Abstract
Tailoring personalized treatments demands the analysis of a patient’s characteristics, which may be scattered over a wide variety of sources. These features include family history, life habits, comorbidities, and potential treatment side effects. Moreover, the analysis of the services visited the most by a patient before a new diagnosis, as well as the type of requested tests, may uncover patterns that contribute to earlier disease detection and treatment effectiveness. Built on knowledge-driven ecosystems, we devise DE4LungCancer, a health data ecosystem of data sources for lung cancer. In this data ecosystem, knowledge extracted from heterogeneous sources, e.g., clinical records, scientific publications, and pharmacological data, is integrated into knowledge graphs. Ontologies describe the meaning of the combined data, and mapping rules enable the declarative definition of the transformation and integration processes. DE4LungCancer is assessed regarding the methods followed for data quality assessment and curation. Lastly, the role of controlled vocabularies and ontologies in health data management is discussed, as well as their impact on transparent knowledge extraction and analytics. This paper presents the lessons learned in the DE4LungCancer development. It demonstrates the transparency level supported by the proposed knowledge-driven ecosystem, in the context of the lung cancer pilots of the EU H2020-funded project BigMedilytic, the ERA PerMed funded project P4-LUCAT, and the EU H2020 projects CLARIFY and iASiS.
Introduction
Lung cancer (LC) is Europe’s most common cause of cancer death, with an estimated 353,000 deaths yearly. LC has the highest economic cost in Europe, with direct costs of caring for patients with the disease amounting to more than €3 billion per year [57]. Although costly, lung cancer therapies can be more effective, and the chances to respond are higher when diagnosed in the early stages [42].
Biomedical data have experienced exponential growth in the last decade; they encode valuable knowledge which can be exploited for accurate disease diagnostics and personalized treatments [51,53]. Nevertheless, lung cancer is a heterogeneous disease whose precise diagnosis requires a holistic analysis of multiple variables, usually collected from data sources represented in myriad formats. Some examples of these heterogeneous data sources include electronic health records (EHRs) comprising unstructured clinical notes expressed in a particular language (e.g., Spanish, English, or German); EHRs offering structured data annotated with controlled vocabularies (e.g., SNOMED/LOINC2
Various computational tasks must be implemented to ensure interoperability across heterogeneous data sources. In the case of unstructured datasets, Natural Language Processing (NLP) techniques are required to recognize biomedical entities and link them to biomedical-controlled vocabularies or ontologies in all these data sources. Additionally, data exchange, sharing, and processing need to respect data privacy and access regulations imposed by the data providers and ethical and legal committees. Lastly, the decisions made during data processing need to be interpretable and verifiable. These data complexities impose requirements that must be solved toward a meaningful analysis of knowledge encoded by integrating these data sources.
To put the role of data integration into perspective, this paper presents patterns between lung cancer treatments and the interactions among the drugs that compose each treatment. The studied therapies are collected from clinical records, while drug–drug interactions are extracted from DrugBank and the scientific literature, and inferred using a deductive system. They represent use cases where analytics on top of integrated data can support a better understanding of the factors that may impact treatment effectiveness.
The process of data integration is also defined using declarative languages R2RML (a W3C standard), RML (the RDF Mapping Language), and FnO (the Function Ontology). The data integration process is declaratively defined as mapping rules in RML + FnO; they express correspondences between data sources, and classes and properties from the unified schema. Transformation functions are expressed in FnO and included as part of the mapping rules. This integrated view of data pre-processing and integration results in a modular and reusable specification of the KG creation process, which can be easily verifiable and traceable. Web APIs have been implemented over the KG and the data processed by each basic DE. The goal is to uncover patterns in the hospital services visited by lung cancer patients that provide insights into the conditions of these patients before the lung cancer diagnosis. The results of these analyses have driven the design of five clinical interventions to identify which of the hospital services visited by lung cancer patients have more potential for diagnosis and may contribute to earlier detection. The reported results uncover patterns in the visited services that provide insights into the potential clinical conditions of patients diagnosed with lung cancer. Although further analyses are required, these patterns can support early diagnosis and prognosis. More importantly, if validated, they will allow clinicians to detect the disease in an asymptomatic phase, reducing complications, which usually increase the complexity of these patients and their response.
