Alshamlan et al. [58] in his study provided a Support Vector Machine (SVM) based Artificial Bee Colony (ABC) technique for precise cancer microarray data classification. The suggested method (ABC-SVM) produces good results for gene selection and cancer classification when compared to other algorithms with an accuracy of 95.83% for Leukaemia and 95.6% for Colon cancer. Generally, the effect of the optimization algorithms for classification problems is very positive, especially ABC has got a slightly higher edge over another algorithm. Many uses modified ABC algorithms also for the better classification results.
Support vector machine (SVM) is a supervised ML classifier model with a very low probability of overfitting problems [59]. The SVM plots the input data points into a high dimensional space Z, and a hyperplane is constructed to separate the potential data points based on the class. The hyperplane should be positioned as close to the data points as possible.
Consider a set of data having n observations,
If ‘g’ denotes the weight vector of the data points and ‘b’ the initial bias of the classifier network, then the hyperplane will be denoted as
A mapping function δ can be used to translate the vector x in the original low-dimensional space into δ(x) in high-dimensional space.
In general, the Kernel function is represented as
Thus, in this model, a polynomial kernel SVM is used for performing the classification. The polynomial kernel is described as
SVM does not inherently enable multi-class classification. By default, it performs the classification of the labels into two groups. The same concept is employed to multi-class classification. The multiple classes are grouped into several binary classes. The class labels are separated by a hyper- plane linearly. This is known as a One-to-One approach, and it divides the multi-class problem into multiple two-class problems. For any binary class, there exists a binary classifier. If there are ‘n’ classes, there will be ‘n’ SVM models.
One-to-Rest is another strategy to investigate. Here there will be an SVM classifier per class. A single SVM can distinguish between two types and perform binary classification. So, according to the two breakdown methodologies, data points from the classes data set can be classified as follows: SVMs can be used by the classifier in the One-to-Rest technique. Each SVM would predict whether a person belongs to one of the classifications. SVMs can be used by the classifier in the One-to-One strategy.
Hence, in the proposed work the multi class SVM uses SVM binary learners and follows a one-versus-one coding design strategy for the classification process.
Experimental setup
The Matlab2019b tool extracts and blends the features in this work. The evaluation metrics considered are Accuracy, Recall, Precision, and F-Measure, derived from the confusion matrix.
The recall objective is to measure the percentage of true positives correctly identified.
Precision is a metric that indicates how accurate is the true positive found.
The model’s accuracy is the proportion of successfully identified data to the total number of classifications made.
The measure of true negative is termed as Specificity,
On a given dataset, the accuracy of the network model is quantified by the F1 Score. Most binary classifier models are evaluated using this metric, also called the F1 score. It is the arithmetic mean of the network model’s recall and precision.
We also use the Cohens Kappa (K-Score) and 95% Confidence interval (95% CI) for the analysis of the model.
The research methodology used is experimental in nature. In the experiments, based on the performance of the individual networks we made the various combinations, which was analysed for obtaining a better performance. From among these combinations, those with better performance metrics than the existing models were retained and few of the combinations having poor metrics were discarded. The proposed work’s best performing multi-CNN model consists of four networks, Xception, MobilenetV2, GoogLeNet, and Darknet53, whose feature maps are concatenated to generate a feature map of 4000×1028. Since performing the classification process with an extensive feature set is computationally complex, we apply the dimensionality reduction process using feature selection methods.
The combined feature set is subject to wrapper-based feature selection with the Artificial Bee Colony (ABC) Optimization method. The ABC algorithm operates by exploiting and exploring the intelligent behaviour of the bees to find the most feasible solution for a particular function. For this reason, we applied the ABC for feature selection in the proposed work. Every iteration, the process is repeated, and the attribute with the lowest error value is added to the feature subset. After the feature selection process, the feature set gets reduced to 1972×1028. The reduced feature map is applied to a polynomial kernel-based multi-class SVM to classify the various classes. The proposed model has an accuracy of 97.76%, a precision of 97%, a recall of 97.67%, specificity of 97.67%, F1score of 97.33% and Kappa Score of 0.9630. The 95% CI of the proposed model have a lower limit of 0.9492 and an upper limit of 0.9784.
When using the multi-CNN network model with ABC and multi-class SVM classifier, the classification is performed based on stratified 3-fold cross-validation outperforms all the other classifiers. The stratified 3-fold cross validation ensures that each fold consist of sufficient number of images from all the classes to ensure efficient training and testing of the model. The stratified cross validation also helps in handling the dataset imbalance effectively. For each fold there will be 343 test images and 685 train images, such that all the classes are there in equal proportion in both train and test images. This leads to a better classification among the various classes and hence better accuracy.
