Abstract
Social networks have become information dissemination channels, where announcements are posted frequently; they also serve as frameworks for debates in various areas (e.g., scientific, political, and social). In particular, in the health area, social networks represent a channel to communicate and disseminate novel treatments’ success; they also allow ordinary people to express their concerns about a disease or disorder. The Artificial Intelligence (AI) community has developed analytical methods to uncover and predict patterns from posts that enable it to explain news about a particular topic, e.g., mental disorders expressed as eating disorders or depression. Albeit potentially rich while expressing an idea or concern, posts are presented as short texts, preventing, thus, AI models from accurately encoding these posts’ contextual knowledge. We propose a hybrid approach where knowledge encoded in community-maintained knowledge graphs (e.g., Wikidata) is combined with deep learning to categorize social media posts using existing classification models. The proposed approach resorts to state-of-the-art named entity recognizers and linkers (e.g., Falcon 2.0) to extract entities in short posts and link them to concepts in knowledge graphs. Then, knowledge graph embeddings (KGEs) are utilized to compute latent representations of the extracted entities, which result in vector representations of the posts that encode these entities’ contextual knowledge extracted from the knowledge graphs. These KGEs are combined with contextualized word embeddings (e.g., BERT) to generate a context-based representation of the posts that empower prediction models. We apply our proposed approach in the health domain to detect whether a publication is related to an eating disorder (e.g., anorexia or bulimia) and uncover concepts within the discourse that could help healthcare providers diagnose this type of mental disorder. We evaluate our approach on a dataset of 2,000 tweets about eating disorders. Our experimental results suggest that combining contextual knowledge encoded in word embeddings with the one built from knowledge graphs increases the reliability of the predictive models. The ambition is that the proposed method can support health domain experts in discovering patterns that may forecast a mental disorder, enhancing early detection and more precise diagnosis towards personalized medicine.
Keywords
Introduction
The COVID-19 pandemic has considerably burdened mental diseases all over the world [60], and eating disorders (EDs) are not an exception [16,46,62,81]. EDs are health conditions severely disturbing eating behaviors and related thoughts and emotions. A recent study by Zipfel et al. [81] reveals that EDs increased during the pandemic by 15.3% in 2020 with respect to previous years. Moreover, outcomes from a systematic literature review by McLean et al. [43] uncover that children and adolescents are the most vulnerable groups impacted by EDs during the COVID-19 pandemic. This burden of mental incidences raises awareness of the need for early detection mechanisms to effectively take action in clinical and healthcare services. More importantly, these studies provide evidence of the urgent need for scalable methods for effectively supporting communities increasingly suffering from mental disorders.
Social media is increasingly used as a dissemination channel to announce novel treatments or conditions to respond to health-related problems and even to discuss natural disasters [64]. Moreover, social media networks are utilized to promote or prevent the administration of certain interventions, e.g., in mental conditions like eating disorders [9,39]. Furthermore, Artificial Intelligence (AI) has gained momentum in the healthcare field [8,80], and AI-based solutions have been developed for disease prevention [17], pathology detection [73], and treatment prescription [57]. Analyzing the discourse on social networks such as Twitter can help to find answers to relevant problems by applying various ML techniques. Specifically, in mental health, predictive models have successfully been applied [12,63]. Exemplary contributions include the detection of depression [29], suicidal mental tendencies [21], bipolar disorders [24] and other mental disorders [20]. Moreover, the models proposed by [38] can uncover patterns of anorexia in datasets obtained from social networks. These results put in perspective the role of automatic classification in the scalable and effective detection of patterns of communities suffering from mental disorders [75].
