Abstract
Recent and intensive research in the biomedical area enabled to accumulate and disseminate biomedical knowledge through various knowledge bases increasingly available on the Web. The exploitation of this knowledge requires to create links between these bases and to use them jointly. Linked Data, the SPARQL language and interfaces in natural language question answering provide interesting solutions for querying such knowledge bases. However, while using biomedical Linked Data is crucial, life-science researchers may have difficulties using the SPARQL language. Interfaces based on natural language question answering are recognized to be suitable for querying knowledge bases. In this paper, we propose a method for translating natural language questions into SPARQL queries. We use Natural Language Processing tools, semantic resources and RDF triple descriptions. We designed a four-step method which allows to linguistically and semantically annotate questions, to perform an abstraction of these questions, then to build a representation of the SPARQL queries, and finally to generate the queries. The method is designed on 50 questions over three biomedical knowledge bases used in the task 2 of the QALD-4 challenge framework and evaluated on 27 new questions. It achieves good performance with 0.78 F-measure on the test set. The method for translating questions into SPARQL queries is implemented as a Perl module and is available at
Introduction
Recent and intensive research in the biomedical area enabled to accumulate and disseminate biomedical knowledge through various knowledge bases (KBs) increasingly available on the Web. Such life-science KBs usually focus on a specific type of biomedical information: clinical studies in ClinicalTrials.gov,1
Nowadays, creating connections between these KBs is crucial for obtaining a more global and comprehensive view on the links between different biomedical components. Such links are also required for inducing and producing new knowledge from the already available data. There is a great endeavour in the definition of Open Linked Data to connect such knowledge and in taking advantage of the SPARQL language to query multiple KBs jointly. Particularly, the creation of fine-grained links between the existing KBs related to drugs is a great challenge that is being addressed, for instance, by the project Linked Open Drug Data (LODD).2
We start with the presentation of some related work.
Querying Linked Data requires to define the end user interfaces which hide the underlying structure of the KB as well as the SPARQL syntax. Usually, three ways are identified for querying Linked Data: knowledge-based specific interface, Graphical Query Builder and question answering system. However, it has been demonstrated that natural language interfaces are the most suitable [12]: indeed, for querying KBs and Semantic Web data, the use of full and standard sentences is preferred to the use of keywords, menus or graphs.
Another important distinction is related to the types of Linked Data which are processed, i.e. typically, general [5,15,32] or specialized [1] KBs, and the purpose of this processing. Concerning the purpose, two kinds of work can be distinguished:
Transformation of questions into queries
The main objective of this kind of works is to propose methods for a more efficient transformation of natural language questions into SPARQL queries. The evaluation of these approaches focuses on the syntactic correctness of the generated queries, but does not take into account the URIs which should be found through these queries. Most of the existing approaches rely on patterns or templates.
The question answering system AutoSPARQL is based on active supervised machine learning independent of the KB [18]. The SPARQL query model is learnt from natural language questions. The authors report that the 50 questions of the QALD-1 challenge are successfully transformed with the system. Another method is based on modular patterns for parsing the questions [26]. Semantic relations are identified on the basis of the first keywords detected. The method is tested on 160 movie-related questions. One advantage is that the method requires only four general and modular query patterns, while in a previous work of the authors, twelve patterns were necessary [25].
The use of resources automatically derived from ontologies or KBs also provides the possibility to transform questions into formal queries, such as those defined using the SeRQL language [31]: questions undergo a set of treatments (e.g. linguistic analysis, string similarity computation). Applied to a set of 22 questions, the method can interpret and transform correctly 15 questions (68%).
Existing tools can also be utilized. For instance, a multilingual toolkit, called
A manually written grammar together with ontological knowledge allow processing 145 questions out of 164 (88% coverage) [9].
Finally, [1] aims at translating natural language questions into SPARQL queries. The proposed method relies on a hybrid approach: a SVM machine learning-based approach, which is used to extract the characteristics of the questions (named entities, relations), is combined with patterns to generate the SPARQL queries. The method is applied to medical questions issued from a journal. The evaluation is carried out on 100 questions. The method achieves a precision of 0.62.
