Abstract
Phenotype is the set of observable traits of an organism or condition. While advances in genetics, imaging, and molecular biology have improved our understanding of the underlying biology of Parkinson’s disease (PD), clinical phenotyping of PD still relies primarily on history and physical examination. These subjective, episodic, categorical assessments are valuable for diagnosis and care but have left gaps in our understanding of the PD phenotype. Sensors can provide objective, continuous, real-world data about the PD clinical phenotype, increase our knowledge of its pathology, enhance evaluation of therapies, and ultimately, improve patient care. In this paper, we explore the concept of deep phenotyping—the comprehensive assessment of a condition using multiple clinical, biological, genetic, imaging, and sensor-based tools—for PD. We discuss the rationale for, outline current approaches to, identify benefits and limitations of, and consider future directions for deep clinical phenotyping.
Keywords
INTRODUCTION
Two centuries after its seminal description, Parkinson’s disease (PD) is now the world’s fastest growing major neurological disorder [1, 2]. Progress in genetics, imaging, and molecular biology have increased our understanding of the condition [3–5]. However, despite advances in technology, many basic clinical features of the disease remain elusive to specialists and researchers. This ignorance impairs our efforts to identify the etiologies of the disease, evaluate new therapies, and provide better care [6, 7].
In this paper, we examine the shortcomings in our current understanding of PD, introduce the concept of deep phenotyping, highlight existing efforts, introduce our own research, examine the benefits and limitations of this new approach, and discuss future directions.
SUPERFICIAL UNDERSTANDING OF PARKINSON’S DISEASE
Today, the principal means for assessing PD are categorical, episodic assessments conducted in the clinic [8]. The 99.9% of the time during which individuals with PD are not directly observed by clinicians, until recently, has not been assessed [9, 10]. Diaries completed by patients concerning their response to medications provide a window into how individuals are affected by the disease in their natural environment, have helped evaluate the benefits of new treatments, and have been the basis for new approvals [11–13]. However, these diaries are episodic, burdensome, primarily focus on motor function, and often require individuals to evaluate their disease using unfamiliar clinical terms.
In the 19th century, Drs. Parkinson, Charcot, and Gowers provided detailed descriptions of PD based on observation and examination [14–16]. However, basic characteristics, such as what proportion of the day an individual with PD has tremor, are known only to those affected by the disease. These measures, which extend beyond the time that individuals are directly observed in clinic, are poorly characterized in the scientific literature by wide-ranging estimates.
For the past decade, accelerometers have helped quantify motor features of PD [17–22]. However, most of these studies (Table 1) have limited their assessments to tasks performed in the clinic [18]. A 2018 systematic review of 24 wearable sensor studies in PD found only seven that assessed the disease in a “free-living environment” [18].
Large wearable sensor studies in Parkinson’s disease, 2015–2019
Source: PubMed search of wearable sensors to assess neurological conditions, filtered by sample size greater than 100, on 11/15/2019 (“parkinson disease” [MeSH Terms] OR (“parkinson” [All Fields] AND “disease” [All Fields]) OR “parkinson disease” [All Fields] OR (“parkinson’s” [All Fields] AND “disease” [All Fields]) OR “parkinson’s disease” [All Fields]) AND (wearable[All Fields] AND (“Sensors (Basel)” [Journal] OR “sensors” [All Fields])).
Wearable sensors and smartphone embedded sensors are providing glimpses into how PD affects individuals in everyday life [23]. A recent ambulatory monitoring study of individuals with newly diagnosed PD employed a wrist-worn sensor (Global Kinetics; Parkville, Australia) and detected clinically meaningful tremor among a few participants for over half the day. Studies of smartphone embedded sensors, some large-scale, have captured remote, objective assessments of PD (Table 2). In general, though, these studies are limited by small sample size and short duration, rely on participants to conduct active tasks (e.g., tapping on the screen), and lack companion sensors which can paint a more complete picture of the disease [24].
Large smartphone studies in Parkinson’s disease, 2015–2019
Source: PubMed search of smartphones using sensors to assess neurological conditions, filtered by sample size greater than 100, on 11/15/2019 (“parkinson disease” [MeSH Terms] OR (“parkinson” [All Fields] AND “disease” [All Fields]) OR “parkinson disease” [All Fields] OR (“parkinson’s” [All Fields] AND “disease” [All Fields]) OR “parkinson’s disease” [All Fields]) AND (“smartphone” [MeSH Terms] OR “smartphone” [All Fields] OR “smartphones” [All Fields]).
