Abstract
Knowledge graphs are freely aggregated, published, and edited in the Web of data, and thus may overlap. Hence, a key task resides in aligning (or matching) their content. This task encompasses the identification, within an aggregated knowledge graph, of nodes that are equivalent, more specific, or weakly related. In this article, we propose to match nodes within a knowledge graph by
Introduction
The Semantic Web [3] offers tools and standards that facilitate the construction of knowledge graphs [17] that may aggregate data and elements of knowledge of various provenances. The combined use of these scattered elements of knowledge allows access to a larger extent of the available knowledge, which is beneficial to many applications, such as fact-checking or query answering. For this conjoint use to be possible, one crucial task lies in
In the present work, we focus on matching specific nodes within an aggregated knowledge graph represented within Semantic Web standards. We view such a knowledge graph as a directed and labeled multigraph in which nodes represent entities of a world – also named individuals – (

Outline of our approach. Gold clusters are computed from existing alignments between the nodes to match in the knowledge graph (
We propose to match specific individuals that represent n-ary relationships through an approach that combines graph embedding and clustering, outlined in Fig. 1. Graph embeddings are low-dimensional vectors that represent graph substructures (
To suggest alignment relations from node embeddings, we apply a clustering algorithm on the embedding space and consider nodes that belong to the same cluster as similar. The resulting clusters are evaluated by comparison with
Within our approach, we particularly investigated the interplay between GCNs and domain knowledge through the two following aspects. First, similarly to existing works with different embedding models [18], we applied various inference rules associated with domain knowledge (
Our approach based on GCNs was motivated by the need to align pharmacogenomic (PGx) knowledge that we previously aggregated in a knowledge graph named PGxLOD [22]. The biomedical domain of PGx studies the influence of genetic factors on drug response phenotypes. As an example, Fig. 2 depicts the relationship

Representation of a PGx relationship between gene
The remainder of this paper is organized as follows. In Section 2, we outline some works related to node matching in knowledge graphs and graph embeddings. We detail the core of our matching approach (node embeddings and clustering) in Section 3, and how inference rules associated with domain knowledge are considered in Section 4. In Section 5, we conduct experiments with this approach on PGxLOD, a large knowledge graph we built that contains 50,435 PGx relationships [22]. Finally, we discuss our results and conclude in Section 6 and 7.
Matching
Numerous papers exist about knowledge graph matching. The interested reader could refer to the book of Euzenat and Shvaiko [12] for a formalization of the matching task, and a detailed presentation of the main methods. In the following, we focus on graph embedding techniques. Such techniques have been successfully applied on knowledge graphs for various tasks such as node classification, link prediction, or node clustering [5,32]. Interestingly, the task of matching nodes can be alternatively tackled as a link prediction task (
Graph embedding
Existing papers about graph embedding differ in the considered type of graphs (
Graph Convolutional Networks (GCNs)
The approach adopted in this article is based on Graph Convolutional Networks (GCNs). GCNs have been introduced for semi-supervised classification over graphs [20] and extended for entity classification and link prediction in knowledge graphs [29]. In contrast with TransE and RDF2Vec that work at the triple and sequence levels, GCNs compute the embedding of a node by considering its neighborhood in the graph. Hence, as aforementioned, we believe GCNs are well-suited for our application of matching reified GCNs and the Soft Nearest Neighbor loss are further detailed in Section 3.2.
However, previous methods do not consider inference rules associated with domain knowledge represented in knowledge graphs on the contrary of recent papers [27]. For example, Iana and Paulheim [18] evaluate the RDF2Vec embedding model when inferred triples associated with subproperties, symmetry, and transitivity of predicates are added to the knowledge graph. Interestingly, the addition of inferred triples seems to degrade the performance of RDV2Vec embeddings in downstream applications (
These related works and our preliminary results [23] inspired the present work where we investigate how
Matching nodes with Graph Convolutional Networks and clustering
Approach outline
Our approach is outlined in Fig. 1.
