Abstract
Reusing ontologies and their terms is a principle and best practice that most ontology development methodologies strongly encourage. Reuse comes with the promise to support the semantic interoperability and to reduce engineering costs. In this paper, we present a descriptive study of the current extent of term reuse and overlap among biomedical ontologies. We use the corpus of biomedical ontologies stored in the BioPortal repository, and analyze different types of reuse and overlap constructs. While we find an approximate term overlap between 25–31%, the term reuse is only <9%, with most ontologies reusing fewer than 5% of their terms from a small set of popular ontologies. Clustering analysis shows that the terms reused by a common set of ontologies have >90% semantic similarity, hinting that ontology developers tend to reuse terms that are sibling or parent–child nodes. We validate this finding by analyzing the logs generated from a Protégé plugin that enables developers to reuse terms from BioPortal. We find most reuse constructs were 2-level subtrees on the higher levels of the class hierarchy. We developed a Web application that visualizes reuse dependencies and overlap among ontologies, and that proposes similar terms from BioPortal for a term of interest. We also identified a set of error patterns that indicate that ontology developers did intend to reuse terms from other ontologies, but that they were using different and sometimes incorrect representations. Our results stipulate the need for semi-automated tools that augment term reuse in the ontology engineering process through personalized recommendations.
Keywords
Reuse in biomedical ontologies
The biomedical research community has been one of the earliest adopters of ontologies to tackle the challenges of efficient knowledge organization, optimized information retrieval and effective annotation of datasets. Researchers have used ontologies for various purposes such as knowledge management, semantic search, data annotation, data integration, exchange, decision support and reasoning [5,32]. For example,
Over the years, ontology development has become a reuse-centric process [34,38]. All methodologies strongly encourage reuse while building new ontologies, be it at the level of an ontology, or at the level of individual terms [2,7]. In the literature, we may find two areas that benefit from reuse:
Several large, collaborative efforts are trying to streamline the development of interoperable, logically well-formed and accurate biomedical ontologies. They deal with ontological term overlap and reuse in different ways. For example, one of the key aims of the Open Biological and Biomedical Ontologies (OBO) Foundry [36] is to create a set of
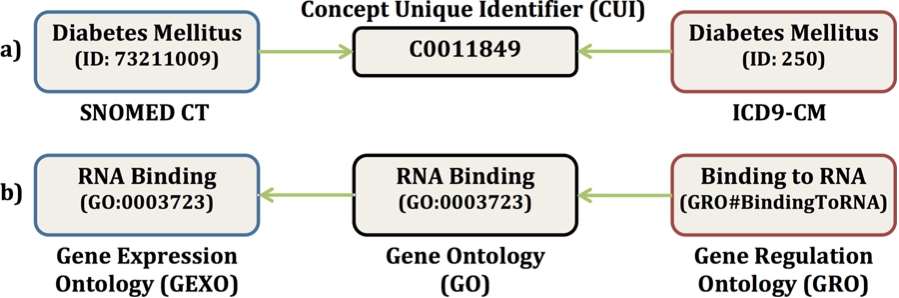
Types of Reuse: (a)
For the purpose of this work, we define a
For this research, we use the entire set of biomedical ontologies stored in BioPortal [45], an open content repository of biomedical ontologies and terminologies. The key contributions of this research can be described as follows:
We provide a systematic study of the current state of reuse and overlap across biomedical ontologies.
We propose and implement a new approach to determine term overlap across ontologies using composite mappings.
We develop a clustering method to help identify patterns of reuse using semantic similarity among ontology terms, and validate the results using the BioPortal Import Plugin logs.
We implement a Web application that can search for similar and reused terms in Bioportal ontologies, and that can visualize reuse dependencies and overlap among ontologies.
We discuss the state and challenges of reuse in biomedical ontologies.
All results of this paper, as well as all developed visualization tools, are available online at:
The paper is structured as follows: Section 2 describes the related work to this research. Section 3 presents the methods that we used for our descriptive study. Section 4 details the results of applying the research methods, and then we discuss our findings in Section 5.