Despite advances in our ability to assess motor features using technology, details, like when the tremor occurs, for how long, and at what amplitude and frequency, are still lacking. In addition, most sensor studies only assess tremor in one limb, even though its presence in multiple limbs has been known for centuries [14, 25].
Similarly, the extent of intra- and inter-day disease variability remains largely unassessed in the 21st century. A recent smartphone study found that the magnitude of inter-day variability of motor impairment was far greater than the overall average progression of motor features over six months [26]. Variability in non-motor features like sleep, neuropsychiatric, and autonomic dysfunction is even less well characterized. Moreover, assessing the impact of these features on the lives of individuals with PD is almost exclusively limited to pen-and-paper surveys [27–29].
DEEP PHENOTYPING OF PARKINSON’S DISEASE
What PD needs is deep phenotyping. Phenotype is a set of observable traits of an organism or condition [30]. In medicine, phenotyping comes from a medical history, physical examination, imaging, and blood tests. Deep phenotyping “shows the different dimensions of a disease” and leverages all available data sources to gather details specific to the individual [30]. Current phenotyping efforts that rely on subjective characterizations about an individual’s walking ability, for example, have been described as “sloppy or imprecise” [30]. In a 2015 Nature piece, deep phenotyping is described as gathering “details about disease manifestations in a more individual and finer-grained way, and uses sophisticated algorithms to integrate the resulting wealth of data with other kinds of information” [31]. This approach results in large data sets that require data fusion algorithms to integrate the results [30, 32].
Many deep phenotyping efforts have used molecular tools, such as genomics, proteomics, and metabolomics, or electrophysiological tools, like electroencephalography, to develop new insights [33, 34]. Such studies have advanced our understanding of pathophysiology in diseases ranging from diabetes to osteoarthritis [35, 36]. In addition to evaluating those with a disease, this approach can track individuals as they transition from a healthy to a diseased state [35].
In this paper, we instead focus on deep clinical phenotyping, which can be integrated with molecular approaches. Deep clinical phenotyping has four principal characteristics. First, the effort begins with the history and examination as in a traditional clinical appointment. Second, deep clinical phenotyping uses sensors or other tools to provide objective measurement of the individual or the condition. Third, multiple domains (e.g., activity and respiratory function) of the disease of interest are assessed. Fourth, these assessments extend to real-world settings like the home. The depth of the phenotype is a function of the quality of the assessment of each domain, the number of areas evaluated, and the duration of observation.
New sensors can generate large volumes of data that are measured in number of observations per person per second [37]. These tools can thus provide high-definition descriptions of previously unobservable clinical features. For example, a recent study used digital sensors, including a smartphone, smartwatch, a digital assessment application, and a bed sensor, to provide a “minute-level behaviorgram” of individuals with either Alzheimer’s disease, mild cognitive impairment, or no evidence of cognitive deficit [38]. This behaviorgram combined assessments of physical activity (e.g., steps taken), sleep (e.g., duration of sleep), and phone usage (e.g., text messages received) to a paint a picture of how cognitive impairment affects individuals in their daily lives.
CURRENT DEEP PHENOTYPING STUDIES IN PARKINSON’S DISEASE
Deep clinical phenotyping studies in PD have begun (Table 3) [39, 40]. For over a decade, the Parkinson’s Progression Markers Initiative has sought to identify biological markers of disease progression and in so doing has collected a wealth of clinical, genetic, imaging, and biological data [41]. Recently, it added sensors, including a smartwatch and a smartphone application, to the current clinical assessments. The expectation is that the sensor data will provide more objective and frequent assessments of PD both in and outside the clinic. Together with the biological assessments, a more complete picture of cohorts with and at risk for PD can be created.
Current studies of deep phenotyping in Parkinson’s disease
Additional studies are using smartwatch embedded sensors to assess PD. The 650-person Personalized Parkinson Project in the Netherlands has started enrolling individuals with PD who will be followed annually in clinic and with a smartwatch [39]. In addition, the study is collecting multiple biological samples, including blood, cerebrospinal fluid, and stool, and capturing structural and functional MRIs from research participants. The study aims to address the stagnation in our understanding of the etiology, pathophysiology, and progression of PD by developing “deep and repeated multi-dimensional phenotyping” of PD enabled by continuous monitoring with a wearable device for two years [39].
A separate U.S. study called Watch-PD is using a different smartwatch along with wearable sensors in the clinic to assess PD. In contrast to the Personalized Parkinson Project, which has broad inclusion criteria, Watch-PD is focused on developing novel digital assessments of individuals with early, untreated PD [42]. The hope is that the resulting digital assessments could be used as outcome measures or endpoints in future clinical trials in this population.