Knowledge graphs (KGs), and Semantic Web technologies in general, have been accepted as data structures that enable the natural representation and management of the convergence of data and knowledge [22]. The information contained in knowledge graphs is increasingly used in the scientific community to solve different problems [1,3,67]. Specifically, community-maintained knowledge graphs such as Wikidata [71] or DBpedia [33] represent rich sources of structured knowledge not only from general domains but also in biomedicine [10,30,79]. Figure 1(b) depicts the contextual knowledge obtained through structured data in a portion of Wikidata. We define this contextual knowledge as entities related to a given resource within the knowledge graph, e.g., the Wikidata resource
Predictive models can also be built over unstructured data, Bidirectional Encoder Representations from Transformers (BERT) models [15] are exemplary solutions to this problem [2]. Contrary to other models (e.g., BiLSTM-based-Bidirectional Long Short-Memory), BERT models can learn from words in all positions, i.e., from the whole sentence. Figure 1(a) illustrates an example of contextual knowledge extracted from unstructured information for the term “anorexia”. Here, this contextual knowledge is based on the information around the word anorexia within a given sentence. These models usually receive texts and labels as input to achieve predictive models that classify texts. By applying this model, vector embeddings are also obtained, but in this case, the encoded contextual content encodes unstructured data (lower part of Fig. 1(a)). BERT models have exhibited high performance in almost any prediction problem where contextual knowledge is extracted from text. Nevertheless, albeit the large number of covered domains and high-quality predictions, BERT models may perform poorly over short texts [35,78].
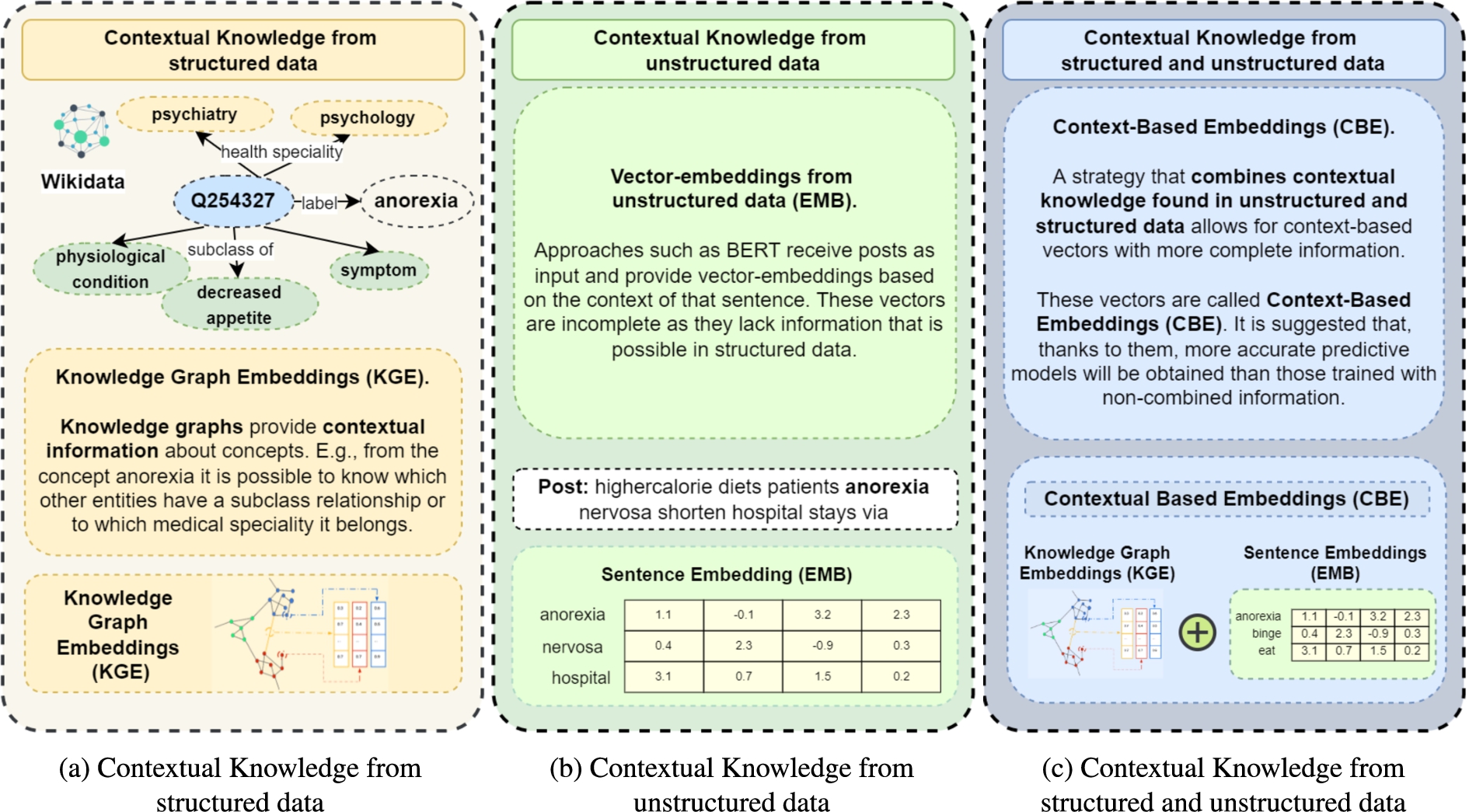
Contextual knowledge (CK). (a) CK is represented in knowledge graphs and encoded in vector embeddings KGE methods. (b) CK is extracted from unstructured data, e.g., social media posts; it corresponds to the words around a particular concept. (c) The proposed approach, CK extracted from both structured and unstructured data and encoded in context-based embeddings (CBE).
