Abstract
Neuro-Symbolic Artificial Intelligence (AI) focuses on integrating symbolic and sub-symbolic systems to enhance the performance and explainability of predictive models. Symbolic and sub-symbolic approaches differ fundamentally in how they represent data and make use of data features to reach conclusions. Neuro-symbolic systems have recently received significant attention in the scientific community. However, despite efforts in neural-symbolic integration, symbolic processing can still be better exploited, mainly when these hybrid approaches are defined on top of knowledge graphs. This work is built on the statement that knowledge graphs can naturally represent the convergence between data and their contextual meaning (i.e., knowledge). We propose a hybrid system that resorts to symbolic reasoning, expressed as a deductive database, to augment the contextual meaning of entities in a knowledge graph, thus, improving the performance of link prediction implemented using knowledge graph embedding (KGE) models. An entity context is defined as the ego network of the entity in a knowledge graph. Given a link prediction task, the proposed approach deduces new RDF triples in the ego networks of the entities corresponding to the heads and tails of the prediction task on the knowledge graph (KG). Since knowledge graphs may be incomplete and sparse, the facts deduced by the symbolic system not only reduce sparsity but also make explicit meaningful relations among the entities that compose an entity ego network. As a proof of concept, our approach is applied over a KG for lung cancer to predict treatment effectiveness. The empirical results put the deduction power of deductive databases into perspective. They indicate that making explicit deduced relationships in the ego networks empowers all the studied KGE models to generate more accurate links.
Keywords
Introduction
Neuro-Symbolic Artificial Intelligence is a research field that combines symbolic and sub-symbolic AI models [3,8,35]. The symbolic models refer to AI approaches based on handling explicit symbols to conduct reasoning and support explainability. On the other hand, AI sub-symbolic systems are based on statistical and probabilistic learning from data mining and neural network models. Symbolic and sub-symbolic systems differ in how they represent and manage data to perform reasoning and prediction. As a result, they aim at solving complementary tasks whose integration has the potential to empower prediction with reasoning supported by symbolic formal frameworks [3,15].
Neuro-symbolic integration aims to bridge the gap between symbolic and sub-symbolic systems; it resorts to translation algorithms to align symbolic to sub-symbolic representations and improve performance [3,8,38]. However, integrating neuro-symbolic into real-world applications is a challenging task. Even in controlled environments, neuro-symbolic integration may not be completed performed [14]. For instance, Fernlund et al. [11] describe systems that use machine learning to learn relations from expert observations. While these systems are successful in learning, they lack the expressive power of symbolic systems. Another example of neuro-symbolic systems combining connectionist inductive learning and logic programming to solve the problems in the molecular biology and power plant fault diagnosis [9]. Furthermore, Karpathy et al. [19] combine convolutional neural networks with bidirectional recurrent neural networks over sentences to recognize and label image regions. Despite these advances in neuro-symbolic AI integration, symbolic processing is not fully exploited, in particular, if reasoning methods are implemented on top of knowledge graphs [38].
A domain-agnostic approach able to empower the predictive performance of sub-symbolic systems with a deductive database system. The deductive system reduces data sparsity issues by inferring implicit relationships in a KG. Consequently, the sub-symbolic system, implemented by KGE models, better represents statements described in the KG into a low-dimensional continuous vector space. An extensive evaluation of our neuro-symbolic system with state-of-the-art KGE models demonstrates the benefit of integrating deductive reasoning and sub-symbolic systems. The evaluation is performed on the problem of predicting the effectiveness of lung cancer treatments composed of multiple drugs, i.e., polypharmacy treatments.
The rest of the paper is structured as follows: Section 2 presents the preliminaries and a motivating example. Section 3 shows the proposed approach and illustrates its main features with a running example. Section 4 applies our hybrid method in the context of predicting the effectiveness of polypharmacy lung cancer treatments. Results of the empirical evaluation of our method are reported in Section 5. Section 6 analyses the state-of-the-art. Finally, we close with the conclusion and future work in Section 7.
V is a set of nodes corresponding to concepts (e.g., classes and entities). L is a set of properties. C is a set of classes
Figure 1(a) depicts a knowledge graph