Table 2 shows that a combination of features extracted from multiple networks performs better than single CNNs. This is because representational power of features extracted from multiple CNNs are better than that of single CNNs. Here in [60] the author proposes a multi-CNN model for classifying the skin cancer using Inception3, Xception, VGG16, MobileNet, ResNet and others with different hyperparameters for achieving higher accuracy. In [61] the author proposes a rank-based ensemble model using GoogLeNet, VGG11 and MobileNetV3_small for classifying the breast cancer lesions using histopathology images. The cervical cancer diagnosis and classification is performed with an ensembled based CNN model with Inception V3, Xception and Densenet169.
Comparison of the performance of the multi-CNN models and the individual models
Comparison of the performance of the multi-CNN models and the individual models
The results show that the proposed multi-CNN model outperforms all the other models analysed [62]. In [63], an automated system using pre-trained ResNet deep neural networks, such as ResNet-50 and ResNet-101, are used for feature extraction in the categorization of skin lesions. The best features are chosen and used to a radial basis function (RBF) SVM for classification once the feature map has been fused. The AlexNet, DenseNet-201, and ResNet-50 models were employed as feature extractors in [64].The features are concatenated to provide a superior feature set for Covid-19’s severity classification using a Cubic Support Vector Machine (SVM).The major advantage of SVM is that the model performs well in the majority of medical imaging applications when the dataset size is medium and has an improved accuracy, classification speed, learning efficiency, and ability to handle overfitting issues for the majority of biomedical image-based classification applications [65]. Figure 4 depicts the confusion matrix of the proposed model.
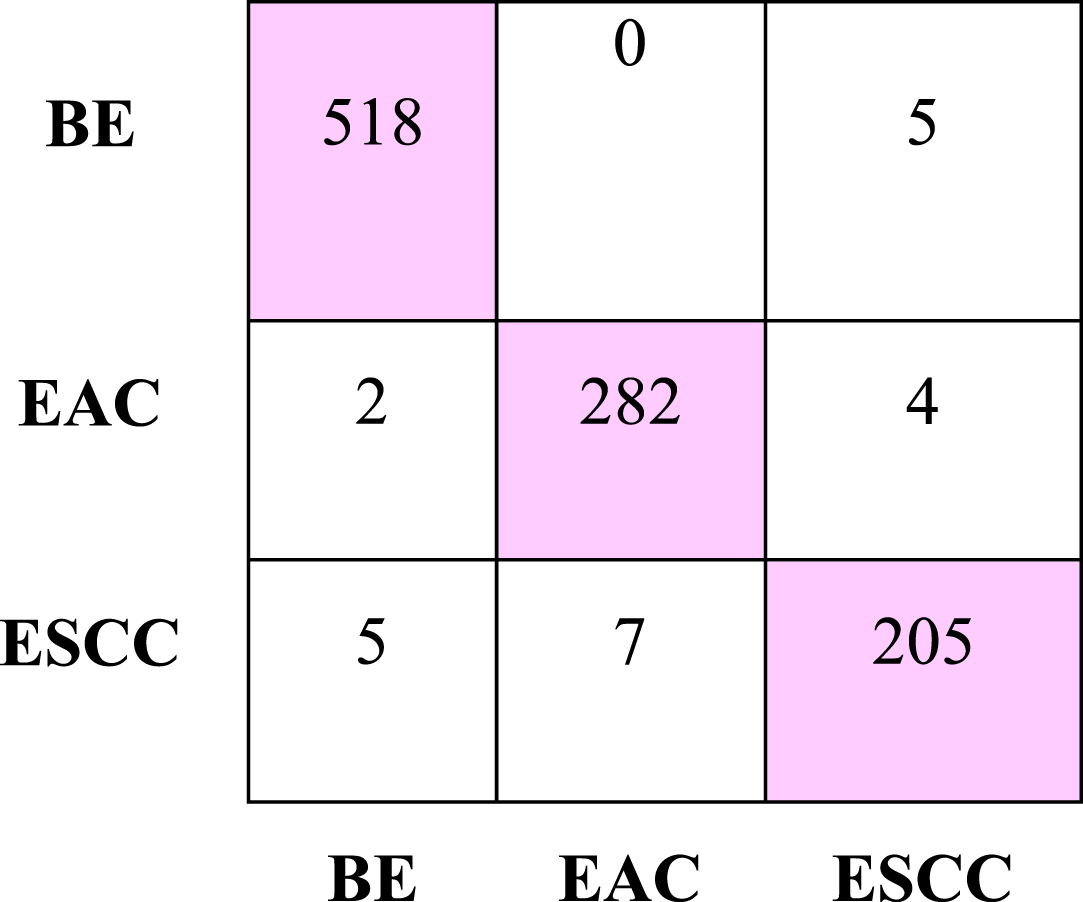
A confusion matrix generated by the proposed model.
