Abstract
The significant increase in data volume in recent years has prompted the adoption of knowledge graphs as valuable data structures for integrating diverse data and metadata. However, this surge in data availability has brought to light challenges related to standardization, interoperability, and data quality. Knowledge graph creation faces complexities from large data volumes, data heterogeneity, and high duplicate rates. This work addresses these challenges and proposes data management techniques to scale up the creation of knowledge graphs specified using the RDF Mapping Language (RML). These techniques are integrated into SDM-RDFizer, transforming it into a two-fold solution designed to address the complexities of generating knowledge graphs. Firstly, we introduce a reordering approach for RML triples maps, prioritizing the evaluation of the most selective maps first to reduce memory usage. Secondly, we employ an RDF compression strategy, along with optimized data structures and novel operators, to prevent the generation of duplicate RDF triples and optimize the execution of RML operators. We assess the performance of SDM-RDFizer through established benchmarks. The evaluation showcases the effectiveness of SDM-RDFizer compared to state-of-the-art RML engines, emphasizing the benefits of our techniques. Furthermore, the paper presents real-world projects where SDM-RDFizer has been utilized, providing insights into the advantages of declaratively defining knowledge graphs and efficiently executing these specifications using this engine.
Introduction
Advancements in data collection devices and methods, such as sensors, wearables, and genomic tests, have resulted in the generation of vast amounts of heterogeneous data, including omics and patient health data. This data is available across various organizations and companies, such as hospitals, universities, and pharmaceuticals. However, the presence of data silos often hampers the combination, analysis, and reuse of this valuable data, which prevents the discovery of insights essential for decision-making. To address this challenge, knowledge graphs (KGs) have gained significant traction in both industrial and academic sectors [32]. KGs provide a unified representation of heterogeneous data, enabling the convergence of data and their meaning. KGs can be defined as data integration systems consisting of a unified schema, data sources, and mapping rules that establish correspondences between the data sources and the unified schema. These declarative definitions of KGs empower modularity and reusability while also allowing users to trace the process of KG creation. Therefore, KGs serve as expressive data structures for modeling integrated data and metadata, and their declarative specifications can be explored, validated, and traced, thus, enhancing transparency and maintenance.
The Semantic Web community has played a relevant role in addressing the challenges associated with integrating heterogeneous datasets into KGs. To tackle this complex task, the community has actively contributed with methodologies, formalisms, and engines aimed at facilitating the creation and maintenance of KGs [6,38,54]. Declarative mapping languages, such as R2RML [21], RML [24], and SPARQL-Generate [40], have emerged as powerful tools within this context. These languages allow knowledge engineers to define mapping rules or assertions [34,44] capable of generating KGs expressed in RDF2 through systematic evaluations. Mapping rules enable the seamless definition of concepts within a unified schema, encompassing classes, properties, and attributes. This is achieved by harnessing data from diverse sources presented in various formats, including tabular, CSV, JSON, or RDF. The use of declarative mapping languages significantly enhances the flexibility and efficiency of the KG creation process. By providing a standardized approach to mapping data to RDF, these languages empower organizations to extract valuable insights from disparate sources, thereby driving informed decision-making. Furthermore, the Semantic Web community has actively contributed to the development of various engines designed to execute mapping rules [10,17,18,48].
Various RML engines are available such as Morph-KGC [3], RMLMapper,3 RocketRML [57], and SDM-RDFizer v3.2 [33]. These engines have implemented techniques to execute RML mapping rules efficiently. However, given the variety of parameters that can affect the performance of the KG creation process [15], existing engines may not scale up to KG creation pipelines defined in terms of complex mapping rules or large data sources. Given the amount of available data, new methods are demanded to scale up to complex scenarios where heterogeneous data sources need to be integrated to provide the basis for knowledge analysis and decision-making.
In this paper, our primary research objective is to address the challenge of KG creation through declarative specifications using RML. We present data management techniques implemented in the latest version of SDM-RDFizer, denoted as SDM-RDFizer v4.5.6. These techniques play a pivotal role in satisfying the requirements of data collection and processing within complex data integration systems. Specifically, they enable the efficient scaling up of KG creation pipelines in real-world scenarios characterized by large and heterogeneous data sources. The significance of these data management techniques is underscored by their effectiveness in handling complex KG creation scenarios encountered in practical use cases. These scenarios are exemplified by the testbeds proposed in the Knowledge Graph Construction Workshop 2023 Challenge at ESWC 2023 [63] and are characterized by parameters reported by Chaves-Fraga et al. [15]. These complex use cases span different domains, such as biomedicine [2,52,59] and energy [37], where the declarative definition of KGs facilitates the smooth integration of large amounts of heterogeneous data, potentially duplicated across different data sources. As a result, SDM-RDFizer v4.5.6 is empowered to scale up data collection, processing, and integration towards efficient KG creation.
This paper extends the work reported by Iglesias et al. [33]. Our new contributions are as follows:
Data structures for RDF data compression and planning techniques for mapping rule execution. Physical operators that leverage these data structures to efficiently handle complex RML mappings, including multiple joins and large data sources with high duplicate rates. A new version of the SDM-RDFizer tool (named SDM-RDFizer v4.5.6) which incorporates these data structures and physical operators, enabling the execution of complex KG creation pipelines. An empirical evaluation using two state-of-the-art benchmarks, GTFS-Madrid-Bench [16] and SDM-Genomic-Datasets.4 Our evaluation encompasses 416 testbeds and compares our new version with Morph-KGC, RMLMapper, RocketRML, and the previous version of SDM-RDFizer (v3.2). The results demonstrate the advantages of the data management techniques proposed in this paper and implemented in SDM-RDFizer v4.5.6. Specifically, the evaluation highlights the significance of data structures and physical operators in executing complex configurations like those found in the SDM-Genomic-Datasets benchmark.