A knowledge graph
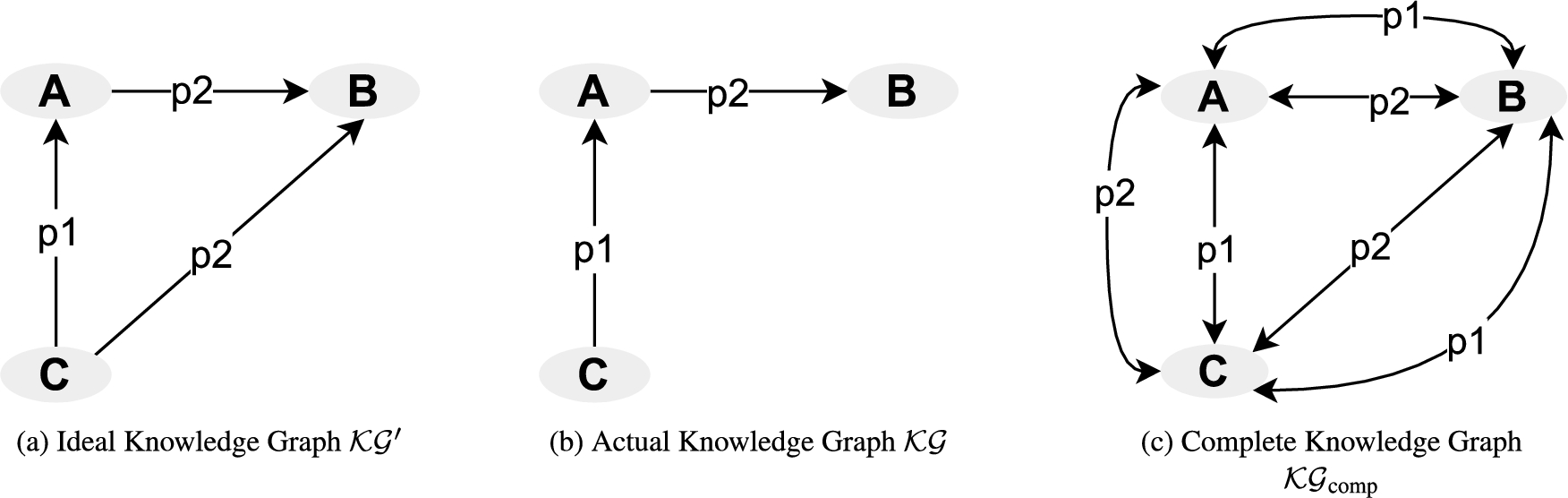
Example of actual, ideal, and complete knowledge graph.
An
r represents a prediction property,
The deductive system
An example of EDB is the set of facts
Suppose the abstract target prediction is defined for the current knowledge graph
We motivate our work in healthcare, specifically for predicting polypharmacy treatment response. Polypharmacy is the concurrent use of multiple drugs in treatments, and it is a standard procedure to treat severe diseases, e.g., lung cancer. Polypharmacy is a topic of concern due to the increasing number of unknown drug-drug interactions (DDIs) that may affect the response to medical treatment. Pharmacokinetics is a type of DDIs, i.e., the course of a drug in the body. Pharmacokinetics DDIs alter a drug’s absorption, distribution, metabolism, or excretion. For example, an increase in absorption will increase the object drug’s bioavailability and vice versa. If a DDI affects the object’s drug distribution, the drug transport by plasma proteins is altered. Moreover, a drug’s therapeutic efficacy and toxicity are affected when a pharmacokinetics DDI alters the object’s drug metabolism. Lastly, if the excretion of an object drug is reduced, the drug’s elimination half-life will be increased. Notice that the pharmacokinetic interactions can be encoded in a symbolic system.
Figure 3(a) shows two polypharmacy oncological treatments encoded in RDF. We extract the known DDIs between the drugs of these treatments from DrugBank. However, polypharmacy therapies produce unforeseen DDIs due to drug interactions in the treatment. Since DDIs affect the effectiveness of a treatment, there is a great interest in uncovering these DDIs. Figure 3(b) depicts an ideal RDF graph where all the existing relations are explicitly represented. Dotted red arrows represent DDI between the drugs