Benefits and challenges of reuse
Ontology reuse is recommended in the methodologies and guidelines outlined by several engineering groups as a means to develop modular, interoperable, accurate and cost-effective ontologies [10,26,38]. Bontas et al. [6] provide several real-world use cases for the benefits of ontology reuse in biomedicine and eRecruitment. By empirically analyzing methodologies, methods and tools currently used, Simperl et al. [34] identify the research and development challenges for ontological knowledge reuse to become a feasible alternative to other ontology-development strategies. In essence, reuse can be increased through the development of pragmatic methods and semi-automated tools that optimally exploit human and computational intelligence for reusing ontologies through a context- and task-sensitive approach [34]. Ontology modularization techniques (i.e., extracting parts of an ontology using some structural or logical properties) are also an important factor in supporting reuse. Researchers have undertaken comprehensive studies of existing modularization techniques [11,29].
Tools to support reuse
There are only a few tools that support term reuse in biomedical ontologies. OntoFox [46] is a Web-based application that allows users to retrieve terms, selected properties, and annotations from the source ontologies, using MIREOT principles [9]. The BioPortal Import Plugin [23,24] is an extension of the Protégé ontology editor [27] that allows the importation of terms, their properties and class subtrees from BioPortal ontologies. The MIREOT Protégé Plugin [15] and DOG4DAG [44] are also Protégé plugins that provide term importations from external ontologies. ProtégéLov [12] allows reuse of terms from the Linked Open Vocabularies repository [42] using
Previous analyses of reuse and overlap
Matentzoglu et al. [22] provide a method to analyze the overlap between automatically-downloaded OWL ontologies from the Web. Ontologies with 90% overlap or containment relations were considered similar. Poveda et al. [30] analyzed the landscape of reuse in the ontologies referenced in Linked Open Data (LOD). The results indicate that over 40% of the terms are reused from other vocabularies, 67% of which are reused by imports, and the rest by referencing the term IRI.
In 2010, a systematic analysis of the member and candidate ontologies in the OBO Foundry indicated that the OBO Foundry had made significant progress over a period of two years towards the goal of orthogonality [14]. However,
Five years later, we conducted a study [20] to investigate the level of reuse across all the biomedical ontologies stored in BioPortal [45]. Both these studies carried out simple lexical comparisons of the term labels to determine term overlap. Even though effective, this naive method tends to leave out terms that represent the same concept but have lexically-different term labels (e.g.,
We found term reuse to be 3.1%, 3.9% and 4.1% for the three reuse types respectively, whereas, we found a term overlap of 14.4%. We also found that most ontologies reuse less than
In this paper, we will extend this research by providing a new approach to determine term overlap, a better metric to estimate term overlap and reuse, and a deeper understanding of how ontology developers reuse terms.
Methods

Worflow of all the steps required to estimate the average term reuse and overlap statistics across the BioPortal Ontologies, as well as clustering and BioPortal Import Plugin Log analysis to detect any reuse patterns. The steps of the workflow are: (1) Ontology Pre-processing, (2) Term Reuse, (3) Term Overlap, (4) Clustering, and (5) Log Analysis.
For our descriptive study, we employed several methods that aim to: (i) estimate the level of term reuse and term overlap across biomedical ontologies, (ii) extract reuse patterns from BioPortal ontologies, and (iii) extract reuse patterns from time-stamped BioPortal Import Plugin logs. These methods are inspired from text mining, graph theory and unsupervised learning. We make the results available through interactive visualizations and a search application (
We used two datasets for our study: (i) a dump of BioPortal ontologies to analyze term reuse (Step 2) and overlap (Step 3), as well as to perform the clustering (Step 4); and (ii) the logs of the BioPortal Import Plugin to analyze the patterns of reuse in user ontologies (Step 5).
BioPortal ontologies
We obtained a triplestore dump of the BioPortal ontologies in N-triples format that contained 509 distinct ontologies as of January 1, 2015. This dump did not contain some ontologies that were deprecated or merged with existing ontologies, or added to BioPortal after January 1, 2015. After removing ontological views (i.e.
BioPortal Import Plugin logs

An anonymized excerpt of the BioPortal Import Plugin Logs
The BioPortal Import Plugin, an extension to the Protege ontology editor, allows users to import terms and sub-trees from BioPortal ontologies into their own ontology [23,24]. The plugin invokes the BioPortal REST API to search the BioPortal ontologies, and also to import terms.
We obtained the logs of REST calls that the plugin made to BioPortal. The logs are time and IP-stamped, and span the period from 26th September, 2011–14th May, 2013 (∼20 months). Listing 1 shows an excerpt of these logs.