The rest of the paper is structured as follows: Section 2 defines and illustrates a KG creation pipeline and the requirements to be satisfied for an RML engine. Section 3 summarizes previous approaches. Section 4 defines the data management methods implemented in SDM-RDFizer v4.5.6. Section 5 reports the results of our experimental studies. Section 6 describes the main characteristics of our tool and, finally, our conclusions and future work are outlined in Section 7.
This section provides the basis for understanding the problem of KG creation. First, we define some basic concepts related to KGs, data integration systems, KG creation pipelines, and RML (RDF Mapping Language). Next, we present the steps involved in declaratively specifying a KG using RML. To ensure the effectiveness of RML engines, we will elucidate the requirements that they need to satisfy, based on existing evaluation studies reported in the literature [15,34,39]. Finally, we will illustrate the problem of KG creation in a use case derived from the data integration challenges in the biomedical area. This example illustrates data integration issues reported by Chandak et al. [12] and observed in the data management tasks of the EU H2020 funded projects iASiS5 and CLARIFY.6
Preliminaries
Data can be siloed and scattered across various data sources. As the volume of available data continues to expand, the prevalence of these data silos is expected to increase, consequently hindering interoperability. Data integration aims to gather information from heterogeneous sources and provide a unified view from which relationships and patterns hidden in isolated sources can be uncovered [11,28]. To illustrate, consider data related to genes collected from tissues from tumors and stored in three data sources. A relational database maintains gene-related information (e.g., created from Genecards7). Further, the results of tumor tissue analysis are maintained in a tabular format (e.g., CSV), and an XML file stores information about the tumors from which the tissues were sampled. When analyzing genes (e.g., BRCA1) related to tissues sampled from specific tumors (e.g., breast tumors), integrating these data sources becomes essential to establish the connections existing between these entities. However, each data source follows a different format and schema, creating interoperability conflicts. Data integration has the objective of providing data management methods to create these holistic views (e.g., integrated views of genes, tissues, and tumors). While these interoperability issues are readily apparent in the biomedical field [39], similar issues persist across industrial and scientific domains where data is autonomously generated. The challenge is to provide data management mechanisms that enable seamless data integration while also ensuring scalability and maintainability to guarantee long-term usability and value of the integrated data.
A data integration process can be specified as a data integration system
KGs are directed edge-labeled graphs that represent statements as entities and their relationships as labeled edges (Gutierrez et al. [29]). Given a KG
R2RML mappings define how tables, columns, and rows in a relational database are declaratively translated into RDF triples. RML is a mapping language that represents mapping rules in RDF; it offers the features of R2RML by allowing the specification of mapping rules over heterogeneous data sources, e.g., JSON, CSV, and XML. RML provides a well-established and standardized approach to mapping heterogeneous data to RDF. The W3C Community Group on Knowledge Graph Construction has proposed the RML ontology [36] as a common agreement on how the mapping rules must be defined. This standardization ensures consistency and compatibility with other RDF-related tools and technologies. Moreover, RML mappings can be reused for different data sources, reducing redundancy in mapping definitions. This reusability streamlines the creation and maintenance of RDF KGs for various applications. Declarative RML mappings are represented in RDF, serving as metadata that can be queried to facilitate the maintainability and reusability of DISs. Consequently, these declarative specifications of KGs as RML DISs promote modularity, supporting maintenance, testing, debugging, and enabling collaborative definition and reusability. RML is currently being used by several companies and public organizations [19,20,50,60] to create their KGs, also including the Google Enterprise Knowledge Graph.9
Fig. 1 depicts RML mapping rules (a.k.a. triples maps). Triples map represents mapping rules where the resources (

The declarative definition of a KG Description of the schema of the data sources Selection of the set For each class Data type properties If The object values of an object property
Requirements of a knowledge graph creation pipeline
Requirements for RML engines are defined based on: parameters analyzed by Chaves-Fraga et al. [15]; testbeds assessed by the KGCW 2023 Challenge [63]; and data integration requirements presented by Kinast et al. [39].
Chaves-Fraga et al. [15] have established that various parameters influence the efficiency of declaratively specified KG creation as a DIS. These parameters are grouped into five dimensions: mapping, data, platform, source, and output. The
The Extended Semantic Web Conference (ESWC) 2023 hosted a Knowledge Graph Creation Workshop (KGCW), which introduced a challenge dataset [63] aimed at evaluating the performance of existing KG creation RML engines. This dataset was designed to assess memory usage and execution time, considering various parameters that influence the KG creation process, as established by Chaves-Fraga et al. [15]. The aim was to create multiple test cases by generating RML triples maps covering a wide range of scenarios. They include: a) size of the data sources based on number of rows and columns; b) number of RML triples maps and their properties; c) complexity of joins among RML triples maps; and d) data diversity (e.g., duplicate rate and empty values). These scenarios each have unique effects on the KG creation process. Data source size and the complexity of joins primarily impact memory usage. Meanwhile, the number of properties, triples maps, and data diversity can influence execution time. How KG creation engines handle these variables determines the specific impact on execution time or memory usage due to data diversity. In summary, this challenge dataset offers a comprehensive examination of KG creation scenarios, making clear the requirements that need to be satisfied to enhance the performance of RML engines.