It takes as input an aggregated knowledge graph
Learn embeddings for all nodes in
Apply a clustering algorithm only on the embeddings of nodes from
It should be noted that gold clusters can result from another automatic matching method or a manual alignment by an expert. For example, in Section 5, our gold clusters are computed from alignments semi-automatically obtained with rules manually written by experts [21]. These alignments can use different alignment relations (
Learning node embeddings with Graph Convolutional Networks and the Soft Nearest Neighbor loss
To learn embeddings for all nodes in
Graph Convolutional Networks (GCNs) can be seen as a message-passing framework of multiple layers, in which the embedding
The number of predicates in
Recall that our objective is to cluster similar nodes, which differs from previous applications of GCNs ( A set A set A temperature Embeddings
Minimizing the SNN loss corresponds to minimizing intra-cluster distances and maximizing inter-cluster distances for the gold clusters of nodes in
The computation of
This step enables to learn embeddings for all nodes in
Matching nodes by clustering their embeddings
After embeddings of all nodes in the graph have been output by the last layer of the GCN, we perform a clustering on embeddings
We conduct comparative experiments with three distinct clustering algorithms presented in Table 1, in regards with three classical metrics presented in Table 2. Within the large set of existing algorithms, our choice has been guided by the constraints of our task that requires to handle an important number of clusters, potentially large, and with uneven sizes (see Section 5.1 and Fig. 3 for the sizes of gold clusters computed on PGxLOD). We decided to arbitrarily limit ourselves to three algorithms, but decided to opt for algorithms that cover some diversity in the various family of algorithms. Our three algorithms differ in their parameters: in particular they require either the number of clusters to find (Ward and Single) or the minimum size of clusters (OPTICS). We used our set of gold clusters to set these parameters. Considering these distinct algorithms allows us to evaluate the influence of inference rules in different settings (see Section 4).
Clustering algorithms applied on the embeddings of nodes in S . Nodes that belong to the same predicted cluster are considered as similar
Clustering algorithms applied on the embeddings of nodes in
Performance metrics used to compare the clusters predicted by the algorithms presented in Table 1 with gold clusters
Semantic Web knowledge graphs are represented within formalims such as Description Logics [2] that are equipped with inference rules. Hence, we propose to evaluate the improvements in the results of our matching approach (detailed in Section 3) when considering such inference rules, independently or combined. Here, we only consider the inference rules associated with the following logic axioms: class and predicate assertions, equivalence axioms between entities or classes, subsumption axioms between classes or predicates, and axioms defining predicate inverses. We consider this limited set of inference rules because they are the only ones actionable in PGxLOD, the knowledge graph that motivated our study. Accordingly, we generate six different graphs (
Visual summary of the transformations of
to evaluate the influence of the application of inference rules associated with domain knowledge on node matching.
is the baseline that corresponds to no inference rules being run and the systematic addition of abstract inverses
Visual summary of the transformations of
For a predicate For a predicate Otherwise, for a predicate
We conducted experiments with PGxLOD,2
PGxLOD presents several characteristics needed in the scope of our study. First, PGxLOD contains nodes whose matching is well-adapted to a structure-based approach such as ours. Additionally, alignments are expected to be found between these nodes. Indeed, PGxLOD contains 50,435 PGx relationships resulting from:
an automatic extraction from the reference database PharmGKB; an automatic extraction from the biomedical literature; a manual representation of 10 studies made from Electronic Health Records of hospitals.