DE4LungCancer has been applied in the context of iASiS,3
In these projects, DE4LungCancer enables the integration of biomedical data sources and provides a knowledge graph from where analytical methods are performed. As a proof of concept, this paper presents some of these methods and reports on the outcomes that have motivated the execution of clinical interventions to enhance treatment effectiveness and lung cancer patients’ quality of life. The portion of the DE4LungCancer KG that comprises open data is publicly available and accessible via a SPARQL endpoint.7
European Health Data Ecosystems target to strengthen the sustainability of health systems across Europe by reducing costs while improving quality and access to care.8
Requirements can be classified into three categories:
The main goal of a lung cancer data ecosystem is to develop analytical tools that give oncologists insights to improve the management of patients with lung cancer during their treatment, follow-up, and last period of life through data-driven techniques. Additionally, they aim to improve patients’ experience, satisfaction, and primary outcomes and save substantial health costs. Moreover, admissions and readmissions due to toxicities and comorbidities present in lung cancer patients need to be traced to reduce visits to emergency care and hospitalizations. A lung cancer data ecosystem should provide the basis to identify the potential side effects of a lung cancer treatment and the adverse events generated by the interactions among the treatment drugs.
There are four different categories of stakeholders in a Lung Cancer data ecosystem. Data are exchanged across these stakeholders, preserving data access and privacy regulations.
DE4LungCancer
The DE4LungCancer framework is devised as a network of data ecosystems (DEs) [20]; it aligns data and metadata to describe the network and its components. Heterogeneity issues across the different datasets are overcome by various data curation and integration methods. Each DE comprises datasets and programs for accessing, managing, and analyzing their data. Interoperability issues across the datasets of the DEs are solved in a unified schema. Mapping rules between the datasets and the unified schema describe the meaning of the datasets. Figure 1 illustrates the components of the DE4LungCancer Data Ecosystem. The metadata layer specifies biomedical vocabularies (e.g., Unified Medical Language System-UMLS10
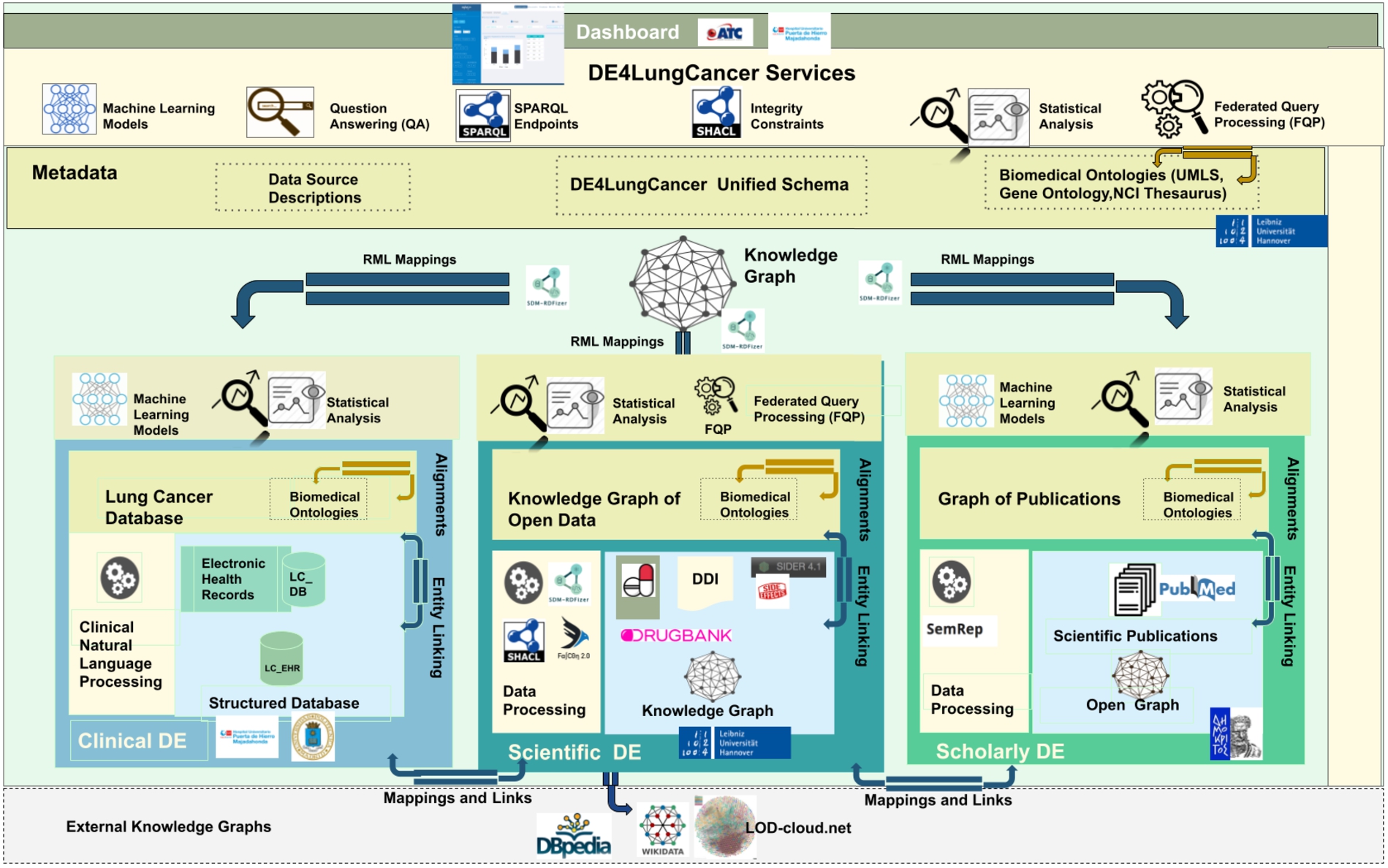
The DE4LungCancer data ecosystem.
Clinical data ecosystem
Clinical data is collected from electronic health records from more than 1,242 lung cancer patients registered in the Electronic Health Record (EHR) system at the Puerta del Hierro University Hospital in Madrid from 2008 to January 2020. The data is extracted from 315,891 notes and 16,550 reports; it represents clinical variables of lung cancer patients and the services consulted by those patients before and after diagnosis. A pseudonymization process maps each patient with a local identifier in the Clinical DE. This way, all the clinical notes and examinations of the same person are integrated. The clinical data providers generate and keep alignments between the actual identifiers and the pseudonymized ones. The statistical analysis performed on EHR follows a stage of Natural Language Processing of raw data to extract patient characteristics and visited medical services at the hospital. The (statistical) analysis performed on EHR concerned KPI-1: Length of hospital stay; and KPI-2: Identification of people at risk of developing lung cancer.
Raw data: 1,242 EHR of patients from 2008–2020, 416 patients were hospitalized 942 times. Out of these 416 patients, 166 had one hospitalization in the first three months after diagnosis. The remaining 250 did not have a hospitalization in the first three months after diagnosis, but they had at least one hospitalization up to six months after the diagnosis.
NLP processing on EHR: Natural Language Processing (NLP) techniques are applied to EHR to extract relevant entities from unstructured fields, i.e., clinical notes or lab test results. The NLP techniques rely on medical vocabularies and rules to perform lemmatization, Named Entity Recognition (NER). The final result is annotating the extracted concepts (i.e., Named Entities) with terms from medical vocabularies. Figure 2 depicts the NLP pipeline that transforms unstructured EHRs into structured data.