Queries generated by such systems can be further used for querying linked KBs, provided that the reference data, i.e. the expected resulting URIs, are available.
Query generation and querying Linked Data
The main objective of the following related work is more complex than work presented in Section 2.1: First, the system has to transform the natural language questions into SPARQL queries; and second, it has to query the KB in order to get the best results possible when querying the Linked Data. The main advantage of this kind of work is that they cover the entire querying process. Moreover, they allow to evaluate the final results (answers extracted from the KBs) and to provide precise evaluation figures. Often, NLP tools and methods are used for the transformation of questions into queries. We can mention three such experiments.
In one study, the system is template-based and relies on NLP tools and semantic resources [32]. The application of the system on 50 questions from DBpedia proposed by the QALD-2 challenge gives competitive results with an average of 0.62 F-measure obtained with 39 questions (the average recall is 0.63 and the average precision is 0.61), but shows a low coverage because 11 questions are not covered by the templates.
Notice that recently, the Question Answering over Linked Data (QALD-4) challenge proposed the task Biomedical question answering over interlinked data,3
Other research questions can be related to the querying of linked data with natural language interfaces. Usually this kind of work aims at improving specific points: identification of different types of SPARQL queries (select, construct, ask, describe) [19], detection of named entities [28], generation of SPARQL templates4
The objective of our work is to propose an end-to-end method for translating natural language questions into SPARQL queries and for querying KBs. The method is based on the use of Natural Language Processing (NLP) tools and resources for enriching questions with linguistic and semantic information. Questions are then translated into SPARQL with a rule-based approach.
Our method goes further in comparison with those presented in Section 2.2. Indeed, we propose to use information available in the Linked Data resources to semantically annotate the questions and to define frames (i.e., linguistic representations of the RDF schema) in order to model and build SPARQL queries. Thus, in comparison with the closest work [32], our method exploits extensively NLP for processing Linked Data. Besides, as our method performs an end-to-end processing, our work is also related to several aspects presented in Section 2.3: identification of different types of SPARQL queries [19], detection of named entities [28], generation of SPARQL templates [13], etc.
The paper is structured as follows. We describe the proposed method in Section 4 and then the semantic resources available and developed for enriching the questions in Section 5. The evaluation of the method is presented in Section 6 and we finally discuss our results in Section 8.