Kinast et al. [39] present the outcomes of a systematic literature analysis, showing the functional requirements for integrating medical data. These functional requirements include several categories, encompassing data acquisition, processing, analysis, metadata management, traceability, lineage, and security. While some requirements are specific to the medical domain, the following are domain-agnostic: i) the capability to collect data in various formats; ii) the use of standardized ontologies, vocabularies, rules, and processes; iii) the possibility of representing integrated data through multidimensional models (e.g., RDF KGs); and iv) the capacity of managing large volumes of data.
We have elucidated the following requirements; they are divided into two categories: data collection and processing.
This section illustrates the problem of defining a KG using a declarative approach where mapping rules are specified in RML. In this example, we focus on a portion of a DIS that defines a biomedical KG [2,52,64,65], e.g., the one created in the context of the EU H2020 projects iASiS5 and CLARIFY6. The data sources, referred to as SDM-Genomic-Datasets [33] vary in size, with 100k, 1M, and 5M rows, and each dataset has a different percentage of data duplicate rate (25% or 75%). The unified ontology10 consists of classes like
Three state-of-the-art RML engines, i.e., RMLMapper v6.0,11 Morph-KGC v2.1.1,12 and SDM-RDFizer v3.228, are utilized to create this portion of the KG; following configurations reported in the literature [13,17,34,35], the engines timed out in five hours. The results of executing these engines in six testbeds are shown in Fig. 2. The execution time was significantly impacted by factors such as the size of the data sources, the percentage of duplicates, and the number and type of joins among triples maps. In fact, two out of the three engines timed out when processing a data source with 5M records. Although SDM-RDFizer exhibited relatively better execution time compared to RMLMapper and Morph-KGC, it still required considerable time to create the KG. In this paper, we present a new version of SDM-RDFizer (v4.5.613) that incorporates data management techniques for planning the execution of triples maps and efficiently compressing intermediate results. These improvements have enabled SDM-RDFizer to scale up to complex scenarios, as reported in Section 5.

This section summarizes the key contributions from the existing literature regarding creating RDF KGs. First, an overview of the data models, formats, and frameworks to transform Web data is presented. Then, existing technologies for KG creation are described, while the following two sections present the main approaches for performing the KG integration process in a virtual or materialized manner [6].
Representing data on the web and transforming web data into RDF
The adoption of the Web as a framework for publishing electronic data [1] has driven the development of semi-structured data models, formats, and languages aimed at facilitating their curation, retrieval, and version control [1]. In this context, XML emerged as a standard proposed by the World Wide Web Consortium (W3C) for representing semi-structured data from diverse sources. XML employs a tag-based syntax that is easily readable by both humans and machines. Additionally, RDF graphs can be represented using XML syntax, with tools like XSPARQL [9] and Gloze [8] supporting the transformation between XML and RDF specifications. CSV, an abbreviation for “Comma Separated Value,” is another commonly used format for representing and exchanging data on the Web. The CSV2RDF14 framework offers standardized procedures for converting CSV data into RDF, while Tarql15 relies on wrappers for CSV data sources. These wrappers enable the execution of CONSTRUCT SPARQL 1.1 queries, facilitating the creation of RDF graphs from CSV data. Relational or tabular data also serve as a prevalent data model for presenting information on the Web. Tools developed by Polfliet and Ryutaro [47], Sequeda and Miranker [56], as well as Auer et al. [7], exemplify solutions for transforming relational data into RDF format. In a broader context, Thakker [61] offers a comprehensive analysis of various data transformation techniques applied to data represented in different models (such as relational or semi-structured databases) and formats (e.g., XML16 or JSON17). These transformations are executed using query languages like SPARQL18 or Gremlin.19 This analysis emphasizes the significance of this topic within both the database and semantic web communities. SDM-RDFizer also facilitates the transformation of data from diverse sources represented in various formats, including JSON, XML, CSV, or relational databases. However, SDM-RDFizer relies on declarative definitions using R2RML or RML to establish correspondences between data sources and RDF. Since different factors can influence the execution of these declarative mapping rules [15], SDM-RDFizer employs various data structures and physical operators to mitigate the impact of data size, mapping rule complexity, and duplicate records.
Knowledge graph creation and existing technologies
The creation of a KG
Virtual data integration in knowledge graphs
The creation of a virtual KG involves generating it dynamically based on a request expressed as a query over a target ontology, with mapping rules used to transform the input query into an equivalent query for the source(s) in
Materialized knowledge graph creation
A KG is constructed by integrating various sources in
SDM-RDFizer: A tool for the materialized creation of knowledge graphs
This section presents the SDM-RDFizer tool, focusing on its architecture and data management techniques that enable the engine to meet the requirements outlined in Section 2.3. The development of SDM-RDFizer follows the Agile software development methodology, which ensures a flexible and iterative approach guided by the requirements for scaling up KG creation. These requirements are gathered from various sources, including the community (e.g., the KGC Challenge at ESWC 2023), ongoing projects, and fundamental findings from the data management field. By embracing the Agile methodology, the development team can effectively adapt to evolving needs and prioritize scalability. The iterative nature of Agile allows for continuous improvement and integration of valuable feedback throughout the development process. Moreover, by actively involving stakeholders and considering their input, SDM-RDFizer aims to address the specific challenges faced by users in KG creation. Through the Agile approach, SDM-RDFizer strives to deliver a more responsive, collaborative, and efficient tool that effectively tackles the demands of scaling up KG creation.