Problem statement
Given an actual knowledge graph
Given a relation,
Proposed solution
Our proposed solution resorts to a symbolic system implemented by a deductive database to enhance the predictive precision of the link prediction task solved by knowledge graph embedding models. The approach assumes that a link prediction problem is defined in terms of an abstract target prediction
The symbolic and sub-symbolic system architecture
Figure 4 depicts the architecture that implements the proposed approach. The architecture receives a knowledge graph

The architecture is composed of two main steps. First, the relationships implicitly defined by the deductive system are deduced by means of a Datalog program. Second, once
Albeit illustrated in the context of treatment response, the proposed method is domain-agnostic. It only requires the definition of the deductive system to enhance the relationships in the ego network of the entities v where

The EDB comprises all the ground facts defined by the ego networks:
The SPARQL query in Listing 1 extracts the ego network

The IDB described by the Datalog program P(1) allows deducing new relationships and increasing the ego networks

Figure 5
As a proof concept, we apply our neuro-symbolic approach to address the problem of predicting polypharmacy treatment effectiveness. We have implemented a deductive system on top of a Treatment Knowledge Graph (
Treatment knowledge graph creation
The P4-LUCAT consortium2
Figure 6 describes a Lung Cancer patient in the Lung Cancer Knowledge Graph. The patient P1 is in stage II and has surgery. Also, P1 received treatment on 10.07.2020 with an effective therapeutic response. In that treatment, P1 was treated with a combination of chemotherapy drugs and one non-oncological drug. Drug-Drug Interactions with the effect and the impact are reported.

The input

For each treatment,
The types Drug, Treatment, DDI, Effect of DDI, and Treatment Response belong to
Drugs, Treatments, DDIs, Effect of DDI, and Treatment Response are represented as instances of V.
Edges in E that belong to
Properties ex:has_response, part_of, ex:precipitant_drug, ex:object_drug, ex:effect, ex:impact, and ex:hasInteraction correspond to labels in L.

Let
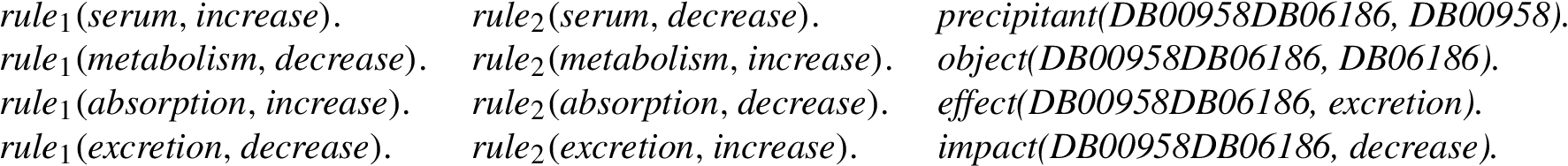
SPARQL queries in Listing 3 and Listing 4 declaratively define the ground


The facts included in the ground predicates precipitant, object, effect, and impact from the EDB are extracted using the CONSTRUCT query of Listing 5. The EDB contains thousands of facts for those predicates; therefore, only a few ground facts are presented.

The above-mentioned
Rule (3) states the base case of the

Once the deductive system
Experimental study
We empirically assess the impact of the DDIs encoded in
Experiment setup
We empirically evaluate the effectiveness of our approach to capture knowledge encoded in
Benchmarks
We conduct our evaluation over three Knowledge Graphs represented in Fig. 9.