The Multi-class Support Vector Machine Classifier achieved the highest accuracy of 97.76% and a recall of 97.67%. The Linear Discriminant Analysis classifier has an accuracy of 95.04% and recall of 94.33%. The K-means clustering classifier has accuracy of 94.65% and recall of 95%. The accuracy is 88.72%, for Random Forest with a recall of 89%. The Ensembled Tree and the Decision Tree classifiers have 84.53% and 81.91% accuracy with a recall of 84% and 78.33%, respectively. Naïve Bayes classifier achieved accuracy and sensitivity of 79.86% and 78.67%. We performed a stratified 3-fold cross-validation on the dataset for the classification. The results are displayed in Table 3.
Comparison of the performance of the classifiers for the proposed multi-CNN model
Comparison of the performance of the classifiers for the proposed multi-CNN model
The various pre-trained networks are initially trained with 3-fold cross-validation using the transfer learning technique. The pre-trained classifier models, ResNet50, DenseNet201, Nasnetmobile, Xception, Darknet53, Darknet19, MobilenetV2, and GoogLeNet, are trained with an epoch and a minibatch size of 6 and a learning rate of 0.0001 with Adam optimizer. An end-to-end training with transfer learning is performed on the various pre-trained network. The proposed model takes 662.27 sec, whereas the computational time for an individual pre-trained network may vary from 1200 sec to 5500 sec. The comparison of the various pre-trained networks with the transfer learning technique and the proposed model is shown in Table 4.
Comparison of the proposed model with the pre-trained CNN models with transfer learning
Comparison of the proposed model with the pre-trained CNN models with transfer learning
Ebigbo et al. [23] performed a tissue-based analysis to classify BE and EAC with a ResNet network using a LOOCV. With a precision of 78.06 % and recall of 77.58%, where the author in [24] proposed a DeepLabV3 + model for EEC segmentation and classification. In [26], the authors introduced an automated CNN model for diagnosing esophagitis, BE and EAC. By concatenating local features acquired by the Gabor filters and the DenseNet network, the computerized system with Faster RCNN enhances the performance. In [28], the author proposes a real-time AI-based CAD system with a ResNet-based DeeplabV3 to classify BE and Esophageal cancer. Here the last layers or the ResNet consist of dilated convolutions with spatial pyramid pooling for better classification. Zhan Wu et al. [37] proposed an Esophageal Lesion classification Network EL-Net for the automatic classification and segmentation of the various esophageal lesions. The Dual stream network performs the classification of the lesions into normal, Esophagitis, Barrett, and Cancer. The classification is done by using the global and local features of the lesion images. Liu et al. [38] introduced the Inception- ResNet model, classifying precancerous tissue and cancer tissues with 85.83% accuracy. Du et al. [40] proposed a deep learning model for classifying four major EC stages starting from BE to EAC. The model achieved a classification performance with an accuracy of 90.63%, precision of 89.71%, recall of 89.26%, and F1score of 89.45%. Mohammad et al. [42] proposed a fuzzy C means-based method with morphological analysis with three fully connected hidden layers to detect esophageal cancer. Yu et al. [43] proposed a multi-task deep learning model to diagnose Esophageal lesions. The proposed model with the multi- CNN and ABC optimization technique with SVM has a better result than all the above methods in classifying precancerous stage BE, squamous cell carcinoma (SSC), and advanced cancer stage (EAC). Table 5 depicts the comparison of the performance of the proposed model with the existing methods.
Comparison of the performance of the proposed model with the existing methods
Comparison of the performance of the proposed model with the existing methods
The comparison of the proposed model with the some of the popularly used nature inspired optimization methods is illustrated in the Table 6. From among these the CNN combination with the ABC algorithm outperformed all the other optimization methods analysed in the work.