The SDM-RDFizer architecture
SDM-RDFizer implements multiple data structures that optimize different aspects of the KG creation process such as duplicate removal, join execution, and data compression, and operators that execute various types of triples maps efficiently. Additionally, SDM-RDFizer is able to plan the execution of the RML triples maps to reduce execution time and secondary memory consumption.
Fig. 3 depicts the SDM-RDFizer architecture in terms of its components. The SDM-RDFizer comprises two main modules:

The Triples Map Planning (TMP) module reorders RML triples maps so that the most selective ones are evaluated first, while non-selective rules are executed at the end. TMP organizes the triples maps and data sources so that the number of RDF triples kept in the main memory is reduced. As a result, the KG creation process consumes the minimum amount of memory and execution time. TMP defines two data structures, the
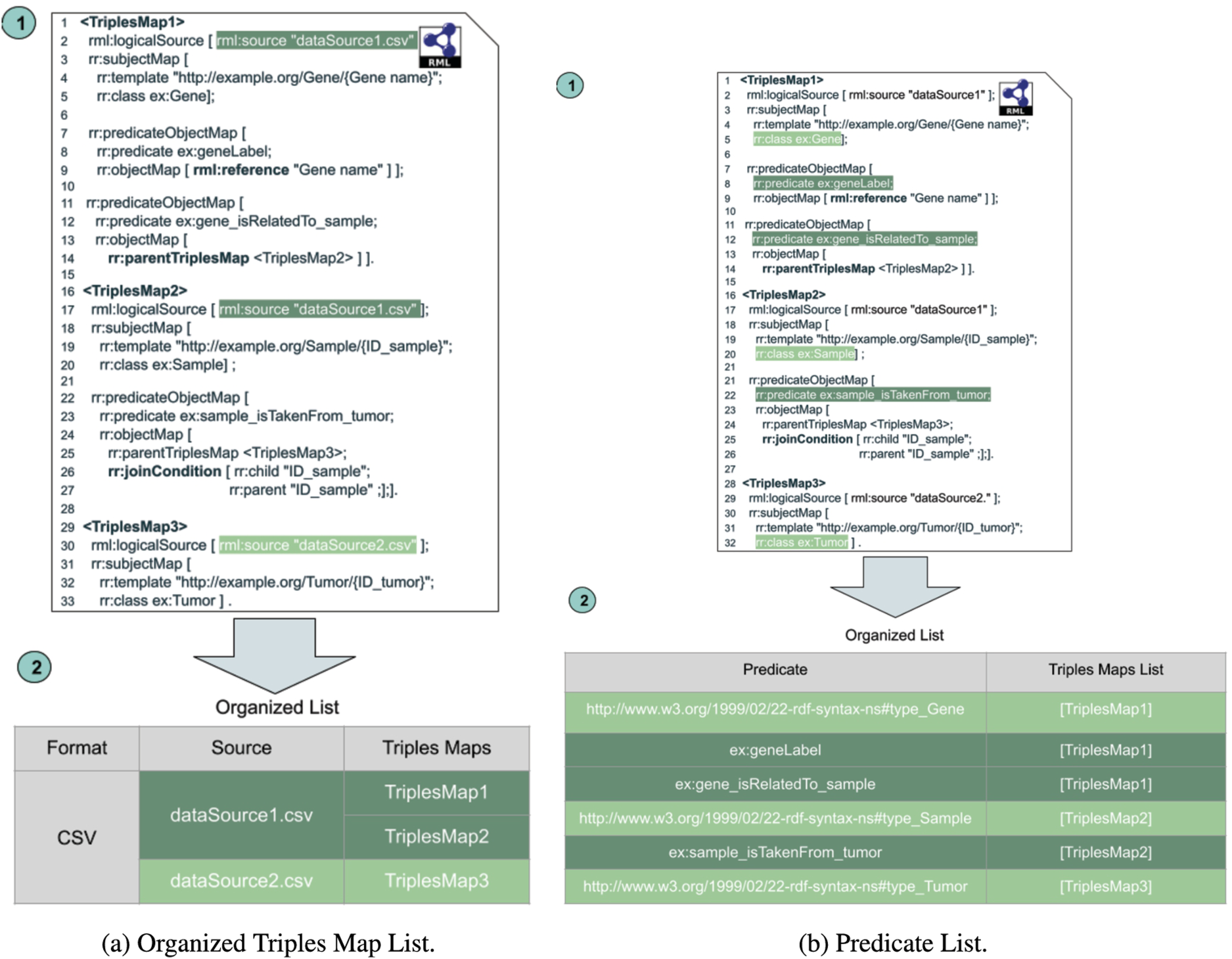
During the TMP phase, triples maps are classified based on the logical data source format (i.e., CSV, JSON, and XML). Afterward, they are grouped by their data source; thus, a data source is opened once to execute all the triples maps. Implementing this data structure into the SDM-RDFizer causes the processor to adopt a hybrid approach. A data-driven approach is used for triples maps with file data sources, while a mapping-driven approach is used for triples maps with relational databases. Figure 4a depicts the OTML for the triples maps in Fig. 1. Since all these triples maps are over CSV files, only one group is created.
This section explains the main data management methods implemented in SDM-RDFizer. The Triples Map Execution (TME) module generates the KG; it follows the order established by the TMP module when executing the RML triples maps. TME introduces multiple data structures,
The SDM-RDFizer data structures
SDM-RDFizer implements three data structures to efficiently manage and store the intermediate RDF triples generated during the execution of RML triples maps. These data structures avoid the generation of duplicated triples. Intermediate results are stored in these data structures independently of the format of the data sources. Thus, SDM-RDFizer exhibits a performance agnostic of the format of the data sources.