Question translation into SPARQL query

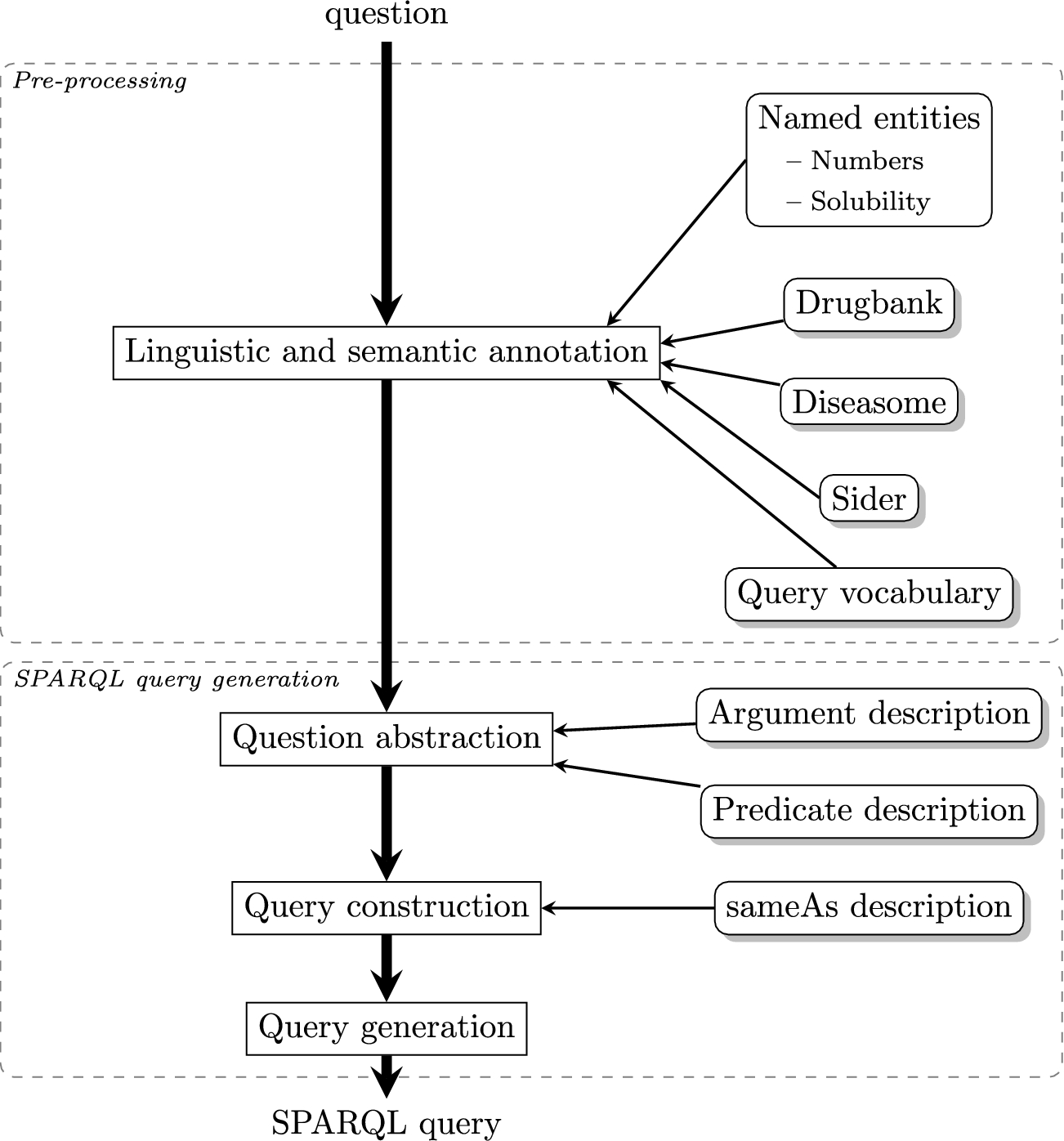
Linguistic and semantic annotation process. Square boxes represent the steps of linguistic and semantic analysis of questions. Rounded boxes indicate the resources used for the semantic annotation.
To translate natural language questions into SPARQL queries, we design a four-step rule-based approach, that relies on NLP methods, semantic resources and RDF triple descriptions (see Fig. 1):
Natural language questions are
Linguistic and semantic information is used for
The abstracted questions are used for
This graph pattern representation is used to
We design our approach on 50 questions proposed by task 2,
Note that the source questions are kept as provided despite the misspelling (first question for instance).

Examples of the linguistic annotation of questions issued from the QALD-4 challenge dataset. The source questions are kept as provided despite the misspellings they may contain (question (a) for instance). Gray rounded boxes represent words. Subscript text indicates Part-of-Speech tags computed by TreeTagger [29].
The annotation step aims at associating a linguistic and semantic description with words and terms from the questions (see Fig. 2).
First, the linguistic annotation aims at parsing questions in order to identify numerical values (such as numbers and solubility values) and words. During this step, part-of-speech tags and lemmas are associated with words. To achieve that, we use the TreeTagger POS-tagger [29]. Figure 3 illustrates the obtained linguistic annotation of questions. POS tagging errors are intentionally kept in the examples (e.g.

Examples of the question pre-processing. Gray rounded boxes represent words and semantic entities. Bracketed subscript texts are semantic types associated with semantic entities.
The objective of the semantic annotation is to identify semantic entities, i.e. terms together with the associated semantic types representing their meaning. Figure 4 displays the obtained semantic annotation of illustrated questions. This step relies on semantic resources, such as DrugBank, Sider and Diseasome (see Section 5), used in order to recognize semantic entities, such as disease names and side effects. The semantic entity recognition is based on the TermTagger Perl module.5

Question abstraction process. Square boxes represent the abstraction steps of questions. Rounded boxes indicate the resources used.
However, because semantic resources often suffer from low coverage [3,22], we also extract terms which usually correspond to noun phrases relevant for the targeted domain from the questions in order to improve the coverage of our approach. For instance, the terms  6
6 . They take into account the morpho-syntactic variation and represent basic syntactic dependencies within terminological entities. Each term is represented in a syntactic tree, while its sub-terms are also considered as terms in the current configuration (e.g.