The Luxembourg Parkinson’s Study is another deep phenotyping study that is addressing the “substantial gaps in our understanding of the underlying mechanisms and the complex clinical presentation” of PD [43]. To do so, the Luxembourg study is utilizing a smartphone application and a shoe sensor to develop “rater-independent” measures of the clinical disease features. These will be combined with detailed biological assessments from at least blood, saliva, and urine to develop a “data driven approach” to evaluate the disease [43].
A NEW EFFORT
To complement these efforts, we launched a study, Super-PD, that uses multiple sensors to enable deep clinical phenotyping of individuals with and without PD (Fig. 1). We are enrolling fifty individuals, thirty-five with PD, and fifteen age- and sex-matched controls in a two-year prospective cohort study.
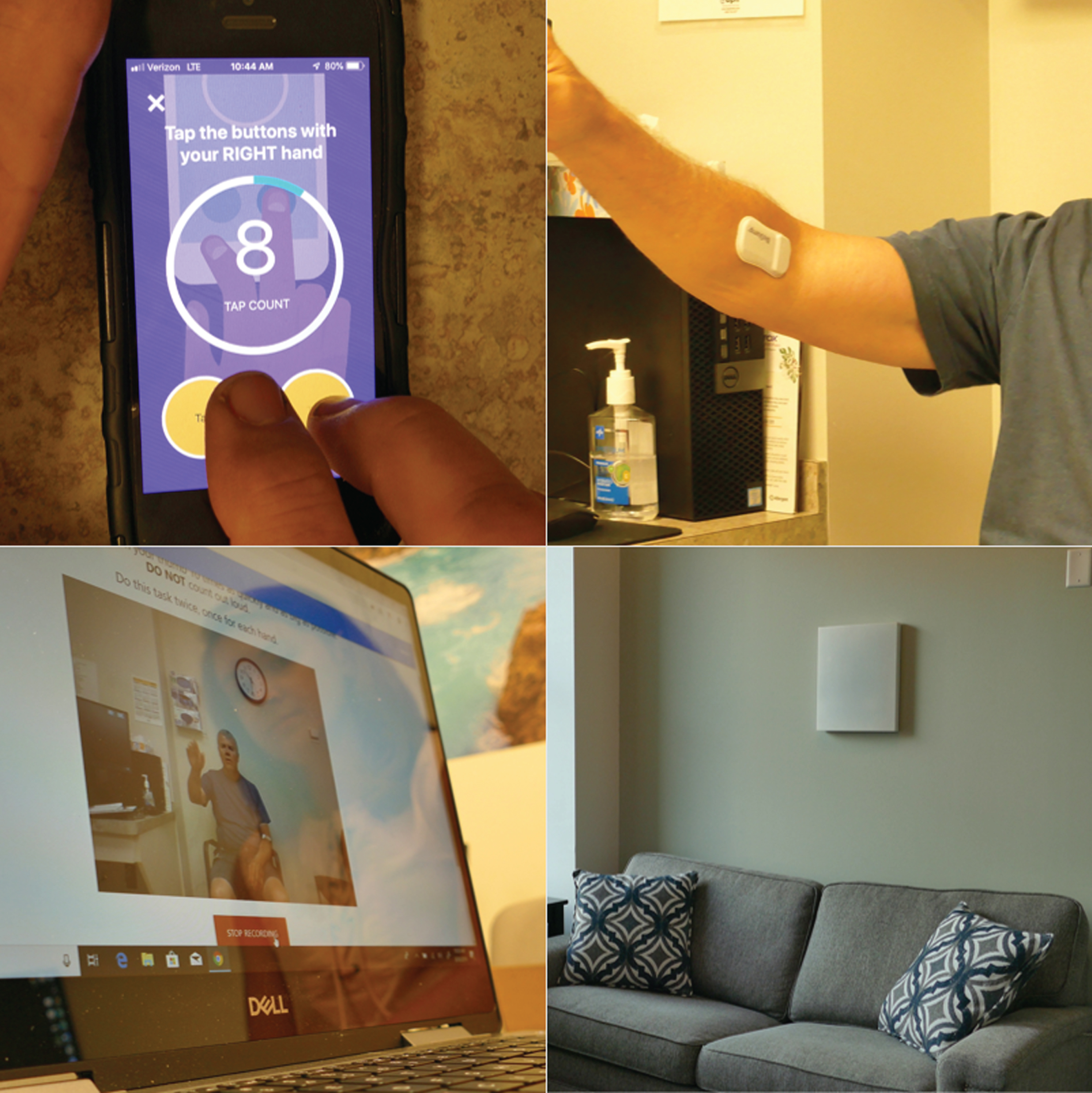

Picture of the digital devices that will be used in our deep clinical phenotyping study.