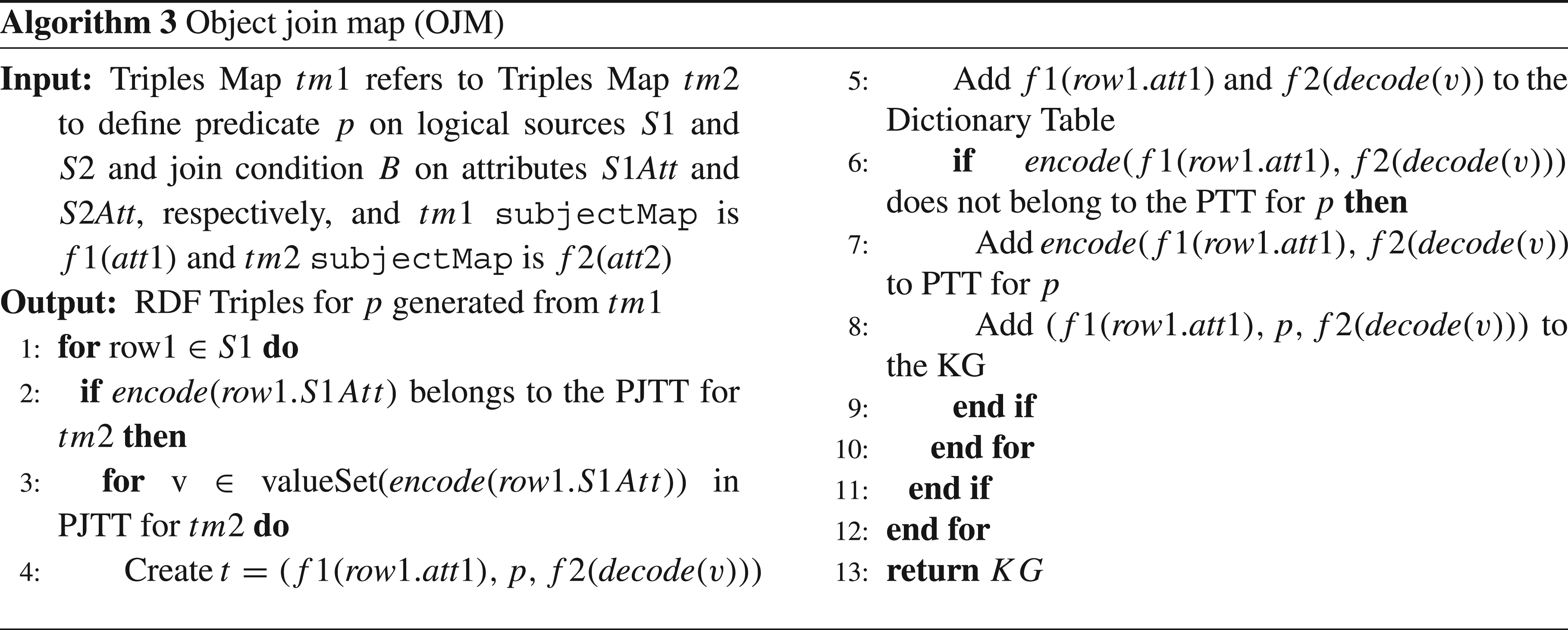
SDM-RDFizer resorts to three physical operators to efficiently execute triples maps in a KG creation pipeline.


Requirements for data integration and SDM-RDFizer
RML allows for the definition of mapping rules over structured data (e.g., CSV and relational databases) and semi-structured data (e.g., XML and JSON). SDM-RDFizer is an RML-compliant engine that can process all data source formats that RML covers, fulfilling RE1-Heterogeneous data and RE6-Semi-structured data. Additionally, by being able to process RML mapping and their corresponding data sources, RE7-Standardized data integration has also been covered. SDM-RDFizer is developed on Python, and given the nature of the programming language, there is no hard cap on how much memory a process can consume (only being limited by the environment in which the engine is executed), which allows data sources of all sizes to be processed, thus fulfilling RE2-Large data. The engine can also upload all the corresponding attributes of the data sources, hence covering RE3-Fragmented data. SDM-RDFizer implements multiple data structures that optimize different aspects of the KG creation process, like duplicate removal and join execution. In the case of duplicate removal, the engine presents PTT, a hash table that stores all the triples generated by that point in time, and all new triples are compared to the corresponding PTT to determine if it is a duplicate. The triple is discarded if it is a duplicate, thus fulfilling RE4-Duplicated data. Finally, the engine introduces three operators (SOM, ORM, and OJM), each transforming a different type of mapping rule, therefore proving RE8-Mapping complexity. Furthermore, the OJM uses a data structure called PJTT; it stores the result of executing a join between two triples maps, so executing the same join multiple times is unnecessary, thus proving RE5-Data diversity.
Empirical evaluation
This section presents the main results of the experimental evaluation conducted on SDM-RDFizer, aiming to address the following research questions:
To provide a comprehensive overview of the empirical assessment and the observed results, this section includes the definition of the experimental configuration. This configuration encompasses the selection of benchmarks, metrics, engines, and the description of the experimental environment used to evaluate state-of-the-art RML engines. Each experimental configuration is repeated five times, and the average execution time is reported as the outcome. The obtained results are carefully analyzed to identify the strengths and weaknesses of SDM-RDFizer in comparison to other engines.
