Abstract
Infectious diseases can inflict immense losses and suffering on the human population. As at 23
Introduction
The Coronavirus Disease 2019 (COVID-19) is an emerging respiratory infectious disease caused by SARS-CoV-2 (also known as 2019-nCoV), which first occurred in early December 2019 in Wuhan, China. According to the [1], COVID-19 situation report of 23
The total cases of COVID-19 in Nigeria. Source: Worldometer, Accessed: June 23, 2020.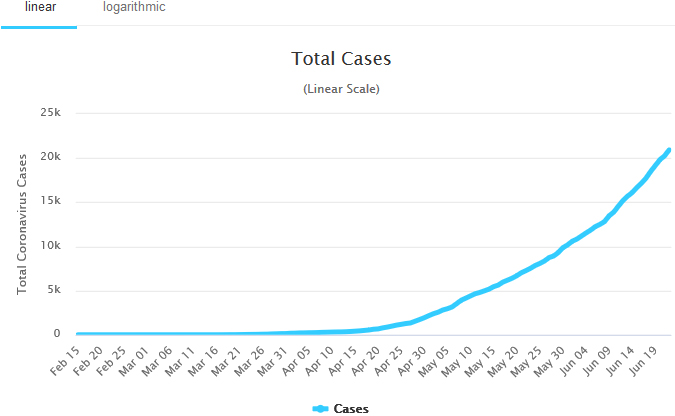
The active cases in Nigeria. Source: Worldometer, Accessed: June 23, 2020.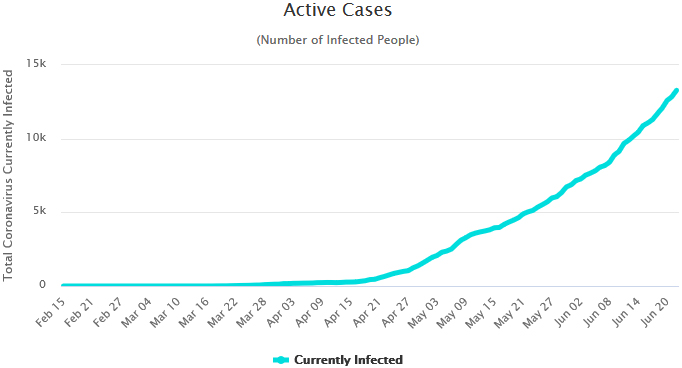
Daily deaths in Nigeria. Source: Worldometer, Accessed: June 23, 2020.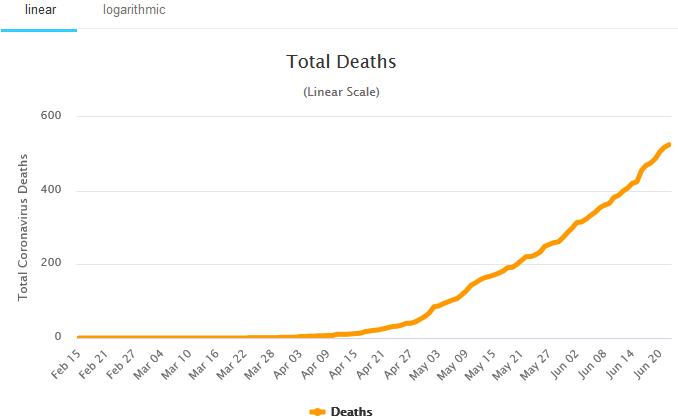
Adaptive Cluster Sampling (ACS) is a sampling design that target hidden events that are spatially inconsistent, and can be used to collect valuable information with respect to temporal and spatial characteristics of individuals and areas [5] reviewed in [6].
Adaptive designs can offer significant ethical and cost advantages over standard fixed techniques. Naturally, adaptive sampling scheme maximizes survey effort where it is most valuable and minimizes survey effort elsewhere. It performs better for sparse and clustered populations than standard grid cell sampling methods.
Authors have modeled COVID-19 cases, but there is a dearth of information on estimating the total number of hidden COVID-19 carriers in Nigeria. Adaptive cluster sample was used for exploring populations of hidden COVID-19 carriers in Nigeria. The data on daily cases of COVID-19 were extracted from NCDC website. Nigeria population was partitioned into 37 regions (36 states and FCT). We considered a model based approach in Bayesian framework to estimate the number of COVID-19 carriers in Nigeria.
Let Nigeria population (R), be defined to consist of 774 local government areas (LGA). Then,
Considering that the COVID-19 carriers are sparse and clustered, let
Based on the above prior knowledge, we considered a Bayesian-based model, the prior distribution of
Region inclusion probability
Suppose Nigeria population U with COVID-19 carriers is partitioned into regions (36 States and FCT), then
Assume
Region inclusion probabilities based on an initial simple random sample
The network inclusion probabilities based on an initial simple random sample (SRS) will be as follows.