. They take into account the morpho-syntactic variation and represent basic syntactic dependencies within terminological entities. Each term is represented in a syntactic tree, while its sub-terms are also considered as terms in the current configuration (e.g.
The processing of questions also requires to identify expressions of negation (e.g.
The question abstraction step aims at identifying relevant elements within questions and at building a representation of these elements (see Fig. 5). It relies on linguistic and semantic annotations associated with the question words detected at the previous step.
Before the identification of relevant elements, annotations are post-processed in order to disambiguate the generated semantic annotations. Indeed, annotated semantic entities may receive conflicting, concurrent or erroneous semantic types. For instance, in Fig. 4(g),
Besides, in order to choose the correct predicate it may be necessary to consider semantic types in the context of words or phrases corresponding to the predicates. For instance, the phrase Available at http://cpansearch.perl.org/src/THHAMON/RDF-NLP-SPARQLQuery-0.1/etc/nlquestion/SemanticTypeCorresp.rc.
In that respect, we manually analyze the predicate names in order to define rewriting rules and to adjust (modify or delete) semantic types associated with a given entity according to its context. Other rules may also modify or delete the entity itself. In total, we defined 44 contextual rewriting rules8
We define the question topic as the type of semantic entity which is the major context of the question, characterizing the user interest [8]. For instance, in Fig. 4(a), the question topic is
For performing question abstraction, we identify information related to the query structure:
Definition of the result form: Negated terms and information related to coordination markers, aggregation operators, and requirements on specific result forms are recorded and will be used at the end of the