Experimental settings
Datasets and configurations of triples maps. The table describes each data source and configuration of TMs used in the experiments and their corresponding benchmarks. Configuration of TMs in bold are considered complex cases. They include several types of TMs of various complexity and complex joins (e.g., five-start joins)
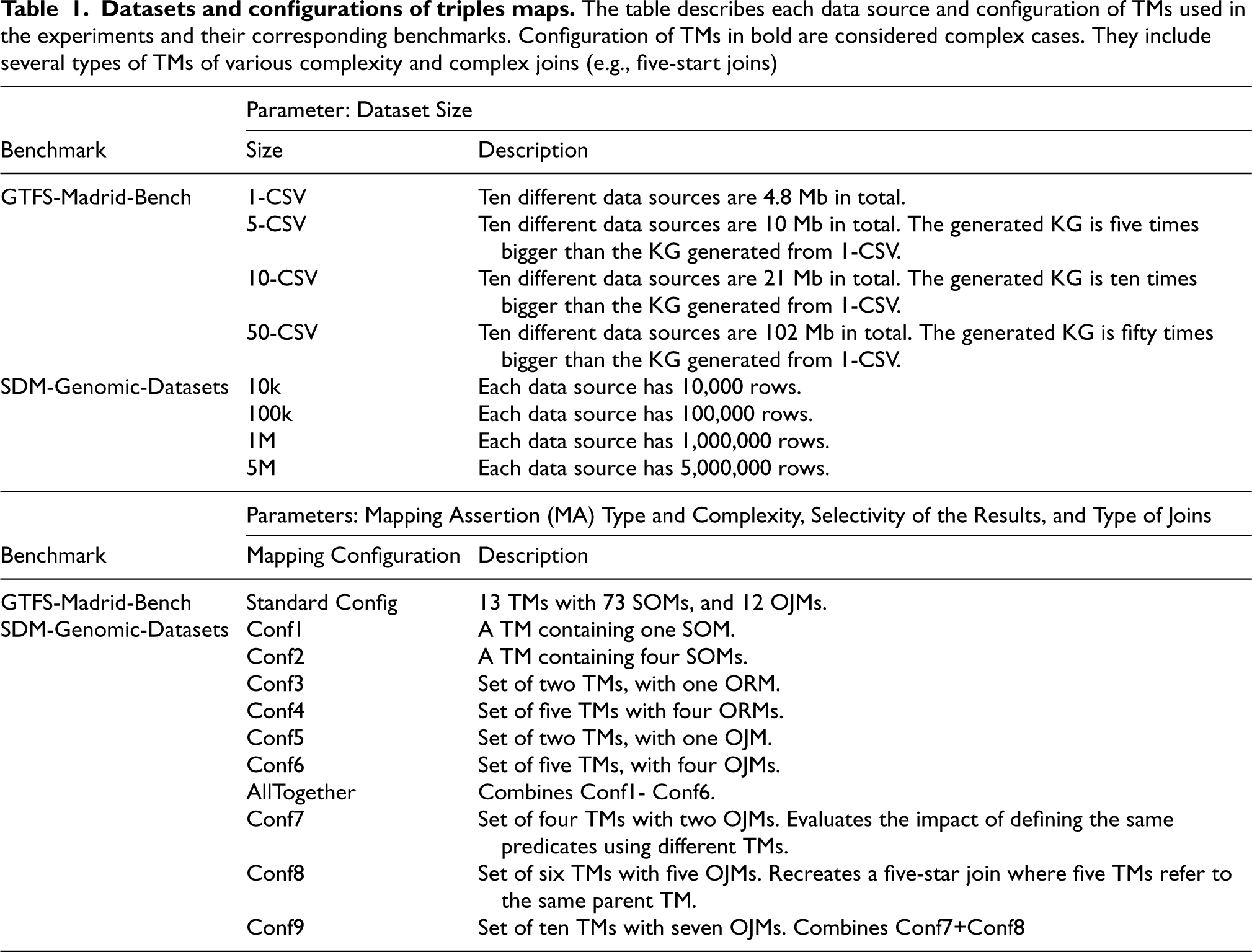

The other state-of-the-art engines were chosen because they excel in handling one of the requirements mentioned above. Morph-KGC [3] was chosen because it presented an extensive study that compares the execution time of Morph-KGC against several RML/R2RML engines in GTFS-Madrid-Bench and SDM-Genomic Datasets, outperforming all of them in most cases. Morph-KGC divides the triples maps into smaller triples maps and executes them in parallel (requirements RE3-Fragmented data and RE8-Mapping complexity). RMLMapper handles heterogeneous data well (requirements RE1-Heterogeneous data and RE6-Semi-structured data), and in the study conducted in Arenas-Guerrero et al. [4], it presents the best level of conformance with respect to test cases defined by RML [30]. Finally, RocketRML was chosen because it has multiple specialized implementations focused on improving join execution between mappings [57] (requirement RE8-Mapping complexity).
The performance of the RML engines is evaluated in terms of the following metrics.
This experiment aims to illustrate the performance increase regarding execution time and memory consumption the proposed data structures and operators will bring when implementing them into a KG creation engine. We evaluate the performance of different versions of the SDM-RDFizer. The previous version of SDM-RDFizer (i.e., SDM-RDFizer v3.2) only contains the operators for triples map transformation and the data structures for duplicate removal and join execution. In contrast, the much more complete version of the SDM-RDFizer (i.e., SDM-RDFizer v4.5.6, a.k.a. SDM-RDFizer+HDT+Flush+Order) contains all the proposed data structures and operators. We also include other combination of the proposed techniques: one only applies data compression (i.e., SDM-RDFizer+HDT), and the other has data compression and main memory flushing but no ordering (i.e., SDM-RDFizer+HDT+Flush).