Let
For a region of size
When including network
A model for number of COVID-19 carriers in 36 states and FCT of Nigeria
We considered Nigeria population (
The variables
Autocorrelation plot.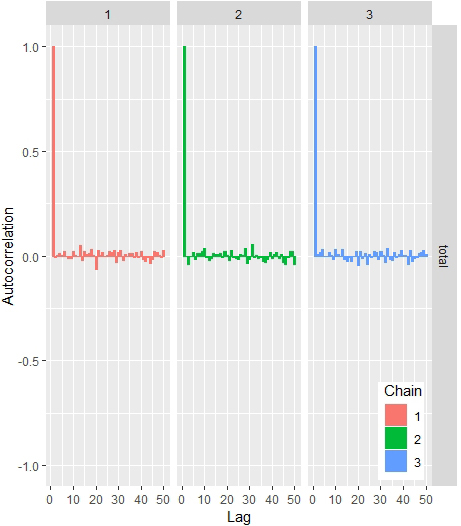
As June 22
We have three unknown parameters
Confirmed cases and estimates
Confirmed cases and estimates
Gibbs sampler was used to fit the joint distributions with the following steps;
State initial values for the unsampled components Generate Generate Generate Generate ( Iterate from ii.
The R package was used to fit the models and Winbugs by [11] software was also used to implement the MCMC algorithm.
MCMC diagnostics
The main goal of MCMC is to gather a representation of the true but incalculable posterior distribution. The diagnostics give confidence that samples are converging to a reasonable representation of the targeted posterior. In this study, Trace plot, correlogram, Geweke, Gelman-Rubin and Raftery-Lewis were used for the diagnostics check.
Autocorrelation plot
This is useful to check to get how correlated the proposed parameters are through the iterations, there is need to plot the autocorrelation of the chains. It will give the sense of the efficiency of the MCMC algorithm.
Effective sample size
The effective sample size in this context is defined as the appropriate number of people to be tested for COVID-19 that could give actual estimate of total number of COVID-19 carriers in Nigeria. The effective sample size,
where
Trace plot.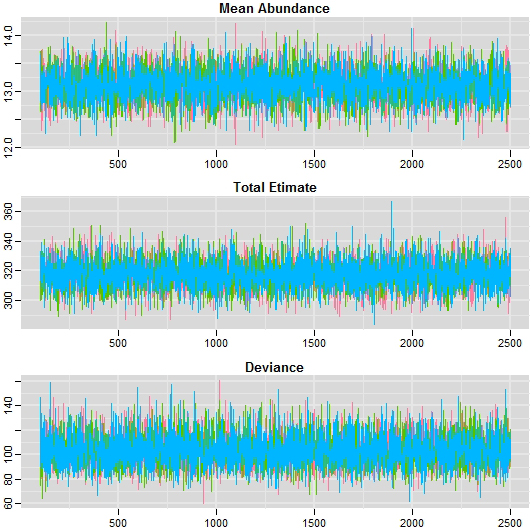
MCMC kernel density estimate plot.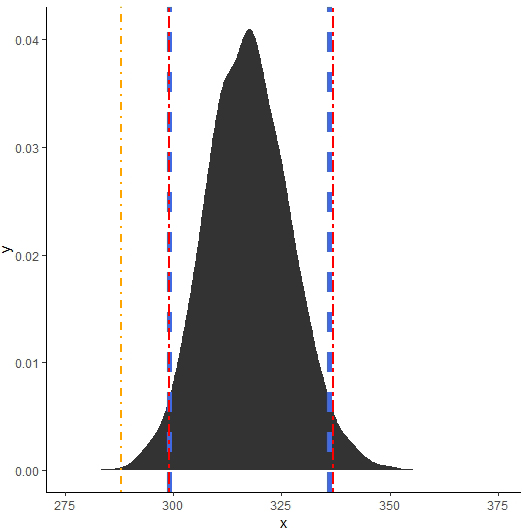
The fitted model showed that the sample were converged to reasonable representation of the target posterior (Fig. 6). From diagnostics check, the trace plots showed that the chains were mixed extremely fast and concentrated around the mode of the posterior distribution and also stationary as shown Figs 5 and 6. Plotting the Autocorrelogram in Fig. 4, the three chains autocorrelation is quite similar at each lag.
Table 1 shows the number of tested individual for COVID-19, the confirmed COVID-19 cases, estimated total of hidden COVID-19 carriers in Nigeria, the appropriate number of individual to be tested in Nigeria such that all carriers would be in the selected samples and the remaining number of unidentified COVID-19 carriers in Nigeria for days 1
The relevance of the estimation is to guide statisticians to know the estimated number of COVID-19 carriers within a population together with appropriate sample to be selected strategically and adaptively such that all carriers would be captured. Adaptive Cluster Design is a specialized sampling procedure for identifying hidden events such as COVID-19 carriers in a population. The proposed approach and contact tracing can work concretely by involving Statisticians in the contact tracing team. Statisticians will use their professional skills to handle all challenges of not at home, hard to reach, non- response due to sensitive issues, hard core, lack of knowledge of the estimated number of COVID-19 carriers and lack of appropriate knowledge and strategy by medical personnel for identify Covid-19 carriers during contact tracing.
Conclusion
We have developed a model for estimating total numbers of COVID-19 carriers in Nigeria. The fitted model shows that all COVID-19 carriers will only be captured at once if contact tracing is combined with methodology designed in this work. The existing contact tracing team lacks involvement of Statisticians. This work recommends involvement of Statisticians in contact tracing team so as to handle all challenges of not at home, hard to reach, non- response due to sensitive issues, hard core, lack of knowledge of the estimated number of COVID-19 carriers and lack of appropriate knowledge and strategy by medical personnel for identify Covid-19 carriers during contact tracing.
Footnotes
Acknowledgments
We thank the National Center for Disease Control in Nigeria for making number cases of COVID-19 patients available online. We also acknowledge the reviewers for their constructive corrections.