The resulting embeddings are used in various state-of-the-art predictive models. The methodology followed to implement the proposed approach is composed of the following steps:
The remainder of this paper is organized as follows. Section 2 presents the fundamentals of RDF, knowledge graphs, knowledge graph embeddings, entity recognition, and entity linking. Section 3 analyzes related approaches from the state of the art. Then Section 4 defines our problem statement and states our proposed solution. This section also describes the main components of the architecture that implements our proposed method. Section 5 reports on the experimental evaluation and discusses the observed outcomes. Finally, we close the paper in Section 6 with conclusions and an overview of future work.
Wikidata – Community-maintained knowledge graph
In this research, we use community-maintained knowledge graphs and, more specifically, Wikidata, to obtain information related to the concepts contained in short texts. The Wikidata knowledge graph is stored internally in JSON format and can be edited by any user thanks to the web interface of this knowledge graph. Wikidata can be downloaded in RDF, however, a subject-property-object triple is annotated with qualifiers [41], representing metadata about the triple. Lastly, although Wikidata is a source of encyclopedic knowledge, Waagmeester et al. [72] report the main characteristics of Wikidata as a relevant provider of knowledge from Life Sciences.
Entity recognition and entity linking, making use of knowledge graphs
In our approach, we use named entity recognition and linking tools to extract the entities found in the short texts used as input data and link them to the concepts contained in Wikidata. A named entity linker implements the tasks of named entity recognition (NER) and named entity linking (NEL), allowing the identification of entities in a text and their corresponding resources in a knowledge graph or controlled vocabulary. Although our approach is named entity linker agnostic, as a proof of concept, we make use of the EntityLinker1
Knowledge graph embeddings (KGE) are low-dimensional representations of the entities and relationships that make up a knowledge graph. KGEs make possible a generalizable representation of an entity based on its context on a global knowledge graph, in our case, Wikidata. This context allows inferring relationships between concepts. The KGEs provide us with information about the relationships between different terms related to our problem. For example, in an ED context, KGEs provide us with the relationships between ‘green tea’, ‘day’, ‘good’, ‘anorexic’, ‘binge-eating’ and ‘nah’. Thanks to the magnitude of information contained in Wikidata, the KGEs provide information about the interactions between the concepts contained in the short texts we want to classify.
This information, added to the information found in the tweets themselves, is of vital importance to improve the predictive models that classify these short texts. There are many configurations and methods for calculating these knowledge graph embeddings. In our approach, we use of RDF2Vec [56]. Inspired by Word2vec [44] which represents words in vector space, RDF2Vec applies this method within a knowledge graph. RDF2Vec allows receiving different ways to create sequences of RDF nodes that are then used as input for the Word2vec algorithm. One of the most commonly used strategies is random walks in an RDF graph.
Extracting knowledge graph embeddings through RDF2Vec
RDF2Vec, as proposed by Ristoski et al. [56], adapts the language modeling approach in Word2vec for latent representations of entities in RDF graphs. Ristoski et al. [56] demonstrate that projecting such latent representations of entities into a lower dimensional feature space shows that semantically similar entities appear closer to each other in the knowledge graph where these entities are represented. The following steps are proposed to generate RDF2Vec embeddings from a corpus of short texts:
The entities of each of the texts in the dataset are obtained, as explained in Section 2.2. The set of unique entities is used to generate a vocabulary. An extension of RDF2Vec (implemented by the authors) is used to generate knowledge graph embeddings of a set of entities. This procedure calculates the embeddings based on the entities that are in the neighborhood of each entity within Wikidata. This enables us to obtain word vectors based on the contextual knowledge within the entities found in the collected dataset.