Even though we did not have access to the user ontologies into which these imports were performed, these logs were an important source of information of terms that were reused together in user ontologies. We used these logs to identify patterns of reuse (Fig. 2, Step 5).

Cartoon representations of the (a)
For the purpose of this work, we define as
To identify term reuse (Fig. 2, Step 2), we used the BioPortal corpus (Section 3.1.1), and defined three reuse constructs:
We iterated over all the axioms in each of the 377 BioPortal ontologies to extract class term IRIs, their labels, synonyms, UMLS CUI reuse was excluded, as we could not identify the source ontology for a CUI.
The percentage of terms reused using the first two constructs from other ontologies (IRI and
The total number of ontologies reused from,
The percentage of terms reused by other ontologies,
The total number of other ontologies reusing terms,
CUI-mapped terms among other ontologies,
Reuse among all distinct pairs of ontologies.
Using these metrics, we determined those ontologies that reused the maximum number terms from other ontologies, and also those ontologies whose terms were reused the most.
An example of a composite mapping. A column represents the term shown in the header. The content of a column contains different labels (preferred labels, synonyms, etc.) associated to the term. An arrow indicates that a label of a term is mapped to the label of another term. This example shows how we can map Term A defined in
We generated a graph
– the graph module containing only IRI edges (an undirected edge links two terms with same IRI),
– the graph module containing only
– the graph module containing only CUI edges (an edge links two terms that are mapped to the same CUI).
Sources for labels and synonyms to generate composite mappings
For each reuse module, we calculated the term reuse across all biomedical ontologies using the equation given below, where
The above equation serves as a better metric to estimate term reuse as compared to the previous metric (Eq. (1)). The equation calculates the percentage of terms in BioPortal that are not unique, but are reused, unlike the previous metric, which did not include the count of reused versions of a term.
For the purpose of this work, we define
In our initial approach [20], we normalized the term labels by converting them to lowercase and then removing all non-alphanumeric characters. We performed naïve string matching to determine the potential
To overcome this limitation, we considered using composite mappings in the current approach. Given a mapping from
We extended this notion to generate graphs of such composite mappings (
We normalized the labels and synonyms, by first removing a set of 126 common English stop words (e.g.
We generated 5 different overlap modules from different combinations of composite mappings:
The
The final
In the next step, we identified those terms that had the same source ontology and identifier, but a different IRI representation, and no explicit mappings (e.g.,
We calculated term overlap for each overlap module using the metric described in Eq. (2), where all nodes (terms
For each the five overlap modules, we conducted an empirical analysis on the composition of the term labels of 100 randomly selected components to determine the threshold of the maximum distance (mapping hops) between two leaf nodes, for which any component
We called the components that have mappings exceeding the maximum distance
An example of a hybrid component
, composed of terms
.
can be broken into two smaller, relevant components
and
that are connected by an incorrect mapping caused due to a synonym of term
An example of a hybrid component
We calculated another term overlap estimate, which we called
One goal of this work is to investigate whether the reuse within biomedical ontologies occur in certain patterns that can be identified algorithmically. To this end, in Step 4 of our workflow (Fig. 2), we used a two-phase clustering approach on the IRI module that we defined in Section 3.1.1. As a reminder, the IRI reuse module contains only IRI edges that link terms that share the same IRI.
We excluded the
Using the terms in the
As our term–ontology matrix
For each pair of terms in each cluster, we computed a similarity score as follows:
In the equation above,
As can be seen, the similarity measure is a weighted distribution of common ontologies and Jaccard semantic similarity.
We generated a term–term affinity matrix
Analyzing BioPortal Import Plugin logs
In step 5 of our workflow (Fig. 2), we analyzed the logs generated by the BioPortal Import Plugin (see Section 3.1.2). We used this analysis for two purposes: (1) to gain knowledge on other reuse patterns that occur in user ontologies, and (2) to validate whether the insights generated from our clustering analysis are accurate.