We utilize eleven models to compute latent representations, e.g., vectors, of entities and relations in the three KGs and then employ them to infer new facts. In particular, we utilize three main families of models:
Tensor Decomposition models such as HolE and RESCAL. Geometric models such as RotatE, QuatE, and the Trans* family models TransE, TransH, TransD, TransR. Deep Learning models such as UM, SE and ERMLP.
The symbolic-sub-symbolic system proposed is implemented in eleven embedding models from different families [31]. Holographic embeddings (HolE) [23] computes circular correlation, denoted by ⋆ in Table 2, between the embeddings of head and tail entities. RESCAL [24] is an algorithm of relational learning based on tensor factorization, where it models entities as vectors and relations as matrices. In RESCAL, the relation matrices
Scoring function and complexity of embedding models . Adapted from [31]
The PyKEEN (Python KnowlEdge EmbeddiNgs) framework [2] is used to learn the embeddings. The hyperparameters utilized to train the model are epoch number 200 and training loops: stochastic local closed world assumption (sLCWA). The negative sampling techniques used are Uniform negative sampling and Bernoulli negative sampling. The embedding dimensions and the rest of the parameters are set by default. To assure statistical robustness, we apply 5-fold cross-validation. For evaluating the performance of embeddings methods, we measure the metrics:
The pipeline for predicting polypharmacy treatment response has been implemented in Python 3.9. Experiments were executed using 12 CPUs Intel® Xeon(R) W-2133 at 3.60 GHz, 64 GB RAM, and 1 GPU GeForce GTX 1080 Ti/PCIe/SSE2 with 12 GB VRAM. We used the library pyDatalog3
Table 3 shows the statistics of the three KGs. We considered the metrics, Number of Triples (T), Entities (E), and Relations (R), to measure the size in KG. The metrics Relation entropy (
Statistics of knowledge graph . Metrics to measure size, diversity, and sparsity in knowledge graph
The metrics
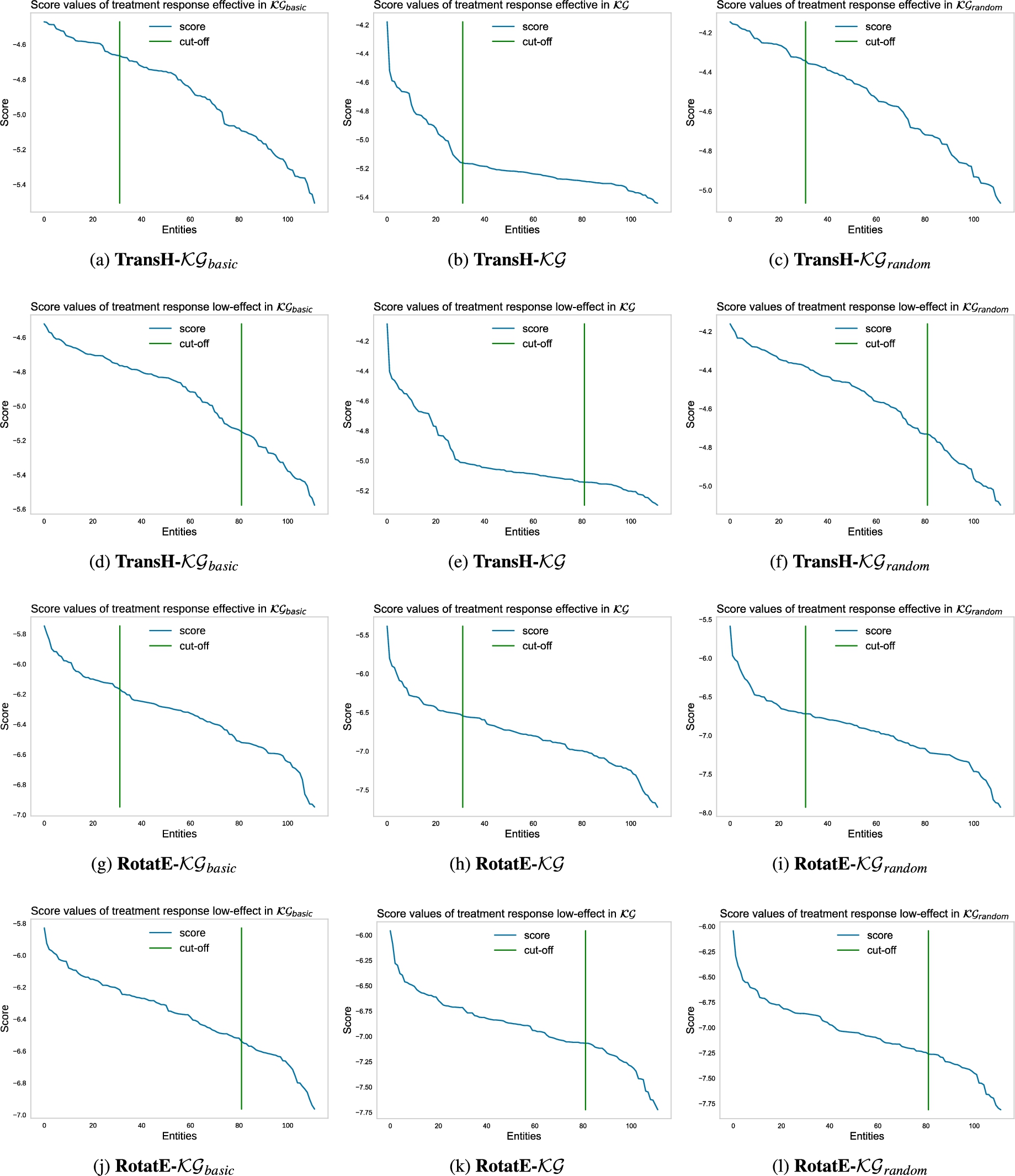
Figure 10 shows the behavior of the scoring function for the entities predicted by TransH and RotatE embedding models. For the purpose of brevity, we only show the score value results for two embedding models. The evaluation material is available .6
The selected portions of entities predicted are measured precision, recall, and f1-score on average because of cross-validation. Figure 11 and Fig. 12 show the evaluation of the Link Prediction task through Uniform negative sampling and Bernoulli negative sampling, respectively. Uniform sampling randomly chooses the candidate entity based on a uniform probability between all possible entities. Bernoulli sampling corrupts the head with probability p and the tail with