Compared to prior work, this study combines multiple sensors that significantly increase the depth and breadth of signals collected. The first measure is the second generation of the mPower smartphone application originally released on Apple’s open-source ResearchKit platform in March 2015 [44]. The initial study enrolled over 15,000 participants with and without PD throughout the U.S. and used the smartphone’s sensors to assess voice, finger tapping, gait, and balance [44]. The second- generation application is much like the first but has additional tremor assessments and allows for passive monitoring using the Global Positioning System (GPS). Analysis of results from a similar application with similar assessments generated a “mobile Parkinson’s disease score,” which allows a frequent, objective assessment of disease severity by anyone with a smartphone anywhere in the world [26, 45].
The second measure is a set of wearable sensors (MC10; Lexington, MA) that can be placed almost anywhere on the body. In an initial study, five sensors (one on each limb and one on the chest) were used to quantify what proportion of a day individuals with PD are lying, sitting, standing, or walking [25]. These sensors can also assess tremor, gait, sleep, and potentially dyskinesias. The second generation of the sensor adds electrocardiogram (ECG) capabilities. Compared to the smartphone assessment, the wearable sensors, which will be applied using double-sided adhesives to the chest and most affected arm and leg, provide a more accurate measure of movement for the body parts where they are placed. However, over the course of the study, participants will likely wear the sensors less often than they carry their smartphone.
The third assessment is a video analytics tool, broadly available via computer browser, which uses machine learning rather than raters to assess movements in PD. Participants are asked to perform elements of the motor portion of the Movement Disorder Society—Unified Parkinson’s Disease Rating Scale (MDS-UPDRS) in front of a web camera on a laptop computer [8]. Using machine vision, a computer algorithm can automatically and precisely identify subtle movement of the facial muscles, hands and fingers and characteristics of spoken words and syllables and correlate these features with manually annotated MDS-UPDRS scores. Based on a task that asks participants to maintain a neutral face, the computer algorithm has identified facial microexpressions around the lips and eyebrows that can differentiate those with PD from those without [46].
Finally, the Super-PD study utilizes a radio wave sensor called Emerald (Cambridge, MA) that is installed in an individual’s home and uses electromagnetic waves to assess individuals in their natural environment [47, 48]. Low power radio waves transmitted by the device reflect off non-stationary objects, most notably people and pets, and are used to detect location, movement, gait trajectories, sleep, and other activities in the home. An initial study quantified gait speed and tracked sleep patterns, including the time, duration, and interruptions when individuals were in bed [47]. The entirely passive sensor monitors up to a range to of 1,200 square feet (110 square meters) and works best when only 2–3 individuals regularly occupy the space.
Participants will also undergo in-clinic assessments with traditional rating scales to assess motor function, mood, sleep, and cognition and provide a blood sample to be stored for analysis as part of the Parkinson’s Disease Biomarker Program [8, 49–54]. The scales will be augmented by the recently developed, online Patient-Reports of Problem (PROP) assessment. The PROP assessment, which was developed to give participants the opportunity to express themselves in their own words, covers three domains: general health, psychosocial wellbeing, and Parkinson’s disease. Each PROP survey asks individuals what bothers them, how the problems affect daily functioning, the severity of the problems, and what they do to alleviate each problem [55]. Natural language processing, standardized clinical curation, and machine learning are used to quantify and analyze responses [56]. Together, the PROP and digital measures will give a robust portrayal of what individuals with PD say and do.
BENEFITS OF DEEP CLINICAL PHENOTYPING
Deep clinical phenotyping can provide new insights into various aspects of PD (Table 4). This approach uses sensors to examine how PD affects people in their daily lives and extends beyond biological markers to give a more complete characterization of the disease.
Deep phenotyping measures of Parkinson’s disease
Smartphone applications can detect subtle tremor that is missed by trained raters [57]. In addition, smartphone embedded sensors can readily assess gait and global motor function. In one study, a smartphone application detected that individuals with PD walked about 25% less than those without the disease [57].
Wearable sensors can also quantify gait speed, which is reduced in PD and fluctuates in response to levodopa and deep brain stimulation [58–62]. In geriatric medicine, “gait speed” is the “sixth vital sign” [63], and reductions are associated with increased mortality [64, 65]. For the past decade, Dr. Jeffrey Kaye at Oregon Health Sciences University and colleagues have outfitted homes of older adults with multiple sensors to assess motor, functional, and social behavior [66]. They have quantified the gait speed of individuals in their natural environment, and they and others have found that declines in walking speed predict cognitive impairment and dementia [66–68]. Similarly, reductions in daily physical activity are associated with an increased risk of Alzheimer’s disease and potentially PD [69–71].