NLP pipeline for knowledge extraction in the clinical data ecosystem.
Scholarly data are obtained by harvesting scientific publications from PubMed (i.e., article abstracts) and PubMed Central (i.e., article full-texts), along with scholarly metadata such as the author list, journal, and publication year. To retrieve publications only related to lung cancer, the Entrez Programming Utilities API12
Natural Language Processing is applied to the article abstracts or the whole text of the article, where it is freely available. Analyzing scientific publications’ text, triplets consisting of two entities connected by a relation (e.g., Hemofiltration-TREATS-Patients) are being produced. The process is performed with industry-standard software. Metamap16

NLP pipeline for knowledge extraction implemented in the scholarly data ecosystem.
Scientific databases (e.g., DrugBank and SIDER) and encyclopedic knowledge bases (e.g., Wikidata and DBpedia) are the main sources of open data. These sources encode knowledge about drugs, their approved indications, the side effects, and the drug–drug interactions and effects. All these features are present as short textual descriptions. In the provided version of the Scientific Open DE, we have collected data from DrugBank18
The techniques of named entity recognition (NER) and named entity linking (NEL) enable the identification of biomedical entities from textual attributes. The rule-based entity linking engine, FALCON [49], performs NER and NEL on this data ecosystem. FALCON is configurable for linking entities to diverse controlled vocabularies or knowledge graphs (KG), e.g., UMLS, DBpedia, or Bio2RDF. FALCON recognizes entities by mapping instances of a word within a short text, i.e., surface forms into the textual representation of entities in a controlled vocabulary or KG. FALCON resorts to a knowledge base and a catalog of rules for recognizing and linking entities. The knowledge base integrates various sources, e.g., DBpedia, Wikidata, Oxford Dictionary, and Wordnet. Additionally, it comprises alignments between nouns and entities in these sources. Alignments are stored in a text search engine, e.g., ElasticSearch, while the knowledge sources are maintained in an RDF triple store accessible via SPARQL endpoints. Moreover, the catalog of rules encodes the English morphology; they are represented as conjunctive rules and provide a forward chaining inference process for entity recognition in English short texts. The main feature of FALCON is the ability to split a short input text into a minimal number of entities that more precisely represent the words in the text. Thus, FALCON is devised to solve the optimization problem of maximizing the number of words linked to an entity/relation while minimizing the number of recognized entities/relations. This feature is extremely relevant for the scientific open data ecosystem entity, e.g., a drug or a disease can be expressed with several words, e.g., thoracic aortic aneurysms. In Section 5.1, we report the results of analyzing the effects of DDIs in a treatment’s response; the NER and NEL method performed by FALCON is named DrugBank because DDIs are extracted from this database. Additionally, the deductive system (DS) proposed by Rivas and Vidal [46] uncovers DDIs resulting from combining several drugs. The extensional database comprises the DDIs extracted by FALCON from DrugBank. At the same time, the intensional rules state the conditions to be met, among the interactions of a group of drugs, to generate new DDIs. We name this method DS prediction in the evaluation reported in Section 5.1.
As proposed by Geisler and Vidal et al. [20], the nested DE4LungCancer Data Ecosystem is defined as a 6-tuple
Datasets
The DE4LungCancer Data Ecosystem integrates three categories of data sources collected from the basic data ecosystems:

An exemplary RML mapping rule calling one FnO function named
Biomedical ontologies and controlled vocabularies describe the data and provide a unified description and annotation. These annotations represent the basis of the data integration methods to merge the data into a KG. The values in the datasets are annotated with terms from the Unified Medical Language System. These annotations enable entity alignment and provide the basis for integrating the datasets into the KG. The KG includes 3,862,429 terms from the semantic groups “Anatomy”, “Disorders”, “Physiology”, “Procedures”, “Concepts & Ideas”, “Chemicals & Drugs”, “Living Beings”, “Activities & Behaviors”, “Objects”, “Devices”, “Phenomena”, “Occupations”, “Organizations”, “Geographic Areas”, and “Genes & Molecular Sequences”. A unified schema provides an integrated view of the data sources. The DE4LungCancer unified schema is expressed in the W3C standard data model RDF. This increases interoperability and facilitates reusability of existing vocabularies and ontologies, e.g., the RDF Schema22
The correspondences between the data sources and the unified schema are defined using the W3C standards RDF Mapping Language (RML) [13] and R2RML. R2RML and RML mapping rules can comprise transformation functions expressed in existing ontologies (e.g., the Function Ontology-FnO). These mappings are expressed in RDF and can be stored in a triplestore (e.g., Virtuoso or GraphDB). Exemplar SPARQL queries are presented in Section A. Query in Listing 1 retrieves metadata about the RML mapping rules that define a particular class, while query in Listing 2 collects the functions included in the RML mapping rules. These functions are expressed in FnO and are part of the toolbox EABlock26

RML mapping rules per RDF class in the DE4LungCancer unified schema.