It can be seen in Fig. 7b that SDM-RDFizer v3.2 and SDM-RDFizer+HDT do not have much difference in execution time; this is because compressing data requires time to be executed; thus, any savings that result from this step can only be appreciated in terms of memory consumption (Fig. 7a). On the other hand, SDM-RDFizer+HDT+Flush and SDM-RDFizer+HDT+Flush+Order reduce execution time compared to the two previous versions of the SDM-RDFizer. This reduction in execution time can be attributed to flushing unneeded data from main memory, thus making the duplicate removal process faster. Unfortunately, there are few savings between these last two configurations of the SDM-RDFizer. This testbed contains 13 triples maps, thus making the organization process take longer and negatively impacting the execution time.
Figure 7a illustrates the maximum memory consumption of the different versions of SDM-RDFizer used. It can be seen in Fig. 7a that the SDM-RDFizer v3.2 is the one that consumes the most memory. By applying data compression, there is a significant reduction in memory consumption caused by the data stored in PTT for duplicate removal, which is much smaller than the data stored in the SDM-RDFizer v3.2. Flushing unneeded data reduces the maximum memory used even further, but not as much as with data compression. Finally, SDM-RDFizer+HDT+Flush and SDM-RDFizer+HDT+Flush+Order have the same maximum memory consumption since the only difference between them is the order in which the triples maps are executed. The benefit of executing the triples maps in a predetermined order is that the maximum amount of data is flushed after finishing the execution of a triples map, thus minimizing the amount of memory being used.

This experiment seeks to prove the impact of using real-world data for KG creation. Even though the triples maps defined for SDM-Genomic-Datasets are simpler than those defined for GTFS-Madrid-Bench, they cover all the triples map types defined in Fig. 1. We evaluate the performance of each engine, i.e., SDM-RDFizer v3.2 and v4.5.6 (i.e., SDM-RDFizer+HDT+Flush+Order), Morph-KGC, RMLMapper, and RocketRML, by measuring the overall execution time it took the engine to complete the KG creation process. As it can be seen in Fig. 8, in case of having ORMs (i.e.,

After observing the performance of SDM-RDFizer during the experimental study and considering the established research questions, the following conclusions have been reached. SDM-RDFizer presented a lower execution time when transforming cases with high duplicate rates, as seen in Fig. 8b, Fig. 8d, Fig. 8f, and Fig. 8h. Therefore, the data duplicate rate is inversely proportional to the execution time. These figures illustrate that SDM-RDFizer has the lowest execution time in complex cases (Conf5, Conf6, AllTogether, Conf7, Conf8, and Conf9) with a high duplicate rate, therefore proving that the duplicate rate of the data impacts the performance of KG creation engines and answering
The SDM-RDFizer characteristics and applications
This section provides an overview of the key features of SDM-RDFizer and outlines its involvement in various projects, highlighting the role it has played.
Main characteristics of SDM-RDFizer
The SDM-RDFizer engine presents a distinctive set of characteristics that make it a valuable contribution for users and practitioners who create KGs for their projects and use cases.
The SDM-RDFizer has been utilized in various industrial and research projects to create KGs from heterogeneous data sources. The following list highlights a selection of these projects.
iASiS,34 EU H2020 funded project to exploit patient data insights towards precision medicine. SDM-RDFizer played a pivotal role in the iASiS project, facilitating the creation of ten versions of the iASiS KG within a span of three years. These KGs, encompassing more than 1.2 billion RDF triples, seamlessly integrate data from over 40 heterogeneous sources using 1,300 RML triples maps [64]. The data sources includes clinical records and genomic data from UK Biobank35 for dementia patients, as well as data from lung cancer patients at the Hospital Puerta del Hierro in Madrid.36 Additionally, the iASiS KGs incorporate structured representations of scientific publications from PubMed,37 drug-drug interactions from DrugBank,38 drug side effects from SIDER,39 and UMLS.40 The integration of these heterogeneous and large datasets underscores the role of SDM-RDFizer in achieving the data management objectives in the iASiS project and specifically fulfilling the requirements Lung Cancer Pilot of BigMedilytics,41 where the KG is defined in terms of 800 RML triples maps from around 25 data sources; it comprises 149,484,936 RDF triples. SDM-RDFizer allowed for the integration of structured clinical records of 1,200 lung cancer patients from Hospital Puerta del Hierro in Madrid and the clinical services visited by these patients, with data about the interactions between the drugs that compose their oncological therapies and their treatments for the comorbidities they may suffer. The use of RML was crucial for defining and maintaining the correspondences among the unified schema and 88 different data sources; SDM-RDFizer enabled the evaluation of these mappings to create nine versions of the KG of the lung cancer pilot of BigMedilytics. Similarly in iASiS, the requirements In CLARIFY,42 nine versions of the project KG were created; they integrate data from lung and breast cancer patients collected in various formats (e.g., CSV and relational databases) with structured representations of publications from PubMed, drug-drug interactions collected from DrugBank, side effects from SIDER, and medical terms from UMLS. The KG definition comprises 1,749 RML triples maps establishing the correspondences with 258 different logical sources. The CLARIFY KG comprises 78M RDF triples and 16M RDF resources. In addition to the requirements satisfied in iASiS and BigMedilytics, here SDM-RDFizer allowed for meeting the requirements of P4-LUCAT43 has 676 RML triples maps that define the P4-LUCAT KG in terms of a unified schema