Evaluation of the link prediction task in terms of precision, recall, and f-measure. Utilizing uniform negative sampling.

Evaluation of the link prediction task in terms of precision, recall, and f-measure. Utilizing Bernoulli negative sampling.
The techniques proposed in this paper rely on known relations between entities to predict novel links in the KG. During the experimental study, we observed that these techniques could improve the prediction of treatment effectiveness. Figure 13 shows a box plot of cosine similarity. Considering the KGE model with better performance TransH, we computed the cosine similarity between the embedding entities of type treatment. Five treatments with a low-effect response are selected, and TransH in
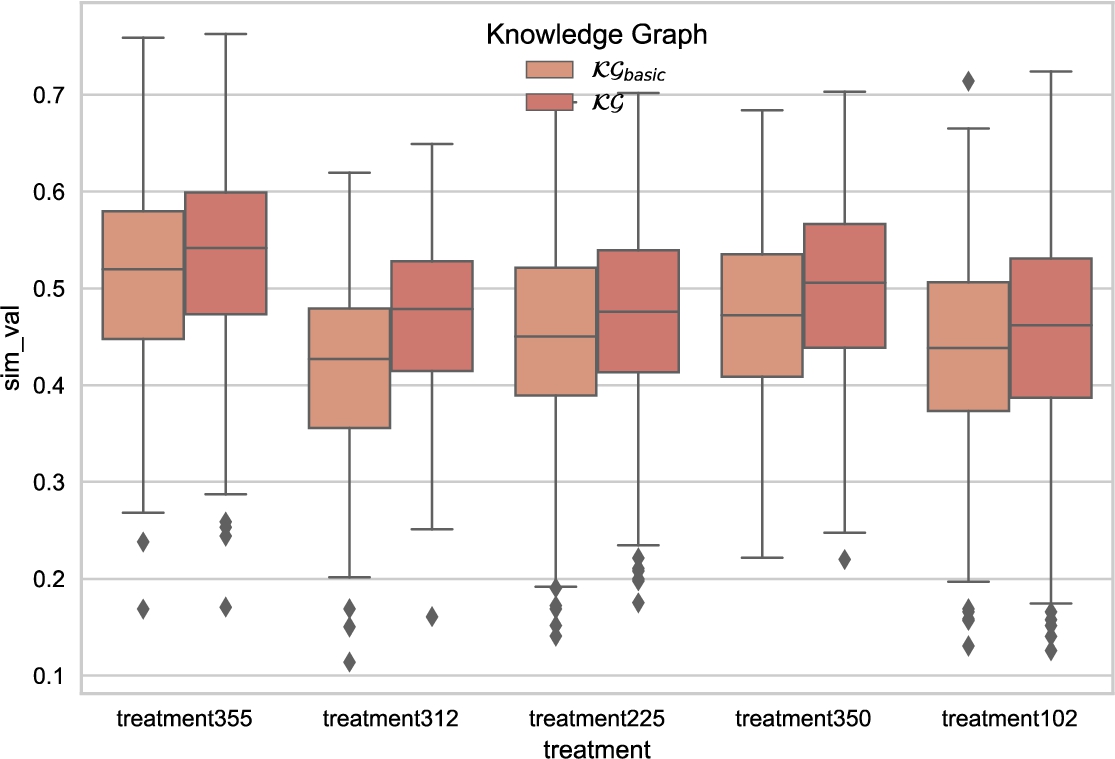