RDF classes in the KG. Annotations of entities extracted from publications (
SDM-RDFizer [26], an in-house RML-compliant engine, is utilized to integrate data from the data sources into the KG following the mapping rules. As a result, a KG of 19,602,972 biomedical entities described in terms of 110,788,660 RDF triples is created. Moreover, 3,900,764 links to DBpedia, Wikidata, and UMLS are part of the KG; they are discovered by the tasks of NER and NEL executed by the FnO function included in the mapping rules and by the NLP processes implemented in each DE. Figure 6 depicts the number of entities of the classes in the KG. The classes

A portion of the DE4LungCancer KG.
The integrity constraints to be satisfied by the data sources are expressed in terms of rules validated with the clinical partners to ensure completeness and soundness. To enhance traceability, the integrity constraints are expressed declaratively using the Shapes Constraint Language (SHACL) [32] and SPARQL [43]. SHACL rules are defined over a class’ attributes (i.e., owl:DatatypeProperties) and constraints on incoming/outgoing arcs, cardinalities, RDF syntax, and extension mechanisms. Inter-class constraints induce a shape network used to validate the KG’s integrity and data quality properties. SHACL relies on validation errors and validation reports using a controlled vocabulary in RDF. A validation report includes explanations about the violations and their severity, and a message describing the violation. In total, there have been defined 67 rules to validate the integrity of the DE4LungCancer KG.
Services
The mapping rules are also utilized to validate the correctness of the process of KG creation. In the DE4LungCancer DE, a quality assessment process guided by the mapping rules is executed on top of the KG. It ensures that each class and predicate in the KG has the same number of instances as the data sources from where the data to populate these classes and properties are extracted. Moreover, different services have been implemented on top of the KG. Several statistical analyses derived from various parameters (e.g., hospitalization, emergency room visits, toxicities, medical tests performed, and oncological treatment types) are integrated as services of the DE4LungCancer dashboard. Additionally, services to traverse the scientific publications associated with a cohort of patients or the drug–drug interactions and side effects of these patients’ treatments can be explored. Thus, the KG acts as a knowledge repository of the DE4LungCancer DE, which empowers the interpretability of the conditions and treatments of the selected cohort. Lastly, DE4LungCancer resorts to a federated query engine to interoperate across the DE4LungCancer KG, DBpedia, and Wikidata; the query processing methods by Endris et al. [14] are implemented to ensure efficiency during query execution over these distributed KGs.
Instances of the classes describing a lung cancer treatment in the open DE4LungCancer KG
Instances of the classes describing a lung cancer treatment in the open DE4LungCancer KG
Considering the data collected from the Scientific and Scholarly DEs, we have created a reduced version of the DE4LungCancer KG; it comprises 34,572,162 RDF triples and is publicly available via a SPARQL endpoint. 7 This open version of the KG comprises 136 classes. These classes represent publications, drugs, drug–drug interactions, and UMLS annotations. Additionally, all the lung cancer treatments reported in the clinical data processed by the Clinical DE have been integrated into the KG. Treatments are documented in terms of the prescribed drugs and the drug–drug interactions (DDIs) among these drugs. Three types of DDIs are also part of the KG (i.e., Literature, DrugBank, and DS). Moreover, the KG also represents the number of patients that observed a particular response when a treatment was administrated. The numbers of instances of these classes are listed in Table 1. Thus, the Open DE4LungCancer KG makes publicly available all the data required to reproduce the analysis reported in Section 5.1. Appendix B presents some exemplary SPARQL queries that enable the exploration of the KG. The ontology and RML mappings required to generate this KG, as well as the SPARQL queries, are available in GitHub.28
Integrating various data sources and ecosystems realized by DE4LungCancer DE, many of which are a result of Natural Language Processing techniques, sets several challenges concerning the data quality (DQ). NLP processing of raw text, e.g., in Electronic Health Records, produces a number of ambiguities on medical terms, and imprecise Named Entity Recognition can result in noisy output information. In this context, we define numerous integrity constraints as part of the DE4LungCancer DE metadata and apply error detection techniques to identify erroneous triples in the ecosystem Knowledge Graph. Moreover, certain mapping rules enrich the DE4LungCancer KG interconnectivity of different sources. At the same time, the data provenance metadata enables the filtering of the information retrieved from the DE4LungCancer DE end-users via the corresponding dashboards. In the following paragraphs, we attempt to investigate the different quality aspects related to each data ecosystem introduced in Section 3.1 and present the data curation and noise detection approaches followed.
DQ in clinical data ecosystem