Identification of the question topic: We assume that the first semantic entity occurring in the sentence, with a given expected semantic type corresponds to the question topic. The expected semantic types are those provided by the RDF subjects and objects issued from the resources. This information will be used during the
Examples of the question abstraction. The left part of the sub-figures displays the graph representation of the identified frames: gray boxes represent subjects and objects of the predicates together with their semantic types, while edges represent predicates and their semantic types. The right part of the sub-figures represents semantic entities and terms identified in questions together with the associated information: question topic (QT), URI, etc.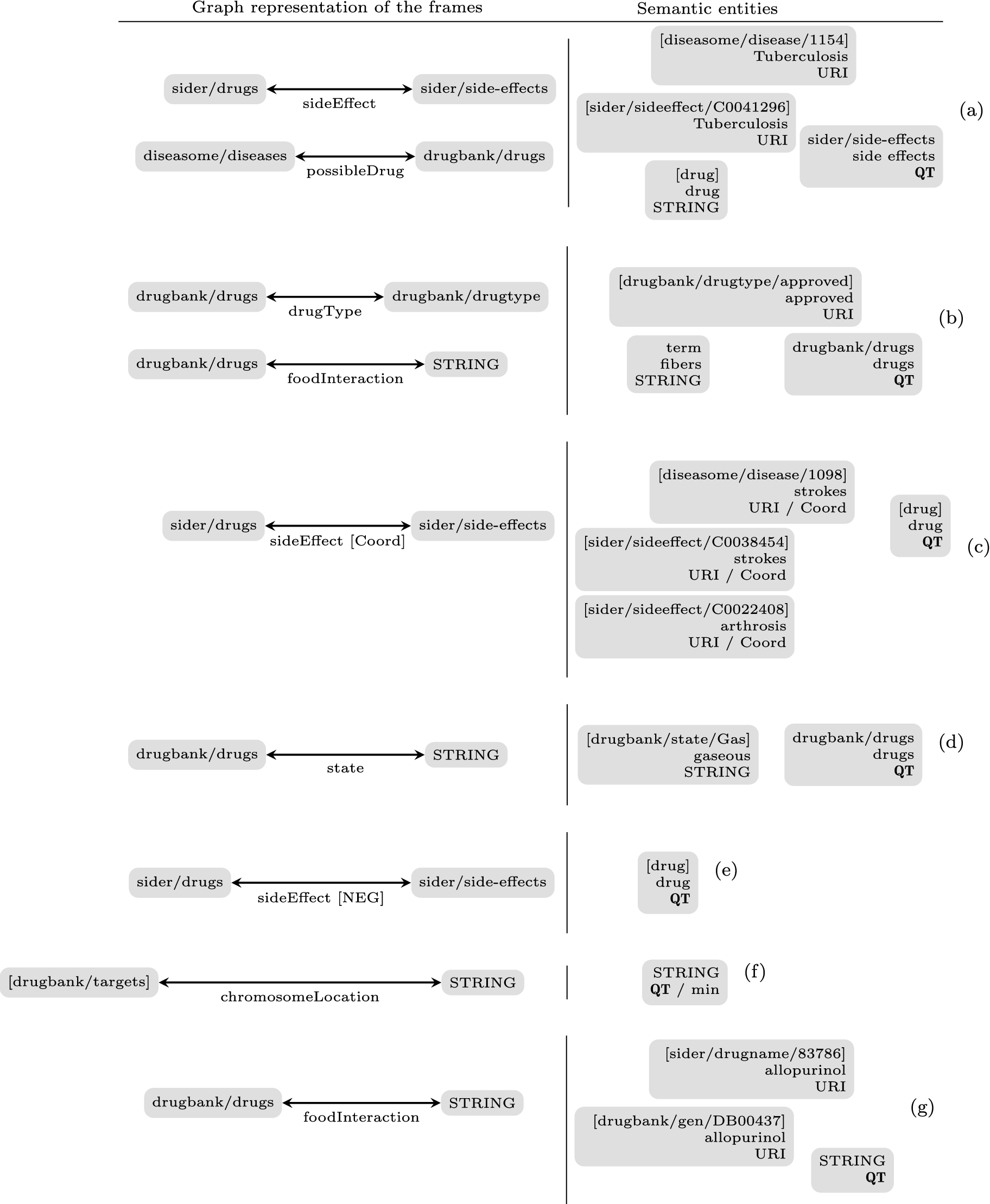
Identification of predicates and arguments: We use linguistic representations of RDF schemas, i.e. frames which contain one predicate and at least two elements with associated semantic types. In that respect, potential predicates, subjects and objects of frames are identified among the semantic entities and then recorded in a table: entries are semantic types of the elements and refer to linguistic, semantic and SPARQL information associated with these elements. Subjects and objects are fully described in the table with the inflected and lemmatized forms of words or terms, the corresponding SPARQL types and indicators on their use as object or subject of a given predicate. Concerning the predicates, only the semantic types of their arguments are instantiated. Subjects and objects can be URIs, RDF typed literals (numerical values or strings) and extracted terms (these are considered as elements of regular expressions).
For instance, in Fig. 6(a), two predicates are identified:
Scope of coordination: Arguments and predicates in the neighbourhood of coordination are identified. These elements are recorded as coordinated, e.g. the semantic entities
Figure 6 presents graph representations and abstractions of the questions presented in Fig. 4.