Figure 14 shows the distribution of DDIs by treatment in

Distribution of DDIs by treatment response.
Neuro-symbolic artificial intelligence
Neuro-Symbolic Artificial Intelligence is a highly active area that has been studied for decades [8]. Neuro-symbolic AI focuses on integrating symbolic and sub-symbolic systems. Several approaches employ translation algorithms from a symbolic representation to a sub-symbolic representation and vice versa [3]. The aim is to provide a neuro-symbolic implementation of logic, a logical characterization of a neuro-system, or a hybrid learning system that contributes features of symbolic and sub-symbolic systems [8,38]. Real applications are possible in areas with social relevance and high economic impacts, such as bioinformatics, robotics, fraud prevention, and the semantic web [3]. Methods utilized in neuro-symbolic integration in some of the aforementioned applications include translation algorithms between logic and networks. Also, the community has focused on studying the systems empirically through case studies and real-world applications. An example of a neuro-symbolic system in the field of bioinformatics is the Connectionist Inductive Learning and Logic Programming (CILP) [9]. In the field of vision-based tasks, such as semantic image labeling, high-performance systems have been produced. Karpathy et al. [19] propose an approach introduced for the recognition and labeling tasks for the content of different regions of the images; it combines Convolutional Neural Networks over the image regions together with bidirectional Recurrent Neural Networks over sentences. Once this mapping of images and sentences in the embedding space has been established, a structured objective is introduced that aligns the two modalities through multimodal embedding. The emerging system performs better than classical approaches, where tasks involving semantic descriptions are associated with databases that contain background knowledge, and computer image processing approaches are based on rule-based techniques.
Despite the progress of Neuro-Symbolic Artificial Intelligence, the scope and applicability of symbol processing are limited. Furthermore, these systems do not examine polynomial overload when integrating both paradigms. Our work leverages the symbolic system, independent of the application domain, and improves the predictive precision of KGE models. Moreover, in our approach, the deductive database is addressed to an abstract target prediction which renders the computational complexity polynomial-time. Thus, we show the positive impact on the overall performance of a predictive model implemented using KGEs considering a deductive system.
Knowledge graph embedding in biomedical field
Knowledge graphs are becoming increasingly important in the biomedical field. Discovering new and reliable facts from existing knowledge using KGE is a cutting-edge method. KG allows a variety of additional information to be added to aid reasoning and obtain better predictions.
Zhu et al. [46] develop a process for constructing and reasoning multimodal Specific Disease Knowledge Graphs (SDKG). SDKG is based on five cancers and six non-cancer diseases. The principal purpose is to discover reliable knowledge and provide a pre-trained universal model in that specific disease field. The model is built in three parts: structure embedding (S) with TransE, TransD, and ConvKB, category embedding (C), and description embedding (D) with BioBERT to convert description annotations into vectors. The best results are obtained when description embedding is combined with structure embedding, specifically with the ConvKB embedding model. Karim et al. [18] propose a new machine-learning approach for predicting DDIs based on multiple data sources. They integrated drug-related information such as diseases, pathways, proteins, enzymes, and chemical structures from different sources into a KG. Then different embedding techniques are used to create a dense vector representation for each entity in the KG. These representations are introduced in traditional machine learning classifiers and a neural network architecture based on a convolutional LSTM (Conv-LSTM), which was modified to predict DDIs. The results show that the combination of KGE and Conv-LSTM performs state-of-the-art results.
The above-mentioned research aims to discover reliable knowledge based on knowledge graphs using KGE models. However, they are limited by the data sparsity issue of the KGE models and the lack of symbolic reasoning. We overcome this limitation by integrating a Neuro-Symbolic AI system, enabling expressive reasoning and robust learning to improve the predictive performance of KGE models.
Polypharmacy side effect prediction and drug-drug interactions prediction
In recent years, there has been a growing interest in Pharmacovigilance. Extensive research has been conducted to predict potential DDI. One approach to predicting potential DDI is based on similarity [12,36,40,43], with the core idea of predicting the existence of a DDI by comparing candidate drug pairs with known interacting drug pairs. These approaches define a wide variety of drug similarity measures for comparison. The known DDIs that are very similar to a candidate pair provide evidence for the presence of a DDI between the candidate pair drugs. Sridhar et al. [36] propose a probabilistic approach for inferring unknown DDIs from a network of multiple drug-based similarities and known DDIs. They used the probabilistic programming framework Probabilistic Soft Logic. This symbolic approach predicts three types of interactions [36], CYP-related interactions (CRDs), where both drugs are metabolized by the same CYP enzyme, NCRDs, where no CYP is shared between the drugs and general DDI from Drugbank. Furthermore, they considered seven drug-drug similarities. Thus, they found