The data quality methodology is composed of four steps: (a) Definition of the constraints; (b) Validation of the constraints; (c) Human validation by the domain experts; and (d) Resolution of the ambiguities. First, the metadata describing the DE4LungCancer data sources and the description of the universe of discourse represented in these data sources are analyzed to identify integrity constraints. Clinical and technical partners were consulted to collect the main constraints to be satisfied. Moreover, the concepts and relations existing in the unified schema were utilized to guide the definition of the constraints. First, constraints describing the properties of the attributes of a class in the DE4LungCancer unified schema were identified, i.e., intra-concept constraints, and next, constraints regarding the relationships existing between these concepts or inter-concept constraints are determined. Intra-concept constraints include (a) data types of the attributes, (b) attribute dependencies, (c) cardinalities, and functional dependencies. Additionally, inter-concept constraints encompass referential integrity, cardinality and connectivity, and mandatory and optional relationships among the concepts in the unified schema. Once the constraints are recognized, they are formally specified as expressions of SQL, SHACL, and SPARQL, and evaluated both over the corresponding raw data and the data integrated into the KG; the SHACL validation engine Trav-SHACL [18] was used to validate the SHACL constraints against the KG.
Moreover, inconsistencies between the results obtained after evaluating the constraints over raw data and the KG reveal errors in the process of integration in the KG. On the other hand, equal numbers of ambiguities in the raw data and the KG evidence a data quality issue in the original dataset or in the extraction process. Finally, when all the issues had been detected and classified, the clinical and technical partners were consulted to find the most suitable way to curate either the raw data or the KG. This methodology implements techniques reported by Acosta et al. [3], Ruckhaus et al. [48], and Mihaila et al. [38]. For the class LCPatient, all the attributes were analyzed, as well as the concepts to which this concept is connected. Table 2 summarizes intra- and inter-concept constraints in the business domain of lung cancer. These constraints have been validated by four knowledge engineers, two experts in the NLP extraction process, and two experts in lung cancer; all these evaluators are partners of the consortium. As a result, 67 constraints are defined and a total of 3.120 ambiguities are detected in the NLP-processed clinical datasets and in their corresponding instances in the KG. Table 2 reports on the distribution of the constraints, attributes and concepts. As observed, most ambiguities are detected in the tumor stages, line of treatments, oncological surgeries, and biomarkers. All these ambiguities were discussed with the clinical partners and curated following their recommendations and directions. The integrity constraints are part of the metadata of the DE4LungCancer DE; they document the quality assessment and curation tasks and trace the changes made during data curation.
Number of constraints and ambiguities in the class DE4LC:LCPatient
Number of constraints and ambiguities in the class DE4LC:LCPatient
In contrast to clinical data that have been manually filled by experts, the knowledge published in scholarly sources may usually be less reliable. Although being reviewed by field experts, published literature can still report preliminary results, observations, and unverified hypotheses. Moreover, given that any NLP software used to automatically extract knowledge from text is far from perfect, we expect a significant amount of inherent noise and unreliable information in the open data graph (i.e., the one resulting from the processing of the scientific publications). As mentioned, we employ two mainstream tools in the field of biomedical knowledge extraction, to perform entity recognition and relation extraction on literature text. MetaMap [4] and SemRep [45] tools have been evaluated on benchmark datasets achieving high precision (>76%) and moderate recall (36%–70%), on various datasets [10,12,31]. The quality of data in the open graph produced by those tools is addressed in two ways. First, each triplet is associated with a quality score, that is related to the confidence scores provided by MetaMap, representing the quality of each concept identification. In specific, a triple-extraction quality score ranging from 0 to 1 (i.e., the higher, the better) has been added, by averaging the concept identification score of the subject and object entities of each triple. These concept identification scores provide the average of the scores for all found instances of the entities in the specific relation, in order to consider the frequency of the concepts found in the scientific publications. Second, to assess the quality of the open graph as a whole, we have developed an error detection methodology [9] that is based on graph topology and theoretic measures to assess the quality of all edges in this graph. This method, called Path Ranking Guided Embeddings (PRGE), combines an extension of the Path Ranking Algorithm [34] (PaTyBRED [37]) with translational graph embeddings (TransE [7]). The aim is to generate confidence-guided graph embeddings identifying erroneous triples by providing global-confidence scores for all automatically generated relations. We evaluate PRGE using two benchmarks and one generated dataset. The AUC score ranges from 0.56 to 0.97, based on the quality of the dataset used, and the followed noise imputation approach, improving in most cases, simple PRA and embedding methods.
Besides the errors imputed by automatic NLP systems, the quality of the information provided by publications can also be dubious. From the end-user perspective, when exploring publications included in the Scholarly DE, it would be appropriate to be able to filter out unreliable publications or focus only on trustworthy institutes and journals. To this end, DE4LungCancer provides the ability to explore scientific literature, using various factors as filters for the information retrieved:
Journal: Different journals have different standards in the review process and the completeness of the published work. As a measure of the quality of each journal, we provide the journal h-index, as well as the SCImago Journal Rank (SJR) indicator. Authors: An expert can be interested in publications by universities or specific authors that can be known for their overall contribution to the field. Thus, filtering can be applied by author name or affiliation. Publication type: Different types of articles are defined according to the different levels of evidence (e.g., scientific review or clinical trial) based on which the represented knowledge is derived. Accordingly, the type of publication is also provided to allow for relevant filtering. Publication year: The age of a publication allow an expert to decide if the results depicted in the publication are up-to-date. Therefore, the publication year, as another useful filter for the end-users, is also provided. Cited By Count: The number of citations for a specific publication can provide a good indication of its quality and trustworthiness.
Table 3 reports the number of annotations from UMLS extracted by the Natural Language Processing techniques implemented in the Scholarly Data Ecosystem. These DE4LungCancer classes correspond to relations in the UMLS Semantic Network.29
Number of UMLS annotations
Out of 1,550,586 drug–drug interactions (DDI) collected from DrugBank, 320 patterns were recognized to evaluate the performance of FALCON in this use case, twelve annotators manually annotated 1,198 DDI descriptions; annotations correspond to Concept Unique Identifiers (CUIs) from UMLS and constitute the gold standard of the evaluation. For example, for the DDI description: “The serum concentration of Lepirudin can be decreased when it is combined with Tipranavir”; Lepirudin and Tipranavir correspond to the extracted entities from the above record, while decrease and serum concentration represent, respectively, the effect and impact of the interaction of Tipranavir with Lepirudin. A 2-fold cross-validation was followed while building the gold standard, and a majority voting solved disagreement. The evaluation indicates a precision of 98%. The 2% where FALCON failed to extract and link the terms correctly are interactions that contain more than one interaction in the same sentence, where FALCON was only considering one interaction. Additionally, the EABlock toolbox has been assessed in Baseline and EABlock pipelines. Three sets of RML mapping rules were evaluated on two datasets of biomedical concepts composed of 10K and 20K entities, respectively. Both pipelines generate the same KGs. However, in Baseline, NER and EL are performed in a pre-processing stage of KG creation, while EABlock functions were executed with the RML rules in the EABlock pipeline. Observed execution time suggests that using the EABlock functions speeds up the KG creation process by up to 40%. Moreover, we created five gold standard datasets considering textual values with frequent quality issues that frequently exist in textual values datasets (e.g., character capitalization, elimination, insertion, and replacement). These datasets are built from DBpedia, Wikidata, and UMLS. The errors are introduced with a certain percentage of the records (i.e., 50% and 80%). The EABlock functions exhibited a F1 score that varied from 0.78 in DBpedia, 0.88 in UMLS, and 0.99 in Wikidata. Table 4 reports on the number of links recognized by EABlock during the execution of the RML + FnO mapping rules that define the DE4LungCancer KG. In total, 12,961, 11,679, and 8,172 distinct DE4LungCancer entities are connected to UMLS, DBpedia, and Wikidata30
Note that an entity (e.g., a drug) may belong to various DE4LungCancer classes, this explains why the total numbers of links do not correspond to the sum of the number of links of each DE4LungCancer class.
Number of links from the DE4LungCancer KG to UMLS, DBpedia, and Wikidata
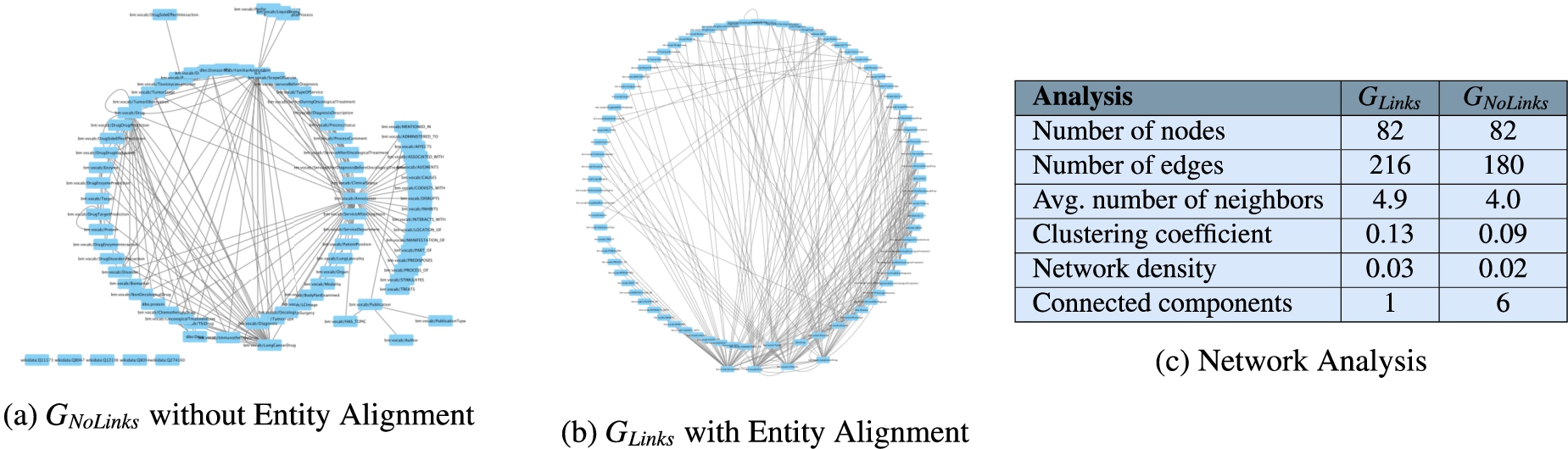
Network analysis to assess connectivity of
The annotations from UMLS, DBpedia, and Wikidata extracted by the NLP techniques implemented in DE4LungCancer enable the creation of entity alignments that define the semantic data integration process executed during the creation of the DE4LungCancer KG. This section presents the impact that these annotations have on semantic data integration. This impact is measured in terms of connectivity or the number of alignments they enable to establish in the DE4LungCancer KG. Two versions of KGs are created:
Assessment of the impact of interaction between drugs in the effectiveness of the lung cancer treatments
The knowledge represented in the DE4LungCancer KG is exploited to understand the impact of the interactions between a treatment’s drugs on the effectiveness of the treatment. The evaluation of treatments’ effectiveness is performed based on the number of toxicities observed in the lung cancer patients and the assessment of a treatment’s response provided by the patients’ oncologists; these results are part of the clinical records processed by the Clinical DE and integrated into the DE4LungCancer KG. The DDIs in a treatment are computed based on three computational methods. The first method (DrugBank) computes the number of DDIs in treatment based on the DDIs reported on DrugBank. We extracted the DDIs from DrugBank and included them in our DE4LungCancer KG. The second method (DS) proposed by Rivas and Vidal [46] deduces new DDIs based on a deductive system implemented in Datalog on top of KG. DS is defined in terms of Datalog rules, and it exploits the fine-grained representation of the DDIs generated by FALCON. The third method (Literature) proposed by Bougiatiotis et al. [8] predicts DDIs based on the Scholarly DE. This method analyses the paths connecting interacting and non-interacting drug pairs in this DE Knowledge Graph and trains a machine learning algorithm (Random Forest) to discriminate between those two classes. Based on the trained model, we then apply predictions to all non-interacting pairs to identify potential DDIs that were not previously known based on the resulting prediction confidence scores.