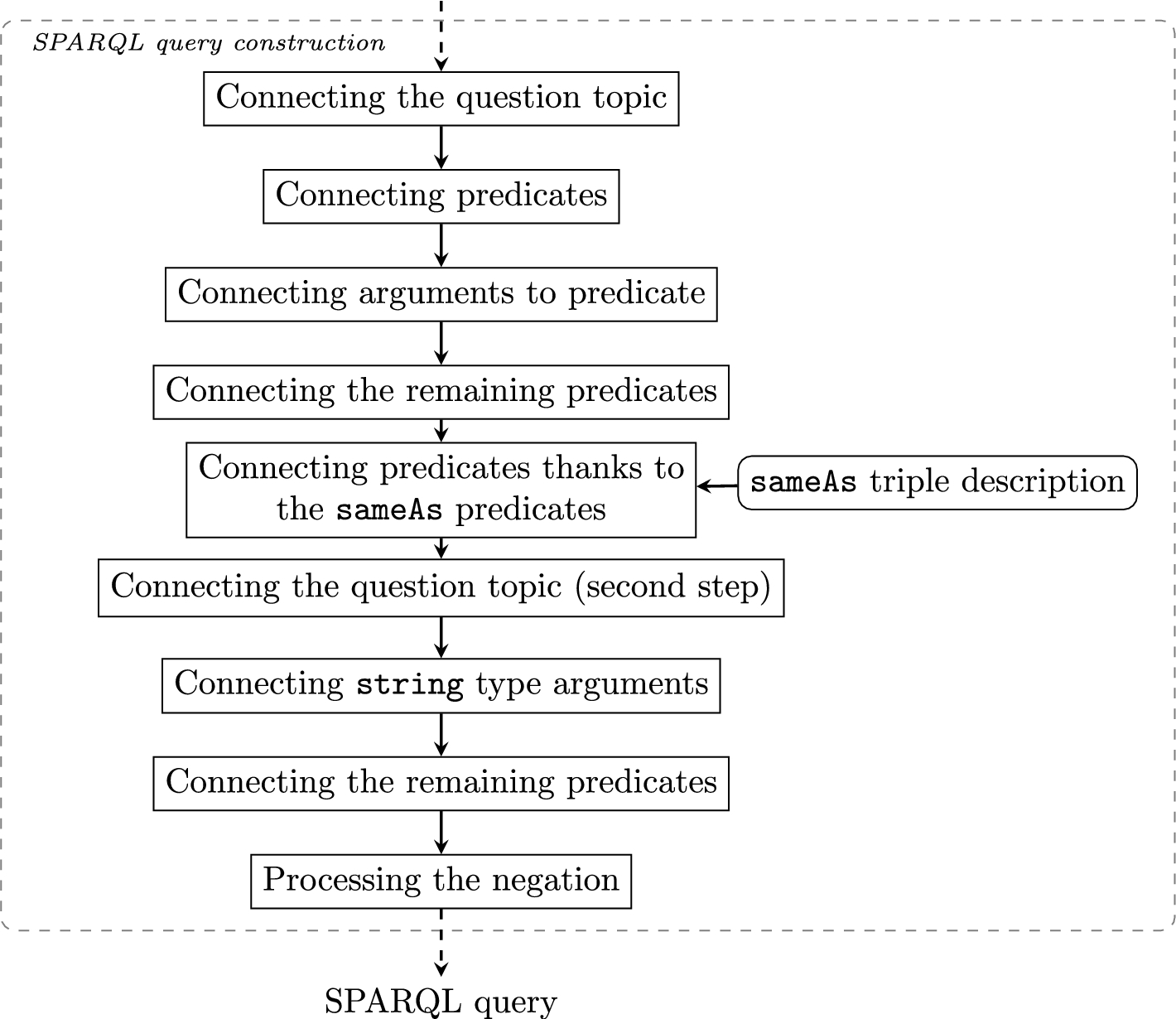
Query construction process. Square boxes represent construction steps of queries. Rounded boxes indicate the resource used.

Examples of the query construction. Graphs represent the constructed queries and consequently, the SPARQL graph patterns. Gray boxes represent subjects and objects of the predicates instantiated by the semantic entities and terms. Variables (
The objective of the
Thus, the predicate arguments are instantiated by URIs associated with the subjects, objects, variables, and numerical values or strings. For each question, we perform several associations:
The question topic is associated with one predicate argument and this is represented through a variable. Hence, this variable is associated with two elements: the question topic and one of the predicate arguments that matches the semantic type of this question topic. Notice that it is not necessary to associate all the predicate arguments that have the same semantic type with the question topic for now. Moreover, at the end of this step, the question topic may remain unassociated with any predicate. In Fig. 8(a), the
The predicate arguments are associated with semantic entities identified during the question abstraction, as they concern elements referring to URIs. Moreover, each predicate with arguments in the coordination scope is duplicated and arguments are also associated with semantic entities, if needed. Thus, in Fig. 8(a),
The predicates are associated with each other through their subjects and objects, and the association is then represented by a variable. For example, in Fig. 8(b), the subject of the
Predicates from different datasets are joined together. We use the
The remaining question topics are associated with arguments of the
The arguments corresponding to the
At this point, the predicate arguments which remain unassociated are replaced by new variables in order to avoid empty literals.