Comparison of the performance of the proposed model for the various optimization methods
Comparison of the performance of the proposed model for the various optimization methods
In the proposed work, the model classifies the stages of EC into three classes namely, BE, ESCC and EAC. Here in the 3-class problem, the proposed model is a multi-CNN model generating a large feature map to analyse the problem. The multi-CNN model with a better initial performance consists of Xception, MobilenetV2, GoogLeNet, and Darknet53. The feature map of each of these CNN models is concatenated to generate the feature map of the multi-CNN model. The combined feature set consists of 4000 features for each image. Since we have many images and each has 4000 image features, we need to reduce the feature set for a more accurate analysis. For this purpose, we use a wrapper -based ABC optimization method selects the relevant attributes so that the image feature set gets reduced to 1972×1028 features. The classification is performed using the optimal feature set mullti-class SVM classifier with a polynomial kernel. The proposed AI-based model has a precision of 97%, a recall of 97.67%, specificity of 97.67%, an accuracy of 97.76%, an F1score of 97.33% and a K score of 0.9630.
Artificial bee colony algorithm was previously used successfully for several disease diagnosis. In the hybrid model proposed by Sharma et al. [54] ABC provided significant improvement in the classification accuracy (98.72%) and reduced computation time. For Zhang et al. [55] hybrid model for the classification of MRI brain images the improved ABC had a performance with reduced MSE and 100% classification accuracy. In Agarwal et al. [56] the ABC and SVM with gaussian kernel had a classification accuracy of 99% for the cervical cancer CT dataset. The ELM-ABC model proposed by Reddy et al. [57] performed better than the latest technology across various datasets used for brain disorder classification. Alshamlan et al. [58] proposed an ABC-SVM based model which produces good results for gene selection and cancer classification with an accuracy of 95.83% for Leukaemia and 95.6% for Colon cancer. Generally, the effect of the optimization algorithms for classification problems is very positive, especially ABC has got a slightly higher edge over another algorithm. Many uses modified ABC algorithms also for the better classification results. The proposed model takes 627.10 sec, whereas the computational time for an individual pre-trained network may vary from 1200 sec to 5500 sec. The 95% CI of the proposed model have a lower limit of 0.9492 and an upper limit of 0.9784.
In the proposed model the ABC is used for the feature selection to avoid redundancy and to obtain the most relevant attributes. The ABC selects 1972 features, and the proposed model performs the classification. The computational time taken for the process is 662.270 sec. from Table 6 when compared the PSO performs with the least computational time of 538.24 s with 1913 features selected. But the ABC has a better performance based on the evaluation metrics, even though the variation is a marginal one. When compared with GA and ACO with 1772 and 1972 features selected with a processing time of 1375.18 and 1703.5 sec the ABC outperforms both the optimization methods. Similarly, when compared with ABO and ACS both have 760 and 476 features selected, the processing time for model to classify the images is very high, such as 728.77 and 1708.4 seconds, respectively. In addition, in this context also the ABC with 1972 relevant attributes ensures a better performance metrics than the ABO and ACS. Thus, to conclude we can say that for a larger FS set the ABC have a upper hand both in the performance as well as the time complexity. Figure 5 depicts the graphical analysis of the performance of the various optimization methods and its time complexity, and the number of features selected.