Toxicity analysis of oncological treatments. Figure 9 shows five bar plots of the toxicities produced by treatments in lung cancer patients. The treatment responses are differentiated by color. The oncological treatments with comorbidity drugs generate more toxicities than those without comorbidity drugs.

Distribution of DDIs by treatment response.
The Spearman’s Rho correlation coefficient analysis between DDIs and responses over DE4LungCancer KG
The Spearman’s Rho correlation coefficient analysis between the number of drugs in a treatment and the number of DDIs among these drugs
A dashboard makes the DE4LungCancer KG available to the clinical partners in the lung cancer pilots of iASiS, BigMedilytics, and CLARIFY, and in P4-LUCAT. Various services are provided to analyze the processed EHRs and the holistic profiles that integrate EHRs with the fine-grained representation of publications and scientific open data. Those services correspond to dedicated REST APIs that provide an integration point with various dashboard versions. The dashboards are available to the project oncologists via certificate-based authentication. The outcomes of the analytical tools provided by the DE4LungCancer KG services through a dashboard have established the basis for the implementation of clinical interventions for the lung cancer patients treated by the team of oncologists of the Puerta del Hierro University Hospital in Madrid. For example, Fig. 11 illustrates a specific example of results when a clinician queries patients’ length of hospital stay based on their gender. The BigMedilytics dashboard provides a statistical analysis of patient hospitalizations in the first three months, based on gender, retrieved from the corresponding DE4LungCancer KG API service. Figure 12 provides another example, exploring the DE4LungCancer KG through the iASiS dashboard. In that case, a clinician requests all possible drug–drug interactions related to a specific drug (documented in Drugbank, deduced, or predicted).

Clinical KPI results, illustrated through the BigMedilytics dashboard. This example is exploring the proportion of patient hospitalizations in the first three months, divided by gender.

Knowledge graph exploration through the iASiS dashboard. This example is providing all documented, predicted and deduced drug interactions for Vinorelbine.
The improvement of the diagnostic pathway and the reduction in the length of hospital stays and emergency rooms represent the essential clinical requirements identified as KPIs. With this aim, the DE4LungCancer KG can be traversed (e.g., through the BigMedilytics dashboard) to identify the most visited services by patients with a new diagnosis of lung cancer in the previous 15 months to diagnosis; in this analysis, four months before diagnosis to avoid consultations related to the diagnostic process strictly, such as medical oncology. Moreover, the services can be explored to identify the prescribed clinical tests.
To this end, we retrieved all the properties of 1,242 patients from the DE4LungCancer KG; 859 patients visited at least one first-attention service between the day of the diagnosis and 15 months before this date. 459 patients saw first-attention services four and 15 months before diagnosis; 331 were in stage III or IV of lung cancer. During the month before lung cancer diagnosis, which we used as our baseline, the most visited services were: Thoracic surgery, Pneumology, Medical Oncology, Internal Medicine, and Emergency Room. When analyzing 15 months ahead of lung cancer diagnosis, excluding the four months before diagnosis, the top-5 most visited services are General Emergencies, Primary Care, Cardiology, Pneumology, and General and Digestive Surgery. Additionally, we have observed that patients have increased the number of first-attention consultations during the 15 months before lung cancer diagnosis. Moreover, the visited services differ entirely from those seen during the month before a diagnosis of lung cancer. The number of tests is also increased during this period. The hospital services visited for the first time by lung cancer patients are grouped into six categories according to the number of months before the lung cancer diagnosis. These groups are denoted as

Evolution of the top-10 hospital services most visited the first time by lung cancer patients prior to the lung cancer diagnosis. Blue indicates that the number of visits to the service increases, and it moves up in the list. Red shows that the number of visits of the service decreases, and it moves down in the list. White shows a service position stays the same with the respect to the previous reported period.

Comparison of the most visited hospital services in the periods

The Jaccard index values. Overlap of most visited hospital services in the periods
We also compute the Jaccard index to quantify the overlap between sets of services visited in distinct periods; Fig. 15 reports these results. The average Jaccard index is 0.62, indicating a relatively high overlap across the studied periods. In particular, corroborating the clinicians’ hypothesis, that the first attention services visited one and four months before the diagnosis are the same (i.e., Jaccard Index is 1.0) and they may be related to the lung cancer diagnosis. Further, the clinical observation that the first attention services should differ from the ones in 0-1 and 0-4 to posterior periods (i.e.,
DE4LungCancer attempts to address the various requirements (Data Management, Clinical, and Ethical) of Health Ecosystems, as introduced in Section 2.1. In the following paragraphs, we describe how DE4LungCancer tackles the requirements of each category.
Data management requirements
The data management techniques implemented in DE4LungCancer enable uncovering the data management requirements: DR1-Data variety; DR2-Integrity constraint satisfaction; DR3-Transparent data management; and DR4-Unified definition of heterogeneous data. Specifically, disparate data sources have been used in the DE4LungCancer (Requirement
Moreover, the definition of the DE4LungCancer unified schema (through the RDF data model) and the KG creation process using declarative languages (e.g., R2RML, RML, and FnO) for data integration brings significant benefits: They empower the reusability and modularity of the data integration process (Requirement
Clinical requirements
Based on the results of the quantitative analysis conducted on the DE4LungCancer, the clinical partners devised five interventions; with those, they attempted to assess the satisfaction of the DE4LungCancer clinical requirements KPI1, KPI2, KPI3, KPI4, and KPI5. These interventions aim at studying a lung cancer patient at various stages of the lung cancer pathway (