Finally, the negation operators are processed: Predicates are marked as negated, while the arguments corresponding to negated terms are included in the new
At this stage, each question is fully translated into a representation of the SPARQL query.
The SPARQL query representation built during the
The query generation process is composed of two parts:
The generation of the result form which takes into account the expected type of the result form (e.g.
The generation of the graph pattern. This part consists of the generation of strings for representing each RDF triple and the filters if the predicates are negated terms. This is the case of the examples from Figs 9(a)–(e) and 9(g). Also, in Fig. 9(d), the
The SPARQL queries are then submitted to a SPARQL endpoint10 For our experiments, we use the SPARQL endpoint provided by the QALD-4 challenge.

Examples of the query generation.
The method described above relies on
Domain-specific resources
To process the set of questions, we used the following three biomedical resources:
DrugBank11 Diseasome13
Example of natural language questions from the new test set. Sider14
The content of each resource is provided in a specific format: RDF triples of form

On the basis of the RDF triples, frames are built from the RDF schemas in which the RDF predicate is the frame predicate, and subject and object of the RDF triples are the frame elements. This also includes the OWL As indicated, subject, object and predicate become semantic entities. At least one of them must occur in questions. This way, the frames are the main resources for rewriting questions into queries; The vocabulary specific to questions is also built. It covers for instance the aggregation operators and the types of questions; RDF literals, issued from the named entity recognizer or the term extractor, complete the resources. The RDF literals are detected with specifically designed automata that may rely on the source KB documentation.
All these entities are associated with the expected semantic types which allow creating the queries and rewriting the RDF triples into SPARQL queries. In that respect, we can process several types of data (URIs, strings, common datatypes or regular expressions) when literals are expected.
Most of the entities are considered and processed through their semantic types, although some ambiguous entities (e.g., interaction or class) are considered atomically. For these, the rewriting rules are applied contextually to generate the semantic entities corresponding to the frames (see Section 4.2). When using the queries, the semantic types become variables and are used for connecting the query edges.
Experiments and results
Training and test question set
Our training set gathers the 50 questions from the training and test sets of the QALD-4 challenge. Separately, they show unbalanced complexity but taken together they provide a balanced training set. The evaluation is performed on 27 new questions. Questions from this new test set are similar to the QALD-4 questions but may differ as for the involved semantic entities or predicates. Our method is applied to this new test set without additional adaptations. Figure 10 presents a sample of questions from the new test set available at the following URL:
Evaluation metrics
The generated SPARQL queries are evaluated through the answers they generate. The evaluation is performed with the following macro-measures [30]:

Performance per sub-step for 50 questions from the training set.
Through the use of macro-measures, we equally consider all the questions independently on the number of expected answers for a given SPARQL query.
Table 1 presents the overall results obtained on the training and test sets. On the test set, the macro-F-measure is 0.78 with 0.81 precision and 0.76 recall, while on the training set, the macro-F-measure is 0.86 with 0.84 precision and 0.87 recall.
Results obtained with the training and test sets
Results obtained with the training and test sets
Our method always proposes syntactically correct SPARQL queries for all natural language questions. On the test set, concerning the answers generated over Linked Data, 20 questions provide the exact expected answers, two questions return partial answers, and five questions return erroneous answers. On the training set, 39 SPARQL queries (out of the 50 questions) provide the expected answers, six questions return partial answers, and five questions return no answers.

System performance for an increasing number of questions (1 to 50) from the training set.
Evaluation results of comparable end-to-end systems
Evaluation results from the QALD-4 challenge
We analyzed the system performance when translating 50 questions from the training set, on a computer with 4 GB of memory and a 2.7 GHz dual-core CPU. Figure 11 presents the run time for each query according to the pre-processing substeps (named entity recognition, word segmentation, POS tagging, semantic entity tagging, term extraction with  and negation scope identification with NegEx) and the question translation into a SPARQL query (Question2SPARQLQuery). Most of the processing time is dedicated to the TermTagger, which aims at recognizing the semantic entities. With the internal Ogmios processing (i.e., mainly the control of inputs and outputs), each question is processed in two seconds on average.
and negation scope identification with NegEx) and the question translation into a SPARQL query (Question2SPARQLQuery). Most of the processing time is dedicated to the TermTagger, which aims at recognizing the semantic entities. With the internal Ogmios processing (i.e., mainly the control of inputs and outputs), each question is processed in two seconds on average.