Strategies for combining embeddings
In the proposed approach, once the knowledge graph embeddings are obtained, it is necessary to combine them, as the number of entities in each tweet may not always be the same and, therefore, the number of KGEs may also be different. As can be seen in the scientific literature, there are many methods of combining embeddings [4,34,52,54], the simplest of which is to find the average of the total number of embeddings obtained in each tweet. However, this does not seem to be the best strategy. In our approach, we have used smooth inverse frequency (SIF) [4]. SIF also considers the most meaningful words within a sentence; it scores from 0 to 5 pairs of sentences, in our case tweets, according to the similarity between them. SIF is used as it is one of the most widely used strategies [49,76].
Related work
Deep learning in social media data
The use of social media data to train predictive models that use machine learning and deep learning techniques is becoming increasingly common in the scientific field. A search for the terms “social media data AND deep learning” in Google Scholar returns approximately 225,000 results since 2017.3
Knowledge graphs such as Wikidata [71] and DBpedia [33] are increasingly used in research. Moreover, studies have shown that the information they contain, is useful and reliable for use in some fields like the Life Sciences [72]. One of the most important problems scientists face in acquiring knowledge through these collaborative knowledge networks is what is known as Entity Linking. In order to obtain a resource that represents the concept “help” from Wikidata, it is necessary to know whether we are referring to the concept of help as cooperation between people4
The contextual knowledge found in knowledge graphs combined with the use of deep learning techniques is being used to solve different research problems. For example, thanks to the semantic information obtained through these knowledge graphs, it is possible to improve the interpretability and explainability of different predictive models powered by deep learning techniques [36]. Improving the interpretability and explainability of predictive models is important, especially when deployed in domains related to health or education. Moreover, knowledge graphs are also being used to help predict relationships between different concepts, such as, the gut microbiota and mental disorders [18] Notwithstanding, there is a lack of studies that have used the contextual knowledge extracted from knowledge graphs combined with the contextual knowledge extracted from unstructured data to classify texts by making use of this information. Our study aims to demonstrate that incorporating semantic enrichment to texts can contribute to improve the performance of predictive models.
Combining contextual knowledge from unstructured and structured data
In this research, we address the problem of effectively classifying short text. We propose a hybrid method based on the combination of contextual knowledge from structured data and unstructured data, resulting in contextual based embeddings. We elaborate below on the problem statement and the proposed solution.

Motivating example. (a) 2 labeled posts, (b) a text classification model using BERT models approach, (c) short text classification using the novel approach making use of context-based embeddings (CBE).

Architecture of the proposed approach. It receives a corpus of posts and knowledge graphs. The first component, called context-based embeddings encoding, represents how unstructured based data is combined with the information obtained through knowledge graphs applying entity recognition and linking to concepts within a knowledge graph. The knowledge graph-based data module shows the process of knowledge graph embeddings (KGEs) extraction from a knowledge base. Then, sentence embeddings from unstructured data are combined with KGE. The second component, called context-based embeddings, is computed, and predictive machine learning models are executed with these embeddings in decoding, resulting in accurate classification models for posts.
This component has three main objectives: (i) to obtain contextual knowledge through unstructured data (sentence embeddings), (ii) to obtain contextual knowledge through structured data (knowledge graph embeddings), and (iii) to combine this textual knowledge into a single vector structure through the combination of both (context-based embeddings). Objectives (i) and (ii) are achieved through modules that are executed in parallel. Objective (iii) is attained once the two previous modules have been executed. The properties of the three modules that make up this component are detailed below.
The recognized entities are: [‘kilocalorie’,‘diet’,‘patient’,‘anorexia’,‘nervosa’]
And the identifiers of these entities in Wikidata are: [‘ Q26708069’, ‘ Q474191’, ‘ Q181600’, ‘ Q254327’, ‘ Q131749’]
This component uses the information contained in the context-based embeddings generated by the first component by decoding them. According to the hypothesis put forward in this study, the complete contextual knowledge contained in these vectors should provide predictive models with a higher degree of accuracy. This knowledge allows the vectors to contain a more accurate inter-post similarity function than vectors generated from unstructured data alone or from unstructured data. In the running example, “anorexia” was close to “nervosa” in the vector obtained from the unstructured data, but not close to “symptom”. The combination of both types of embeddings encodes different contextual knowledge; it offers a contextual-based enhanced vector.