The entries in the BioPortal logs are generated as the user does certain operations in the user interface of the plugin. For example, if the user searches for a term in a BioPortal ontology using the plugin, the log will record a line corresponding to the
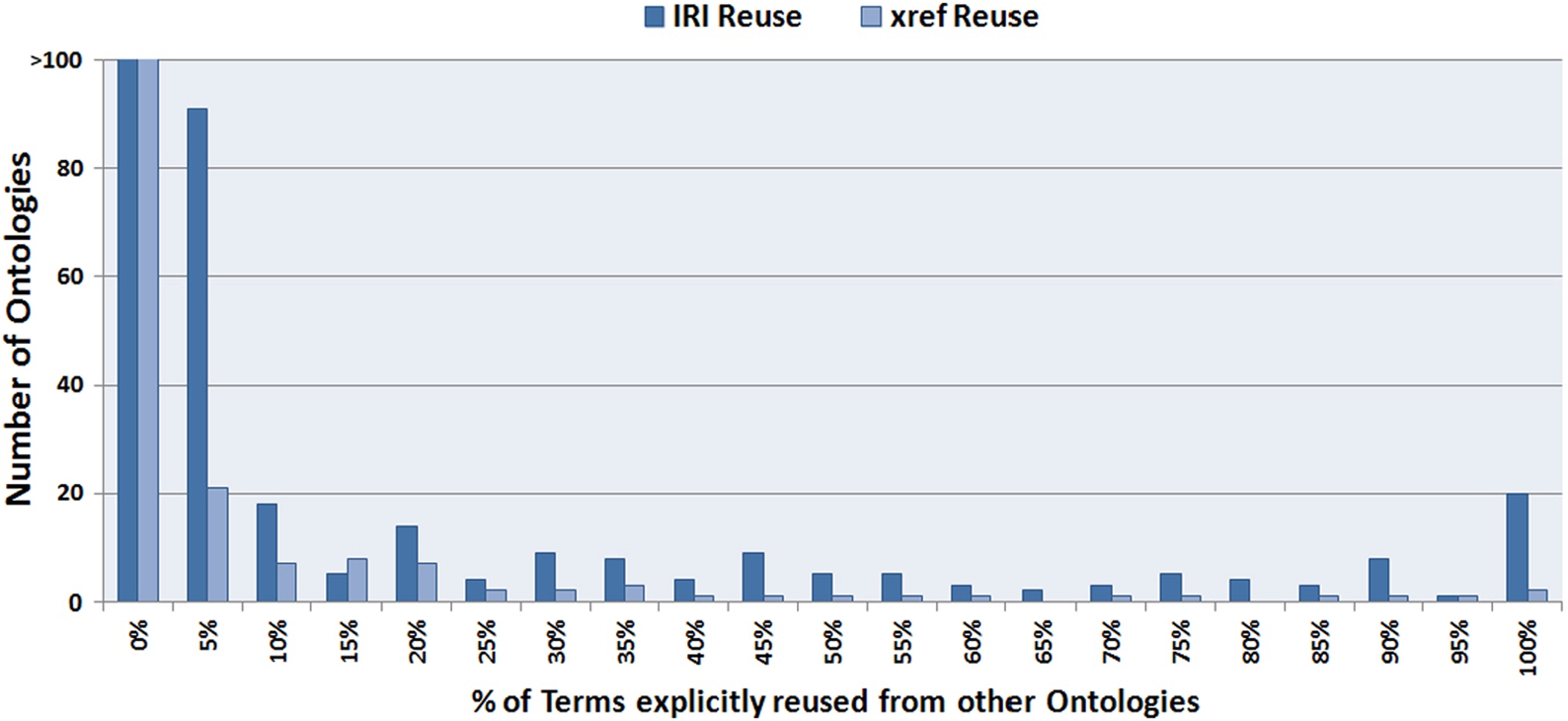
Histogram depicting the number of ontologies that reuse a given percentage (%) of terms
As we do not have access to the user ontologies into which the BioPortal terms have been imported, the only sources we have are the time- and IP-stamped BioPortal call logs. Therefore, we had to reverse-engineer these logs to find out the actions that the users have taken in the user interface, and to identify which BioPortal terms are being reused (i.e., imported) together.
We documented the algorithm we used to reverse-engineer the logs in the additional online materials (
As a result of running the reverse-engineering algorithm on BioPortal logs, we obtained term sets that have been reused (i.e., imported) together in user ontologies. Then, we mapped the extracted terms to existing terms in the current version of the source BioPortal ontology to find the overall depth of tree imports and the location of these terms and subtrees. We used this information as an additional source of reuse patterns, and also to validate the hypotheses made from clustering analysis (Section 3.4).
We now present the results of each of the methods that compose our workflow (Fig. 2), described previously in Section 3.