Figure 12 shows the overall system performance according to the number of questions to be processed. The variation of run time when processing one question and the whole set of questions is less than two seconds on the training set.
Findings
Overall, our approach exhibits good results, with an F-measure of 0.78 on the newly created test set. This value is lower than the F-measure obtained on the training set because no additional rules or adjustments were made for processing the questions from the new set. It is noteworthy that this value would have been higher if the
Comparison with existing work
We propose two ways for comparing our work with existing ones: either the methods or the reference data are comparable. Hence, we can compare our work with those end-to-end works presented in Section 2.2. In Table 2, we indicate the available evaluation numbers for precision, recall and F-measure. We can observe that our system provides competitive results.
We can also compare our system with those that participated in the QALD-4 challenge. In Table 3, we indicate the official results of the challenge: our system rates second among the three participating systems. As already said, the first system exploits a Grammatical Framework (GF) grammar based on formal syntax, while the third one proposes a semi-manual approach combining automatic POS tagging and manual transformation of questions into queries. The results provided by our system are close to the best system of the challenge.
In general, we can observe that the proposed method is competitive by comparison with existing work, and is also portable to new datasets.
Similar work applied on general-language datasets show less impressive results. For instance, a comparable approach (linguistic annotation, syntactic analysis and scoring of the right answers) applied to comparable material (QALD-3 DBpedia) reaches 0.32 F-measure [7]. We assume that processing of data from specialized areas, for which terminologies and semantic resources are available, provides the possibility to describe the involved concepts and scenarios with more detail. As result, the performance of automatic systems can be higher there.
Error analysis
We performed an analysis of erroneous or partial answers, other than those caused by
The annotation of semantic entities. For instance, in
The expected meaning of the terms in the questions. Semantic entities mentioned in some questions may refer to specific entities, while in other questions they may refer to general entities. For instance, in
These two main problems can be solved by using regular expressions in SPARQL graphs rather than URIs. However, we must test the influence of this modification on each query individually.
Other erroneous answers happen during the question abstraction step when the question topics are wrongly identified or when the contextual rewriting rules are not applied. Errors may also occur during the query construction step: The method may abusively connect predicate arguments and semantic entities or, on contrary, it may not consider all the identified semantic entities. Further investigations have to be carried out to solve these limitations.
Besides, during the design of queries, we had difficulties to express some constraints in SPARQL. For instance, the question
Reproducibility of our method
Our method is fully automated, once the rewriting rules have been defined. A key issue related to the reproducibility concerns the evolution of KBs. In case they are updated, it is only required to rebuild the semantic resources used for identifying the semantic entities. Yet, for managing the change of the structure of the KBs, entire frames must be regenerated. This is one direction of our ongoing research work. Moreover, the addition of new resources such as DailyMed15
The method for translating questions into SPARQL queries is implemented as a Perl module and is available at http://search.cpan.org/~thhamon/RDF-NLP-SPARQLQuery/ including the rewriting rules and frames.16 http://cpansearch.perl.org/src/THHAMON/RDF-NLP-SPARQLQuery-0.1/etc/nlquestion/SemanticTypeCorresp.rc.
We proposed a rule-based method to translate natural language questions into SPARQL queries. The method relies on the linguistic and semantic annotation of questions with NLP methods, semantic resources and RDF triple description. We designed our approach on 50 biomedical questions proposed by the QALD-4 challenge, and tested it on 27 newly created questions. The method achieves good performance with an F-measure of 0.78 on the set of 27 questions.
Further work aims at addressing the limitations of our current method including the management of term ambiguity, the question abstraction, and the query construction. Moreover, to avoid the manual definition of the dedicated resources required by our approach (frames, specific vocabulary and rewriting rules), we plan to investigate how to automatically build these dedicated resources from the RDF schemas of the Linked Data set. This perspective will also facilitate the integration of other biomedical resources such as DailyMed or RxNorm [24], and the use of our method in text mining applications.
Footnotes
Acknowledgements
This work was partly funded through the project POMELO (