The experiments conducted in this study demonstrate that machine learning models enhanced with contextual knowledge can improve accuracy in predictive short text classification models. The objective of our particular case study is to improve the classification of eating disorders in social media posts. The empirical study sets out to answer the following research questions:
What is the impact of contextual knowledge extracted from structured data on the performance of the model?
Do BERT pre-trained models based on short texts perform better than other pre-trained models?
What type of contextual information provides the most accurate predictive models – structured, unstructured or a combination of both?
Data availability of the experiments and code of the empirical evaluation of the proposed approach8
The following settings are configured to answer our research questions.
Formulas of the evaluation metrics used
Formulas of the evaluation metrics used
The different steps taken to validate the model proposed in this research using a dataset of our own are detailed below. The working pipeline is divided into 4 phases:
The phases mentioned above are explained in detail in the following subsections.
Collecting and labeling ED dataset
Figure 4 depicts the process followed to collect and label the tweets that compose our corpus.

Data collection and data labeling steps. First, a set of tweets were collected through a keyword-based search. Subsequently, a subset of the tweets was manually labeled into four binary categories.
We employ information-filtering-based methods of social media posts using a set of keywords as search queries. This method of collection allows for faster and less costly access to a bigger volume of data and with a specialized level of detail as to the desired output when compared to what could be obtained through traditional collection techniques such as clinical screening surveys [48,74]. Nonetheless, the datasets produced this way come with their limitations due in great part to the pervasive noise associated with their source, which may impact the quality of the data itself, as well as the reliability and representativeness of the results obtained [5,48].
Our chosen social media platform is Twitter, and we perform the capture of tweets using the T-Hoarder tool [11]. The list of hashtags used to filter tweets is anorexia, anorexic, dietary disorders, inappetence, feeding disorder, food problem, binge-eating, eating disorders, bulimia, food issues, and loss of appetite. These hashtags were selected taking into account the most relevant keywords that yielded tweets about eating disorders in other research [23,25,40,65]. However, this sub-selection could still bias the sampling and results, as we might be making assumptions about the behavior of the users generating the content or overlooking relevant information outside these limits [48]. The collection yielded the capture of 494,025 tweets, and to mitigate issues associated with lexical and semantic redundancy of the content [48], a process of cleaning (removal of duplicates, re-tweets, re-shared content), manual curation and annotation were performed, thus, resulting in the creation of a subset of 2,000 tweets.
To annotate the tweets, we first determine four binary classification problems of interest guided by preexisting ED research [25,40,65]. The resulting categories for each problem are detailed below, followed by the labeling criteria of each tweet:
This dataset is available for download from github [7].
As per the characteristics of the data and the research itself, it is not our objective to profile the sampled users, thus we avoid making assumptions based on ‘gender’, ‘nationality’, ‘socio-economic background’, ‘age’, to name a few features, that could lead us to fall back on perpetuating damaging stereotypes associated with persons who suffer eating disorders, as recent research has shown that they are global illnesses that do not discriminate [47,53]. However, all the collected tweets belong to users that express themselves in English, a result of the predetermined search criteria. Additionally, as part of the 2,000 tweets, 1,567 belong to unique Twitter accounts, meaning that 433 tweets are concentrated in 183 Twitter accounts, a figure that could indicate there is no presence of over-representation of ‘noisy’ social media users, notwithstanding the possibility that this distribution includes accounts that belong to nonhumans (i.e., bots, corporate accounts), multiple users posting from the same account, or the same entity posting from different accounts [6,48].