Reuse
Previously, we found that most ontologies reuse less than 5% of the total terms in their current versions, using either the same IRI or through
Using our
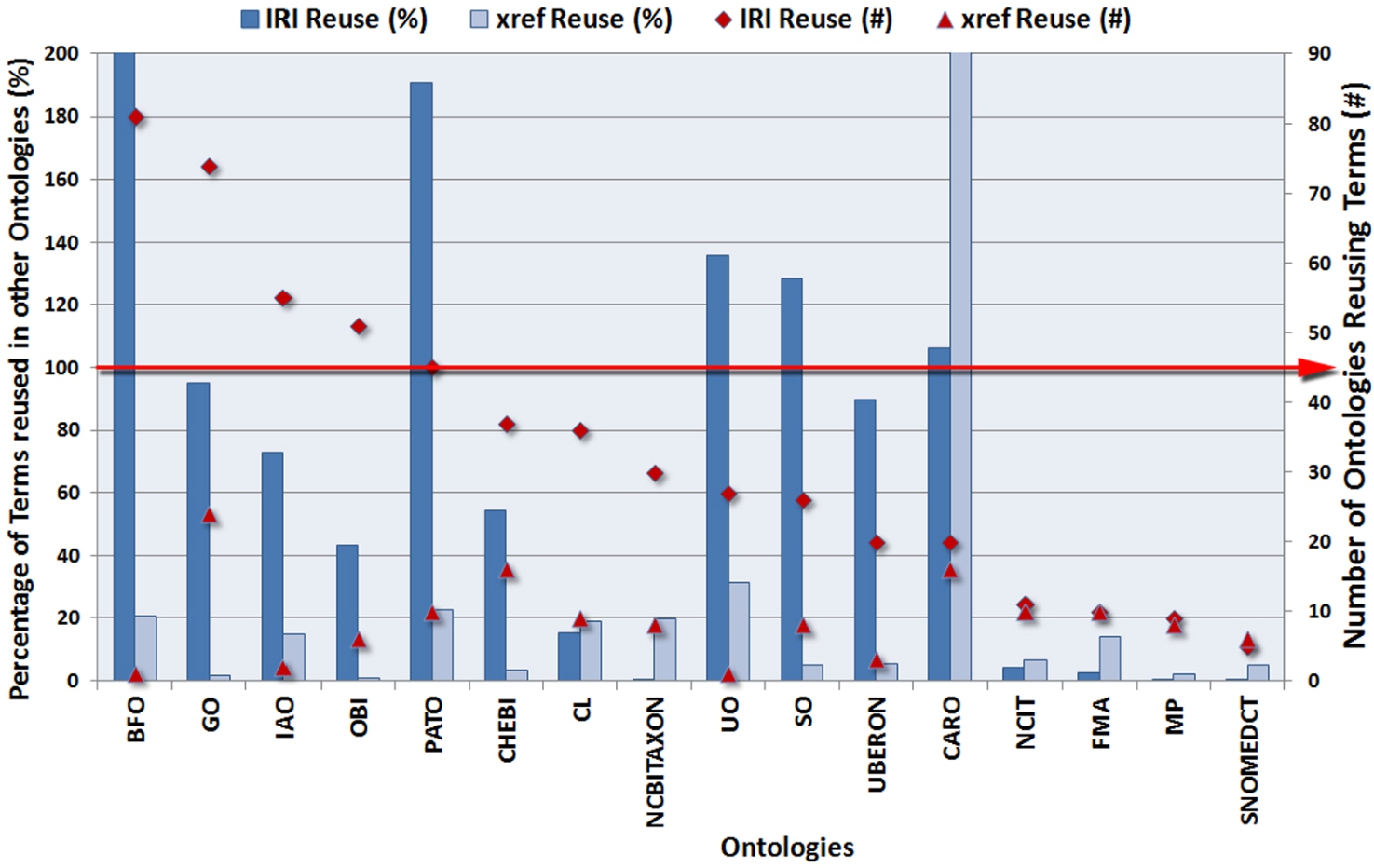
Top 16 ontologies whose terms are reused the most through
Term overlap (actual and hybrid-adjusted) estimated for different overlap modules composed of different mappings
The 16 ontologies whose terms are reused the most from the first 2 constructs (IRI and
It is easily noticeable that most of these are popular or upper-level ontologies, some of which have more than 100% of their terms reused (e.g., we found 101 different versions of Basic Formal Ontology – BFO IRIs, whereas the current version only has 39 terms). As we have discussed [20], this anomaly is due to the fact that ontology developers tend to reuse terms with different versions, notations, or namespaces, that are sometimes incorrect and have no explicit mappings to the original term. We do not consider this case as reuse, but rather an
Using the updated metric described in Section 3.2, we found term reuse to be 6.63% for the
Term overlap
In our previous work [20], we determined term overlap using a naive approach. We found a total of 2,023,854 terms sharing 752,177 unique labels across the BioPortal ontologies. Using the new metrics described in Section 3.3, we can calculate this naive term overlap to be
The

30% term overlap among different BioPortal ontologies. For simplicity, only the OBO Foundry member and candidate ontologies (blue squares), UMLS terminologies (red circles), and a few popular ontologies in BioPortal (green octagons) are shown here.