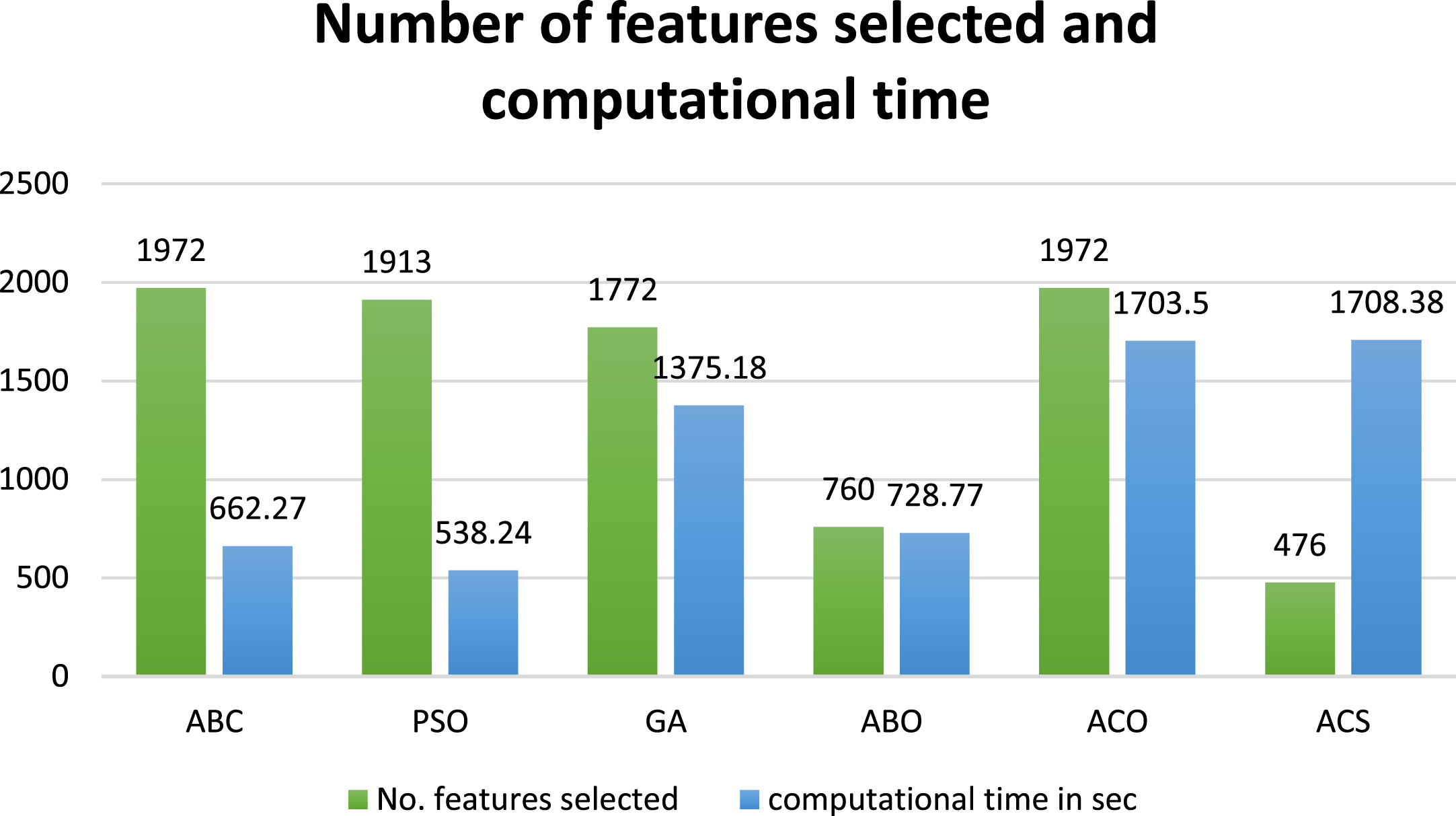
Performance of the various optimization methods based on the no. of features selected and computational time.
The domain of medical image processing and deep learning is one of the essential fields for a variety of reasons, including assisting doctors in disease diagnosis, increasing performance speed and accuracy, and, most importantly, lowering EC fatalities. An efficient deep learning model is required to enable a CAD system to serve as “a second observer” in an endoscopic examination to assist non-experts in detecting EC and reduce missed diagnoses. The CNN models are nowadays used popularly for the medical image analysis. This is because these models are good in learning the low level and high-level semantic features of the input image [66]. The different CNN models learn different levels of feature and accordingly represent the image. So, an ensemble CNN model extracts high quality feature for a better analysis and representation of the image class. The CNN model when finetuned can extract more relevant attributes for a better classification of the images [67]. So, an ensemble finetuned CNN model can generate more powerful framework for the medical image analysis.
One of the major limitations is the lack of endoscopic images from all types of gastroscopic systems. Another limitation is the smaller number of images in the publicly available datasets. Kvasir [44] and MICCAI 2015 [68] are the primary publicly available research datasets. The small dataset and the cost of labelling always pose a drawback for the studies. Due to these factors, more studies will be conducted using real-time videos. Most of the work is performed using supervised learning methods requiring a large training dataset. So, we need to consider self-supervised and semi-supervised learning methods to train the networks. The accuracy and quality of the endoscopic camera are a factor to consider. All the above factors must be considered for developing a more precise and well-performing CAD system.
In the proposed work, the efficacy of the multi - CNN pre-trained network is analysed to classify the precancerous stage BE and the malignant stages ESCC and EAC. The fusion of feature maps obtained from the CNN models is considered for the analysis. A wrapper-based method with the Artificial Bee Colony optimization is utilized for the feature selection in the proposed method. The multi-class SVM classifier is employed to classify the desired classes for the multi-class classification. The multi-CNN network combination with Xception, MobilenetV2, GoogLeNet and Darknet53 outperformed all the other networks. The multi-CNN network model with ABC and the multi-class SVM machine learning model for classification achieved an accuracy of 97.76%. The results prove the efficiency of the multi-CNN model for the classification of EC stages. In the future, more network combinations with new attribute selectors and classifiers need to be analysed for the diagnosis of EC stages.