Through an exploration of the frequency of terms associated with the hashtags collected, we managed to identify a predominant mention of anorexia and other derivative terms such as “anorexia”, “anorexic”, “proana”, “ana”, “anatwt”, “anorexiatips”, (1,034 associated hashtags), while an underrepresentation of hashtags associated with other eating disorders, such as bulimia (182 associated hashtags) is identified. This pattern is also replicated in the analysis of the content of the tweets themselves, as the most frequent terms were also associated with anorexia (more than 428 unigram tokens) over bulimia (109 tokens), a small caveat regarding this will be that binge (282 tokens) eating, is a behavior that could both be identified as a symptom of bulimia or a disorder in itself. While this detangling has not been the focus of this study, the results of this exploration leave the door open to perform further captures that could cover a wider variety of eating disorders or for a fine-grained analysis that could help semantically differentiate these disorders in short text analysis. Figure 5 presents a summary of the data analysis. Most frequent terms with unigram tokens (see Fig. 5(b) and Figure 5(a)) and a breakdown of the annotated categories and the corresponding class count is showed in Fig. 5(c). The top 35 hashtags show a predominant mention of ‘anorexia’ and other derivative terms such as “anorexia”, “anorexic”, “proana”, “ana”, “anatwt”, “anorexiatips”.

Results of the analysis of the dataset, showing the most frequent terms and the distribution of the categories.
Emojis were eliminated from the tweets in the corpus. The following analysis provides evidence of the lack of effect of the emojis’ removal.
The frequencies of the different emojis in the texts were obtained, and it was determined that only 17.9% of texts contained some emoji, i.e., 359 of the 2,000 total texts.
Statistical calculations were carried out on the distributions of emojis according to the four categorizations made. To statistically analyze these data, two statistical techniques were used: (i) overlap analysis and (ii) Spearman’s rho correlation.
It was observed that, in all cases, the similarity between the two distributions was statistically significant. The correlations are shown in Table 2. Note that in category ED III, the Spearman’s rho value is lower because 8 emojis were used more than 90% of the time in tweets categorized as non-informative, compared to those categorized as informative. This difference makes sense, since, when an opinion is expressed, there are emojis that relate more to the feelings of that opinion, and informative tweets are more objective.
Therefore, in line with what has been done in other similar research [55,66], it was decided to eliminate emojis in the pre-processing of the texts.
Frequency of emojis in each of the four categories, and Spearman correlation between the distributions for each category
The texts of the dataset were processed with Falcon 2.0 [58] and Entity Linker [26], resulting in a total number of entities in the Wikipedia knowledge graph, as shown in Table 3.
Number of entities, unique entities, and unique entities appearing two or more times obtained from Wikidata for each of the datasets used
Number of entities, unique entities, and unique entities appearing two or more times obtained from Wikidata for each of the datasets used
From the total number of entities, the total number of unique entities was calculated, and a dataset was obtained consisting of those entities that appeared at least twice in each of the tweets. By manually reviewing the entities obtained in the datasets, some disambiguation errors were detected and manually corrected (Table 4(a)).
In addition, several concepts were added to Wikidata, shown in Table 4(b). After obtaining all the entities, the knowledge graph embeddings were collected using the pyRDF2Vec tool [69] (setting
To validate the proposed approach, 144 tests are carried out, from which the evaluation metrics indicated above, accuracy and f1-score, are obtained. These 144 tests are divided as follows:
Dataset properties in the classification outcome
To verify that our model is not dependent on the dataset used, an analysis has been carried out on the original dataset and on the outputs obtained after applying the different models. This analysis consists of obtaining the Spearman correlation between the distributions of the most frequent terms found in tweets labeled as “0” or “1” in each category in each of the tests performed. A correlation close to 1.0 with a p-value of less than 0.05 would indicate that the distributions of the most frequent terms analyzed are statistically similar, thus being a valid and unbiased dataset for training the models. Likewise, keeping the same distribution at the output may suggest that the models do not increase or decrease the possible bias that could be present in the dataset.
After analyzing the output of the 12 models, applied on the 3 different input datasets, in the 4 tested categorizations, it has been observed that the distributions of the most frequent terms in the dataset are similar in the 144 outputs of the analyzed algorithms with a p-value < 0.001 in all cases, the maximum correlation was 1.0 and the minimum 0.996. This means that the models do not amplify the possible bias of the dataset.
Discussion of observed results
Twelve different classification models to classify tweets in four binary categories are compared using three different approaches: (i) contextual knowledge extracted from unstructured data, (ii) contextual knowledge extracted from structured data (KGE), and (iii) the approach proposed based in contextual knowledge extracted from structured data combined with contextual knowledge extracted from unstructured data (context-based embeddings, CBE). The results are shown in Table 5.