Once we include also the other types of mappings using synonyms (rows 2–6 in Table 4), the term overlap gradually increases all the way up to
Rows 6 and 7 in Table 4 show that after removing all the three reuse modules (cf. Section 3.3), the term overlap decreases—the range is (
As a next step, we investigate how the term overlap reflects on ontology overlap. Therefore, we mapped the nodes in the
We generated a directed sub-graph (Fig. 6) between those ontologies that have more than
Figure 6 shows that there is substantial overlap among ontologies generated independently through the OBO Foundry and UMLS methodologies. The overlap between BFO and the OBO Foundry candidate ontologies is caused by the fact that the candidate ontologies import BFO as their upper-level ontology, but they use different (incorrect) IRI representations. It is also noteworthy to see that the UMLS terminologies for adverse events, namely World Health Organization Adverse Reaction Terminology (WHO-ART), Coding Symbols for a Thesaurus of Adverse Reaction Terms (COSTART), and the Medical Dictionary for Regulatory Activities (MEDDRA), have substantial term overlap. The lower region of the graph shows several anatomical ontologies (CARO, UBERON, XAO, TAO, FMA, MA, TGMA, etc.), in which term overlap is obvious (similar anatomical features), but is debatable – most terms represent anatomical parts that may not be necessarily equivalent, as they belong in different organisms. Finally, the top-right corner shows the overlap between the RxNorm Vocabulary and the Drug Ontology (DRON). These results and the
Clustering
The first step of our two-phase clustering approach was to use a
The primary ontological composition of the clusters was determined from the ontologies common among terms in a cluster, and is shown in Table 5. It should be noted that
Primary ontological composition of the clusters
Primary ontological composition of the clusters

Proportion of term pairs with semantic similarity in a given range for each sub-cluster.

We computed an affinity matrix among all pairs of terms in a given cluster using weights
After executing spectral clustering using the affinity matrix, we divided all the term pairs in each sub-cluster in 2 bins, based on their Jaccard semantic similarity measure (<0.9 in Bin 1, and >0.9 in Bin 2). We plotted the proportion of term pairs in each bin for each cluster. Cluster 4 is shown in Fig. 7. In Cluster 4, a larger proportion of term pairs in any given sub-cluster have a semantic similarity in the range of (0.9–1.0) (>70%), indicating that these are either sibling terms or one term is the direct superclass of another. Generally, we found this to be the case for all the large clusters of the first kind. This finding likely indicates that ontology developers reusing terms from one main source ontology tend to reuse hierarchical subtrees mainly composed of terms with parent–child or sibling relations. This was less evident in the second kind of the large clusters where the proportion ranged between 30–60% of term pairs.
We mapped these sub-clusters to their location in the source ontology. We found that most of these 2-level substructures are located in the higher or upper-middle levels of the ontology. Hence, developers reuse terms from the higher levels in the ontological hierarchy of a small set of popular ontologies, and seldom reuse leaf nodes.
We found a total of 3,538 distinct IP addresses originating from 90 different countries, from which ontology developers used the BioPortal Import Plugin to search and reuse terms from BioPortal ontologies. We were able to isolate 5,755 individual terms and 2,139 ontological subtrees imported from 40 different ontologies in 516 distinct sessions. For an IP address, a
The top 10 ontologies with the maximum number of sessions were SNOMEDCT, NCIT, BFO, ABA-AMB, FMA, GO, RCD, AMINO-ACID, HP and IAO, whereas with the maximum number of terms were in ICD10PCS, SNOMEDCT, NCIT, ICD9CM, LOINC, BIRNLEX, ABA-AMB, FMA, RCD and SHR.