Lung cancer pathway and medical interventions.
Based on the analysis results of the services most frequently visited by patients in the 15 months before diagnosis, first-attention visits to certain services (e.g., General Emergencies, Primary Care, Cardiology, and Pneumology) are considered relevant patterns. As a result, persons who follow these patterns are selected as patients, who may be in an asymptomatic stage and may have the potential risk of developing cancer.
The goal is the identification of people at risk of developing lung cancer and a continuous assessment of a patient’s bypass channels. This intervention has been possible, speeding up appointments for diagnostic tests as well as consultation reviews when it comes to a patient with suspected cancer.
The goal of the intervention is to administrate Palliative Care attention and provide close control in consultations before the next treatment cycle date in a treatment line. This study has allowed for measuring readmission and death at 28 days after discharge to determine the need for external early clinical control. Furthermore, the frequency of this event in advance or initial treatment lines has been assessed. This quantitative analysis indicates that 30% of the patients are over 70 years old; they also suffered from advanced stages of lung cancer and more than three comorbidities. Additionally, they have received more than three lines of oncological treatments. These results are considered as a pattern to promote lung cancer patients to palliative care.
This intervention is defined based on the analysis of the patients who attended General Emergencies and were readmitted, to a new hospital service, in a period of 28 days. The study aims at uncovering combinations of comorbidities and specific treatments that increase the risk of being readmitted to the emergency room. Based on the uncovered patterns, the Oncology Department processes inter-consultations with the departments of the most visited hospital services to identify potential side effects of the prescribed treatments.
General Emergencies have been identified as the most visited medical service once a patient is under follow-up by the Oncology Department. Pain is one of the most common symptoms because pain often changes with disease progression. Despite the importance of pain assessment and management, it is uncovered that pain under treatment is common. Thus, this intervention aims at reinforcing the work of the nurses in assessing pain and favoring early referral to the Pain Unit.
Data sharing, management, and analysis in the DE4LungCancer DE have been conducted following the regulations imposed by Ethical protocol and the Ethical committee of the Puerta del Hierro University Hospital in Madrid. Thus, a legal framework to respect data privacy has been established (Requirement
User acceptance
An initial version of the aforementioned dashboard [33] has been provided to a group of relevant stakeholders to assess its quality and characteristics. Specifically, 44 evaluators have participated in dedicated training and evaluation sessions, running different scenarios, testing the dashboard functionalities, and querying to retrieve data and information of interest. They were then given a questionnaire, asking them to input general profile information and their motivation for utilizing the dashboard. An overview of the basic user groups that evaluated the dashboard can be seen in the results of Fig. 17. The evaluators justified the use of the system and data based on the following reasons:
to analyze the characteristics of special populations for queries related to survival curve analysis; to learn how to use the platform and the various functionalities as a knowledge tool about lung cancer; and to identify some interesting publications and drug interactions
Finally, the evaluators were asked to rate their overall experience with the dashboard (Fig. 18) and report any issues found or propose features that they considered missing.35

The initial question of the evaluation questionnaire, in order to identify the stakeholders’ groups that participated in the training and evaluation sessions.

A general rating from the end-users, evaluating their overall experience with the dashboard.
Conclusions and future work
This paper discusses knowledge-driven data ecosystems (DEs) and their prognostic role in enhancing transparency. DE4LungCancer has been presented as the computational framework to address the data management, clinical, ethical, and legal requirements of the lung cancer pilot of the H2020 EU projects iASiS, BigMedilytics, and CLARIFY, and in the EraMed project P4-LUCAT. DE4LungCancer is a nested framework that comprises three DEs that process and analyze the pilot datasets. DE4LungCancer offers a semantic layer composed of a unified schema, biomedical ontologies, and mapping languages; they provide the basis for transparent data integration into a KG. The hybrid approach that combines the multidisciplinary pilot team with computational tools to validate integrity constraints, the unified schema, and the mapping rules have enhanced the trustability of the outcomes of the analytical services. More importantly, the documentation of the whole process backs up the certification of the process by ethical committees and data protection officers. The project clinical partners can access the DE4LungCancer services through a dashboard. The outcome of the execution of the provided services has enhanced the understanding of the conditions of the hospital services visited by lung cancer patients. Based on the observed results, clinical interventions have been devised. We plan to develop analytical methods to analyze the interventions’ results toward improving the patients’ quality of life.
Footnotes
Acknowledgements
This work has been supported by the EU H2020-funded projects iASiS (GA No. 727658), BigMedilytics (GA No. 780495), the EraMed project P4-LUCAT (GA No. 53000015), and the EU H2020 RIA project CLARIFY (GA No. 875160). Furthermore, Maria-Esther Vidal is partially supported by Leibniz Association in the program “Leibniz Best Minds: Programme for Women Professors”, project TrustKG-Transforming Data in Trustable Insights with grant P99/2020.
Queries to explore RML mapping rules of DE4LungCancer KG
Listing 1 presents a SPARQL query that collects the information about the mapping rules that define the class
Queries to explore the open DE4LungCancer KG
This appendix illustrates some exemplary queries to be executed over the version of the DE4LungCancer KG which is publicly accessible via the SPARQL endpoint. 7 Query in Listing 3 retrieves the drugs that composed a given treatment and the DDIs among these drugs. Query in Listing 4 reports, per treatment, the number of DDIs extracted from DrugBank (?extensionalDDI) and the ones deduced using the deductive system (?intensionalDDI). Additionally, it retrieves the percentage of new interactions deduced by the system. Query in Listing 5 reports, per treatment, the number of DDIs extracted from DrugBank (?extensionalDDI) and the ones from literature (?intensionalDDI). Additionally, it retrieves the percentage of new interactions predicted by the literature. Query in Listing 6 retrieves per treatment the drugs that compose the treatment and the number of DDIs that have been reported in DrugBank for the drugs of the treatment. Query in Listing 7 compares the number of drugs per treatment, and the DDIs extracted from DrugBank and the ones inferred using our proposed deductive system. Query in Listing 8 retrieves the drugs that are more commonly used in the lung cancer treatments. We can observe that Omeprazole, Carboplatin, Atorvastatin, Enalapril, and Simvastatin are the top-5 most prescribed drugs in the therapies registered in the clinical data.