Alignments are expected to be found between such relationships since, for example, PharmGKB is manually curated by experts after a literature review. Recall that PGx relationships are
Second, PGxLOD contains
Third, PGxLOD contains subsumption axioms between classes and between predicates, which makes possible the transformations represented in
Alignment relations considered in each gold clustering to compute the gold clusters used in our experiments. We indicate whether a relation is transitive (T or ¬ T) and symmetric (S or ¬ S)
Alignment relations considered in each gold clustering to compute the gold clusters used in our experiments. We indicate whether a relation is transitive (T or ¬ T) and symmetric (S or ¬ S)
Fourth, some PGx relationships in Number of gold clusters (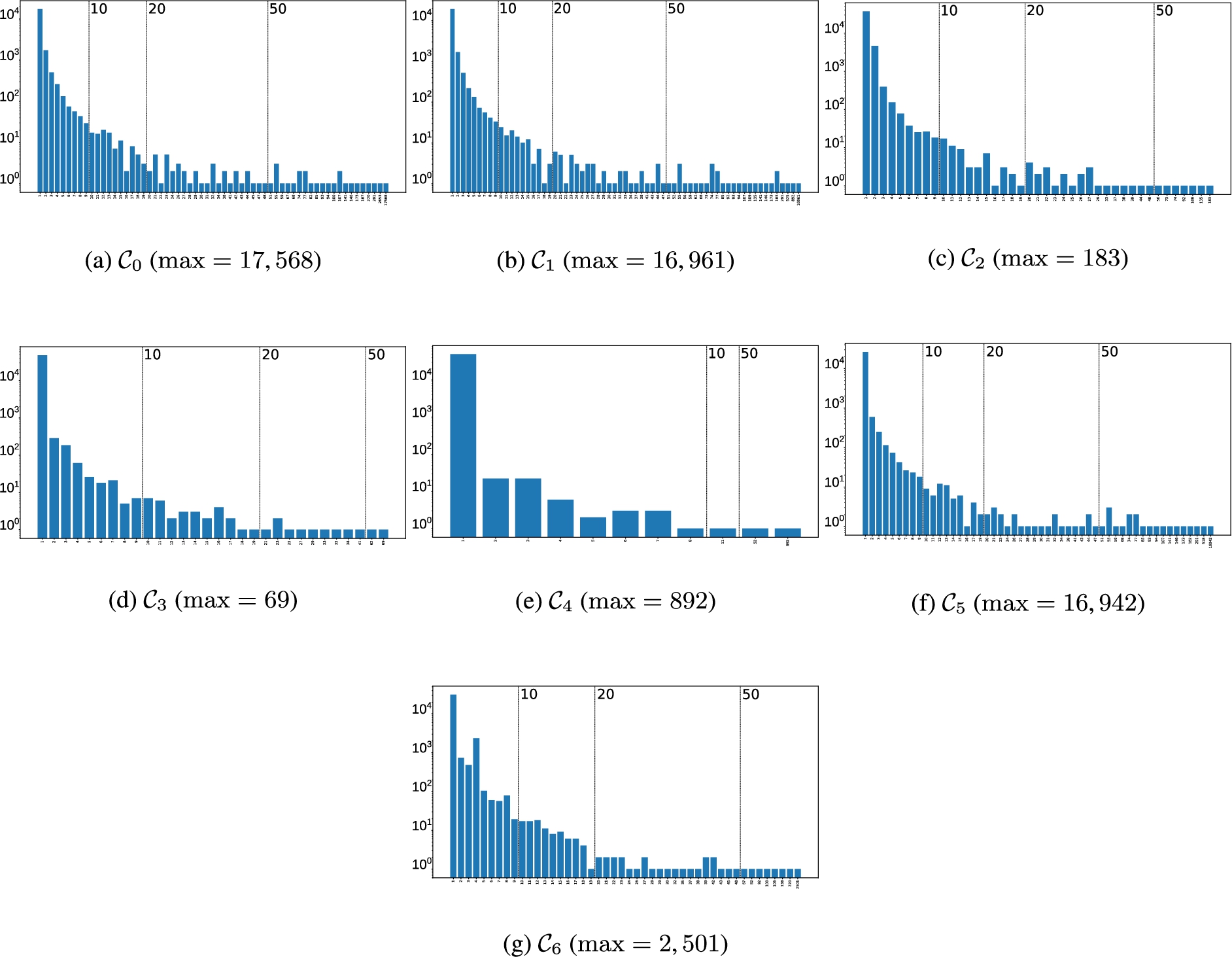
We experimented our approach with different pairs
An architecture formed by 3 GCN layers is used to learn node embeddings. The input layer consists of a featureless approach as in [20,29], The 3-hop neighborhood of a node
Statistics of PGxLOD and its transformations as described in Section 4. Statistics for PGxLOD discard literals and edges incident to literals. As we use a 3-layer architecture, statistics for all
Only the embeddings of nodes in
Clustering algorithms are only applied on the embeddings of nodes in
Results on all gold clusterings and graphs
Results on
During learning and clustering, our model is unaware of the different alignment relations holding between similar nodes. Indeed, the SNN loss only considers labels of gold clusters that do not indicate the alignment relations used to compute these clusters. This is particularly relevant for gold clusterings

Distributions of distances between similar nodes by alignment relation for each test set, the
Impact of inference rules
It appears that
In
Impact of clustering set-up
Clustering performances are generally better for gold clusterings
Among the considered clustering algorithms, Single generally performs better than the others. For
Distance analysis
Regarding the distance analysis of node embeddings, Fig. 4 shows that distances between similar nodes are different depending on the alignment relation holding between them. Recall that our GCN model is agnostic to these alignment relations when computing the SNN loss. Interestingly, distances reflect the “strength” of the alignment relations: strong similarities (
Towards a further integration of domain knowledge in GCNs
Our results highlight the interest of considering domain knowledge associated with knowledge graphs in embedding approaches and seem to advocate for a further integration of domain knowledge within embedding models. Future works may investigate the same targets with additional inferences rules (
Generalization to other knowledge graphs