The ontologies that were reused the most through the plugin, both by the maximum number of sessions or by the maximum number of terms, are shown in Fig. 8. The total number of sessions observed, total number of single term imports, total number of structures imported, and total number of terms imported are shown as a bar plot. The structure of the content imported from each source ontology is shown across the depth of an ontology – the imported structures are shown as translucent blue polygon and the terms imported (either single or as a group) are shown as circular constructs, grouped according to the level. The depth of the ontology was retrieved from BioPortal repository. The width of the structure on each level is indicative of the number of terms imported on that level in log scale. The radius of the circular construct represents the total number of terms on that level. For clarity purposes, we have only shown 4 ontologies – FMA, ICD10PCS, NCIT and SNOMEDCT. The website (
In general, we found that, on an average more people tend to reuse terms from OBO Foundry ontologies (higher number of sessions detected) than UMLS terminologies using the Bioportal Import Plugin, with the exception of NCIT and SNOMED CT. However, the users, who import UMLS terminologies, tend to reuse more number of terms, in the form of complete hierarchical structures, during a single import session.
In the cases of ICD10PCS and ICD9CM, we found that the users reuse the entire hierarchy of these ontologies starting from the root node, into their target ontology. We observed the same pattern also in the case of the BFO, but it is expected as it is an upper level ontology. In almost all the other cases, we found that the ontology developers simply reuse terms from the higher or upper-middle levels in an ontological hierarchy, and the lower leaf nodes and structures are seldom reused. This reuse pattern can be seen in the FMA ontology in Fig. 8. We found the same reuse pattern in GO, CHEBI, NCBITAXON and LOINC (
Reuse and overlap visualization on the Web
One of the contributions of our work is a general-purpose visualization of reuse and overlap among biomedical ontologies that employs the reuse and overlap modules, which we generated as part of this work. The Web application also allows users to search for similar terms by providing any string or an IRI as an input. In case of a string, the application matches the name to the set of the most similar terms that have it as a label or a synonym. We believe such an application is of general interest, and we make it available to the community through our website (
The application does a depth-first search against the
Discussion
Term reuse
As seen in Fig. 4, we are seeing the full spectrum of reuse from 0–100%, but in general, reuse is fairly low. Not only do most ontologies in BioPortal never reuse terms, their terms are also never reused by other ontologies, which is contrary to the reference-application paradigm considered in the ontology engineering process. However, we did find some ontologies that are approaching complete reuse. For example, the Mental Functioning Ontology (MF) [17], reuses 91.33% of its terms from 6 different ontologies. Our clustering analysis shows that not only single terms are reused, but also entire hierarchical structures of the source ontologies are reused. Ontology engineers need semi-automated tools to support both cases.
Generally, well-established ontologies and controlled terminologies do not reuse terms from other ontologies. Usually, these ontology are built by large organizations (e.g., NCI, WHO, IHTSDO). Some of these organizations are making concerted efforts to take advantage of reuse. For example, ICD-11 and SNOMED CT are trying to define a common core ontology to be reused by both [31]. Such collaborations may generate a set of best practices for ontology reuse in the future.
Different kinds of IRI representations observed in BioPortal ontologies and BioPortal Import Plugin logs
(*) marks the recommended representation(s)
Different kinds of IRI representations observed in BioPortal ontologies and BioPortal Import Plugin logs
(*) marks the recommended representation(s)
Through the empirical analysis of the BioPortal Import Plugin logs, as well as, the generated clusters and overlap modules, we found some reuse patterns that show that ontology developers have the intention to reuse terms. Essentially, these are IRI patterns that generally have the same identifier and source ontology, but that are reused from different versions of the source ontology, or represented using different notations or namespaces. These patterns cannot be considered as term reuse, as the IRIs use different, and often incorrect, representations for the same terms, and no explicit CUI or
We found several cases, in which an ontology reuses the same terms from different ontologies, and these terms are not linked by a reuse construct. For example, the BioModels Ontology (BIOMODELS) reuses the same terms from two different ontologies:
Based on the observations from this study that show only modest reuse among biomedical ontologies, we believe that ontology engineers would benefit from better guidelines, along with improved tools, to increase term reuse.