Results after applying 12 machine learning models using 3 different input variables: texts, knowledge graph embeddings, and the context-based embeddings obtained using our approach. The best results are highlighted in bold
Results after applying 12 machine learning models using 3 different input variables: texts, knowledge graph embeddings, and the context-based embeddings obtained using our approach. The best results are highlighted in bold
These results are obtained after training twelve different machine learning and deep learning models to categorize tweets in different categories using three different inputs for our dataset about eating disorders which consist of: (i) 2,000 labeled tweets, (ii) 2,000 knowledge graph embeddings and (iii) context-based embeddings obtained through our proposed approach.
The models trained with the context-based embeddings (CBE) dataset are the ones that obtain the best results in 97.5% of the cases. The highest percentage improvement in the
There are only two cases in which the
From the analysis of the results, it is clear that the use of contextual knowledge extracted from structured has a positive effect on the performance of the generated predictive models. Of all the experiments carried out over a dataset related to eating disorders, in 97.5% of the cases, the experiments carried out using those with knowledge graph exploitation as input data obtained better results. In the remaining 2.5% of cases, the model is equally effective or slightly less effective, not exceeding 1% worse than the model using data without KGE.
Comparing the performance results obtained by each of the seven pre-trained BERT models, the results suggest that the pre-trained model with short texts, TweetBERT, is the one that offers the best results in the metrics evaluated against its opponents in all but one occasion. These results suggest that pre-trained models with short texts may perform better than others.
It is possible to highlight that the approach proposed in this research, based on the combination of contextual knowledge obtained through structured and unstructured information, offers better results than the other two approaches studied. These results suggest that the contextual content that can be obtained from structured information, combined with the textual content of unstructured information, can help us generate more accurate predictive models. This improvement in the accuracy of models may contribute to assist some tasks of early disease detection.
We address the problem of improving the performance of predictive models generated through machine learning techniques for short texts classification proposing a hybrid framework based on the combination of contextual knowledge from unstructured and structured data sources. Given the importance of these text classification models in different scientific and industrial contexts, any contribution that improves the performance of these predictive models is of worldwide interest and, more specifically, in the health field. Thanks to the improvement of these text classifiers, it is possible to detect and diagnose diseases earlier so that interventions can be planned to help reduce the number of patients who may have serious consequences due to these diseases.
This research presents a hybrid approach using a combination of contextual knowledge from unstructured data (e.g., BERT) and contextual knowledge from structured data (e.g., knowledge graphs), obtaining contextual knowledge embeddings. One of the objectives achieved with these embeddings is to improve classification models in short texts. It has been demonstrated on a real health-related problem over a dataset about eating disorders. The tests to validate the proposed approach show that the models trained with contextual-based embeddings have a higher success rate than those obtained with models trained only with contextual knowledge from structured or unstructured data. As a result, 97.5% of the trained models perform better with our approach than the usual ones. It is possible to highlight that using semantic information in knowledge graphs can improve the performance of predictive models in natural language processing and text mining, with the corresponding importance in the health sector. Thus, the proposed method contributes to the portfolio of tools to support understanding the disorders that social media users may suffer. Given the urgent need to help these communities, we hope these results will motivate using our proposed methods in other health-related classification problems.
Given that the approach presented in this research has been tested on a dataset within a particular health domain, eating disorders, further testing is needed to bring more robustness to the validation of the approach. In the future, we will test on a dataset with a larger amount of data to provide further validity, and we will test this approach by adding the information of emojis by combining other Knowledge Bases such as, e.g., Emojinet or Emojipedia. Finally, creating a framework that applies our approach to a given dataset is on our future agenda.
Footnotes
Acknowledgements
Part of this research was funded by the European Union’s Horizon 2020 research and innovation programme under Marie Sklodowska-Curie Actions (grant agreement number 860630) for the project “NoBIAS – Artificial Intelligence without Bias”. This work reflects only the authors’ views, and the European Research Executive Agency (REA) is not responsible for any use that may be made of the information it contains. Furthermore, Maria-Esther Vidal is partially supported by Leibniz Association in the program “Leibniz Best Minds: Programme for Women Professors”, project TrustKG-Transforming Data in Trustable Insights with grant P99/2020.