Despite our approach being motivated by the matching of individuals within an aggregated knowledge graph, its transposition to distinct graphs could be explored. Such a perspective could allow to assess the generalization of our approach and its results. In particular, we could consider knowledge graphs that are not completely independent such as LOD datasets that are connected through major LOD hubs. Recall that our approach is supervised since gold clusters are computed from preexisting alignments. Hence, testing our approach on different knowledge graphs of the LOD would require such preexisting alignments or using ontology alignment systems in a distant supervision process [8]. In this setting, merging the different graphs into one and learning a “global” embedding, as we did, may provide positive results but may pose additional scalability issues.
Conclusion
In this paper, we proposed to match entities of a knowledge graph by learning node embeddings with Graph Convolutional Networks (GCNs) and clustering nodes based on their embeddings. We particularly investigated the interplay between formal semantics associated with knowledge graphs and GCN models. Our results showed that considering inference rules associated with domain knowledge tends to improve performance. Additionally, even if our GCN model was agnostic to the exact alignment relations holding between entities (
Footnotes
Acknowledgements
This work was supported by the
Detailed results of clustering experiments
Detailed results of clustering experiments are available Table 8, Table 9 and Table 10.
Results of clustering nodes that belong to gold clusters whose size is greater or equal to 50 for graphs Results of clustering nodes that belong to gold clusters whose size is greater or equal to 20 for graphs Results of clustering nodes that belong to gold clusters whose size is greater or equal to 10 for graphs Results of clustering nodes that belong to gold clusters whose size is greater or equal to 50 for Results of clustering nodes that belong to gold clusters whose size is greater or equal to 20 for Results of clustering nodes that belong to gold clusters whose size is greater or equal to 10 for
ACC
ARI
NMI
ACC
ARI
NMI
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
ACC
ARI
NMI
ACC
ARI
NMI
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
ACC
ARI
NMI
ACC
ARI
NMI
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
Ward
Single
OPTICS
ACC
ARI
NMI
ACC
↓
ARI
↓
NMI
↓
↓
↓
ACC
ARI
NMI
↓
ACC
↓
ARI
NMI
↓
ACC
↓
↓
ARI
↓
↓
NMI
↓
↓
↓
ACC
ARI
NMI
↓
Ward
Single
OPTICS
ACC
ARI
NMI
ACC
↓
ARI
↓
NMI
↓
↓
ACC
ARI
↓
NMI
↓
ACC
↓
ARI
↓
NMI
ACC
↓
↓
ARI
↓
↓
↓
NMI
↓
ACC
ARI
NMI
↓
Ward
Single
OPTICS
ACC
ARI
NMI
ACC
ARI
NMI
↓
ACC
↓
↓
ARI
↓
NMI
↓
↓
↓
ACC
↓
ARI
NMI
↓
ACC
↓
ARI
↓
NMI
↓
ACC
↓
ARI
NMI
↓
↓