In 2010, a systematic analysis of all the OBO Foundry ontologies outlined consistent term overlap, yet minimum term reuse, and commented on the limitations and challenges to achieve
The OBO Foundry mandates reuse by candidate ontologies from the member ontologies under its orthogonality aim. However, there is still substantial
In our previous analysis, we used a conservative approach to determine term overlap. As a result, lexically-different terms that may be similar, and can be categorized under term overlap, were considered different. Using our approach of tokenization and removal stop words, we were able to map terms with labels such as
Our approach for detecting overlap has certain limitations.
Terms with labels such as
Lexically-similar terms in different ontologies may represent different concepts (e.g., anatomical concepts like
Some biomedical ontologies use different classes for the same concept to show evolutionary or developmental stages (e.g.
Some ontologies may instantiate a synonym relation between terms that can actually have an
Some ontologies use chemical formulas as synonyms. Terms with the same chemical formula may be stereoisomeric molecules or completely different compounds (e.g.,
Hence, the term overlap estimates should be seen cautiously, and can serve as an upper bound to the actual
Clustering
One of the key challenges that we encountered while clustering was the fact that we were dealing with a large number of terms (compared to the features), resulting in a large
From clustering, we claim the following hypotheses:
BioPortal Import Plugin log analysis
As was observed from our term reuse analysis across BioPortal ontologies, ontology developers only import terms from a small set of popular ontologies in BioPortal using the BioPortal Import Plugin. From our analysis of the logs, it is apparent that:
Hence, we can say that the claims made from our clustering analysis (Section 5.3) are validated through our BioPortal Import Plugin log analysis. As future work, we plan to do a more formal validation of this finding. Moreover, for some ontologies that were common between both our analysis (e.g. NCIT, GO and FMA), we found a substantial similarity between some sub-clusters and the reuse structures extracted from the logs (results online). The similarity ranged between 70–100% for NCIT structures. This similarity can suggest either the ontologies developed using the BioPortal Import Plugin were saved back to BioPortal repository, or there are recurrence patterns in some ontologies that are reused frequently in different ontologies.
From this validation, we can postulate that our approach used for the two-phase clustering process, using the similarity equation and the term–term affinity matrices, accurately captures the thought process of the ontology engineer, when she reuses terms, and it can be coupled with the BioPortal Import Plugin to provide reuse recommendations in the future. The clustering only used the terms in the IRI reuse module, and might be biased towards OBO Foundry ontologies, and not generate enough UMLS recommendations (as they are seldom reused using the same IRI). Hence, our initial term–ontology matrix and the similarity equation will need to be extended to deal with this bias.
Future work
All ontology development methodologies encourage reuse with several advantages, such as cost reduction, quality control, semantic interoperability, EHR mining and query federation, cited in favor of reuse [6,19,31,41]. However, our extensive analysis suggests that ontology developers do intend to reuse terms, but often, they are not able to do so correctly. Converting the intent for reuse into actual reuse can help increase term reuse, and reduce term overlap (Section 4.2).
We plan to provide personalized reuse recommendations for ontology developers through a WebProtégé plugin (
We believe that our Web application will allow ontology developers to search for similar terms in other ontologies, while our visualization of overlap and reuse dependencies may guide developers to reuse terms in their own ontology based on the structure of ontologies in related domains. Our composite mappings approach may serve as a complement to the existing BioPortal mappings, which are currently generated through naive string matching algorithms [13]. We also plan to develop a term–centric visualization that summarizes everything known about a particular term in BioPortal, and presents it to developers and domain experts through an interactive interface. Our hope is that this visualization will enable ontology developers to serendipitously discover and reuse existing knowledge.
Conclusion
We estimated the level of reuse and overlap in a corpus of 337 ontologies from the BioPortal repository. We developed novel methods for detecting reuse and overlap in biomedical ontologies. Our findings show a term overlap of approximately 25.31–30.18%, and term reuse of less than 9%. Most ontologies reuse less than 5% of their terms from a small set of popular ontologies, with terms from several ontologies never being reused. We found strong indications that users actually intended to reuse terms, but in many cases they used incorrect representations. We also identified common error patterns in term reuse. Our hope is that the results of this work may be used to develop better guidelines and tool support with the aim to enhance reuse, and minimize overlap among biomedical ontologies.
Acknowledgments
The authors acknowledge Manuel Salvadores for providing a triplestore dump of BioPortal ontologies, and other members of the Protégé Group and the National Center for Biomedical Ontology for their input. This work is supported in part by grants GM086587 and GM103316 from the US National Institutes of Health.