of 318 attributes and 177 classes; it comprises 178M of RDF triples. Data is collected from various sources in different formats, such as CSV and relational databases. As a result, SDM-RDFizer enabled the fulfillment of all the requirements outlined in Section 2.3. The ImProVIT KG44 integrates immune system data into a unified schema consisting of 102 classes, 88 predicates, and 175 attributes. SDM-RDFizer was employed to construct the project KG, generating 6,005,844 RDF triples and 220,414 entities through the evaluation of 577 triples maps. Additionally, for various studies [26], SDM-RDFizer executed multiple versions of these triples maps, resulting in over 40 versions of the ImProVIT KG. The requirements The PLATOON project45 is dedicated to creating the KG for a pilot [37] defined in terms of 2,093 RML triples maps. These mappings define 158 classes and 107 predicates of SEDMOON, the Semantic Data Models of Energy.46 Data were collected from a relational database comprising 600 GB of energy-related observational data collected over six years. The resulting KG comprises 220M RDF triples and 80 million RDF resources. SDM-RDFizer played a crucial role in meeting the requirements of The Knowledge4COVID-19 KG [52] comprises 80M RDF triples integrating COVID-19 scientific publications and COVID-19 related concepts (e.g., drugs, drug-drug interactions, and molecular dysfunctions). It is defined in terms of 57 RML triples maps. All the data was collected from tabular CSV files; SDM-RDFizer allowed for the satisfaction of H2020 – SPRINT47 studies performance and scalability of different semantic architecture for the Interoperability Framework on Transport across Europe. The SDM-RDFizer was used to evaluate the impact of different parameters on the transport domain during the creation of KGs to identify bottlenecks and allow optimizations. Additionally, under this project, the GTFS-Madrid-Bench [16] was also defined, where the SDM-RDFizer was used to materialize the KG used for comparing the performance between native triplestores and virtual KG creation engines. EIT-SNAP48 innovation project on the application of semantic technologies for transport national access points, and SDM-RDFizer allowed the integration of transportation data in Spain. In this specific project, the SDM-RDFizer was integrated in a sustainable workflow to construct KGs based on the Transmodel ontology [51] in a systematic manner [14]. Open Cities49 is a Spanish national project on creating common and shared vocabularies for Spanish cities; SDM-RDFizer executes the RML mapping rule for integrating geographical data for Spanish cities, hence ensuring the interoperability between open data in Spain throughout the created KGs. Virtual Platform for the H2020 European Joint Programme on Rare Disease,50 and SDM-RDFizer merges data collected from the consortium partners, thus, satisfying the requirements in Section 2.3. CoyPU51 is a German-funded project where SDM-RDFizer generates KGs for various events collected from economic value networks in the industrial environment and social context. Specifically, SDM-RDFizer is utilized to create a federation of KGs that integrates data from World Bank, Wikidata, DBpedia, and the CoyPU KG.52 The World Bank dataset comprises 21 topics, ranging from Agriculture and Rural Development to Trade. Each topic has three CSV files associated with it. The CSV files contain annual statistics per country from 1960 to 2022. In total, 63 CSV files are transformed, and a KG comprised of 111 Million RDF triples is generated.
This paper introduces novel data management techniques that leverage innovative data structures and physical operators for the efficient execution of RML triples maps. These techniques have been implemented in SDM-RDFizer v4.5.6, and their effectiveness has been empirically evaluated through 416 testbeds encompassing state-of-the-art RML engines and benchmarks. The results highlight the significant computational power of well-designed data structures and algorithm operators, particularly in complex scenarios involving star joins across multiple triples maps. We anticipate that the reported findings and the availability of the new version of SDM-RDFizer will inspire the community to adopt declarative approaches in defining KG creation pipelines using RML and to explore data management techniques that can further enhance the performance of their engines. For future work, we aim to develop a flushing policy for the Predicate Join Tuple Table (PJTT) to reduce memory consumption by eliminating values of redundant joins. Additionally, we pursue optimizing the Simple Object Map (SOM) and Object Reference Map (ORM) operators to enhance their respective transformation capabilities. Furthermore, we plan to devise efficient data management techniques to empower SDM-RDFizer for executing RML-Star mapping rules; an initial version of this implementation is already available on GitHub53 and extending the current data structures and physical operators is part of our future tasks. We aim to enable the evaluation of observational data, such as sensor data, within the SDM-RDFizer framework. Lastly, we plan to develop parallel operators to empower SDM-RDFizer with fine-grained parallelization – as proposed by Wang et al. [66] – and speed up the process of KG creation, specifically when data sources presented as relational databases are integrated. Extensive empirical assessments with different parallel approaches are also part of our future agenda.
Footnotes
Acknowledgements
Enrique Iglesias is supported by Federal Ministry for Economic Affairs and Energy of Germany (BMWK) in the project CoyPu (project number 01MK21007[A-L]). Maria-Esther Vidal is partially funded by Leibniz Association, program “Leibniz Best Minds: Programme for Women Professors”, project TrustKG-Transforming Data in Trustable Insights; Grant P99/2020. This research was funded by the Galician Ministry of Education, University and Professional Training and the European Regional Development Fund (ERDF/FEDER program) through grants ED431C2018/29 and ED431G2019/04.