five novel DDIs validated by external sources. A framework to predict DDIs is presented in [12]; they exploit information from multiple linked data sources to create various drug similarity measures. Then, they build a large-scale and distributed linear regression learning model to predict DDIs. They evaluate their model to predict the existence of drug interactions, considering the DDIs as symmetric. A neural network-based method for drug-drug interaction prediction is proposed in [30]. They use various drug data sources in order to compute multiple drug similarities. They computed drug similarity based on drug substructure, target, side effect, off-label side effect, pathway, transporter, and indication data. The proposed method first performs similarity selection and then integrates the selected similarities with a nonlinear similarity fusion method to obtain high-level features. Thus, they represent each drug by a feature vector and are used as input to the neural network to predict DDIs.
Other approaches focus on predicting DDIs and their effects [20,22,32,47]. Beyond knowing that a pair of drugs interact, it is essential to know the effect of DDI in polypharmacy treatments. In [20], propose a novel deep learning model to predict DDIs and their effects. They use additional features based on structural similarity profiles (SSP), Gene Ontology term similarity profiles (GSP), and target gene similarity profiles (TSP) to increase the classification accuracy. The proposed model uses an auto-encoder to reduce the dimension of the resulting vector from the combination of SSP, TSP, and GSP. The benchmark used has 1597 drugs and 188’258 DDIs with 106 different types. The model works as a multi-label classification model where the deep feed-forward network has an output layer of size 106, representing the number of DDI types. The results show that the model obtains equal or better results in 101 out of 106 DDI types than baseline methods. Also, they demonstrate how adding the features GSP and TSP increases the accuracy of DDIs prediction. Marinka Zitnik et al. [47] present Decagon, an approach for predicting the side effects of drug pairs. The approach develops a new convolutional graph neural network for link prediction. They construct a multi-modal graph of protein-protein interactions, drug-protein target interactions, and the DDI side effects. The graph encoder model produces embeddings for each node in the graph. They proposed a new model that assigns separate processing channels for each relation type and returns an embedding for each node in the graph. Then, the Decagon decoder for polypharmacy side effects relation types takes pairs of embeddings and produces a score. Thus, Decagon can predict the side effect of a pair of drugs.
All the approaches mentioned above are limited to predicting DDIs and their effects between pairs of drugs. However, in our view, the interactions and their effects need to be considered as a whole and not in pairs in polypharmacy treatments. Our symbolic system resorts to a set of rules that state the implicit definition of new DDIs generated as a result of the combination of multiple drugs in treatment. Since cancer treatment schemes are usually composed of more than one drug, and patients may have several co-existing diseases requiring additional medications, it is of significant relevance to the deduction of DDIs holistically in a given treatment.
Conclusions and future work
This paper addresses the problem of Neuro-Symbolic AI integration, enabling expressive reasoning and robust learning to discover relationships over knowledge graphs. We have presented an approach that integrates symbolic-sub-symbolic systems to enhance the predictive performance of abstract target prediction in KGE models. The symbolic system is implemented by a deductive database defined for an abstract target prediction over a KG. The proposed solution builds the ego networks of the head and tail of the abstract target prediction to deduce new relationships in the ego network; it is able to enhance the ego networks of the abstract target prediction and effectively predict treatment effectiveness. Further, the sub-symbolic system implemented by a KGE model enhances the predictive performance of the abstract target prediction and completes the KG. The performance of the proposed approach is assessed in a knowledge graph for lung cancer to discover treatment effectiveness. Predicting treatment effectiveness is effectively modeled as a link prediction problem, and exploiting DDI Deductive System improves existing embedding models by performing the treatment prediction task. Results of a 5-fold cross-validation process demonstrate that our approach, integrating neuro-symbolic systems, improves the eleven KGE models evaluated. The presented approach using the symbolic system’s reasoning can enhance the ego networks of the abstract target prediction and effectively predict treatment effectiveness. Thus, our work broadens the repertoire of Neuro-Symbolic AI systems for discovering relationships over a KG. As for future work, we envision having a more fine-grained description of the DDIs and a descriptive profile of the patients and improving the model.
Footnotes
Acknowledgements
Ariam Rivas is supported by the German Academic Exchange Service (DAAD). The authors thank the BIOMEDAS program for training. This work has been partially supported by the EU H2020 RIA funded projects CLARIFY with grant agreement No 875160, EraMed P4-LUCAT No 53000015, and Opertus Mundi GA 870228, as well as the Federal Ministry for Economic Affairs and Energy (BMWi) project SPEAKER (FKZ 01MK20011A). Furthermore, Maria-Esther Vidal is partially supported by Leibniz Association in the program “Leibniz Best Minds: Programme for Women Professors”, project TrustKG-Transforming Data in Trustable Insights with grant P99/2020.