Multiple non-motor features, including autonomic function and sleep, can also be evaluated. The physiologic oscillation in the time interval between heartbeats, known as heart rate variability, is reduced in PD [72–74]. Prior studies of heart rate variability in PD have collected observations on up to 150 participants for 24 hours with an ambulatory ECG, generating around 3600 hours of data [72, 75]. In contrast, deep phenotyping studies such as ours, can generate data volumes that dwarf more traditional studies with serial ECG assessments over longer periods of time (e.g., weekly intervals). In addition, as with motor features, the effect of exercise, medication, walking, and sleep on heart rate response can be assessed.
Sleep has also been inadequately assessed in PD. Most studies use polysomnography for single night observations in artificial sleep labs [76–78]. Accelerometers have helped measure different aspects of sleep at home [17, 18]. These studies have quantified various aspects of sleep and have found that individuals with PD turn fewer times but leave the bed more often [79, 80]. In our study, the number and duration of naps, duration of sleep, number of sleep interruptions, and large movements during sleep will all be quantified. The latter could be used to assess rapid eye movement (REM) sleep behavioral disorder, which is an early feature of the disease that can be identified by multiple digital devices [81–83].
Sensor technology also enables functional assessments in the home. The Emerald device can quantify number of visits to the bathroom and time spent in the bathroom. Such assessments can provide new insights into the impact of urinary urgency and constipation and supplement current measures of these symptoms, which rely on diaries, surveys, and patient-reported outcomes [84].
Finally, this study will provide novel insights into how PD disease affects an individual’s interaction with the environment or social function. A small smartphone study used GPS to measure the “lifespace,” or the “geographic area in which a person lives and conducts their activities,” of individuals with PD [85]. The lifespace can be mapped in a de-identified way that measures time and distance away from home without giving the specific location of participants. Such assessments could reflect both motor (e.g., mobility) and non-motor (e.g., depression) features of PD. Inside the home, the radio-wave sensor can quantify how much time is spent at home, where it is spent (e.g., bedroom), and what proportion of time an individual is alone. In older adults, increased time outside the home is associated with superior physical ability and improved emotional state [86]. Conversely, loneliness is associated with decreased time out of the home, motor slowing, functional decline, and death [87–89]. Loneliness not only affects individuals with PD but also their caregivers [90, 91].
In essence, sensors increase the precision of established measures to capture greater variability in symptoms and enable more frequent, seamless data collection. Within the context of deep clinical phenotyping studies, sensors can identify new outcome measures of PD and generate composite digital biomarkers that holistically measure disease symptoms and progression.
LIMITATIONS OF DEEP CLINICAL PHENOTYPING
New efforts confront many barriers and limitations. Among them are who participates in such research studies, their privacy, the ability to capture their data, the validity of the data, and their subsequent analysis and generalizability. Thus far, participants in deep phenotyping studies have not been representative of the general population. They are overwhelmingly white, well educated, and likely have more trust in clinicians and health care institutions than other groups [92]. Social and economic factors also contribute to the digital divide, with differential access to the internet and digital devices [93]. Engaging in outreach, reducing financial and travel burdens, and bringing research tools to participants (e.g., direct-to-consumer genetic testing, smartphone applications, online research platforms) can address some of this selection bias, which plagues both deep phenotyping and traditional research studies [94].
The privacy of participants is another concern. Deep phenotyping requires extensive monitoring of individuals, including continuous passive location assessment and monitoring in their homes. Despite the intrusion, studies that have assessed privacy concerns have generally found only modest concerns [23, 47]. That said, these responses come from those who are willing to share de-identified data. Privacy concerns may also be less among those with PD, who often are either retired or disabled, have guaranteed health insurance (e.g., Medicare in the U.S.), and are invested in clinical research. Privacy views may differ for those without or at risk for PD, who are likely to be younger, employed, and less secure in their health insurance [95].
Some sensors inform researchers not only about the behavior of research participants but also about those that live with them or come in close contact to them. Sensors that allow for easy pausing of data collection could help protect privacy. In addition, the Emerald device, for example, filters all movement patterns to isolate those from just the study participant. In other cases, sensors may collect information about all activity in its range. In these cases, researchers need to ignore such data where feasible and discard it where not. To assist with the conduct and oversight of such studies, ethicists, consent specialists, and voices of those affected by the disease are also valuable, if not essential.
Interest in studies utilizing sensors is high, including among older individuals [18, 97]. However, maintaining that interest is difficult as participation often wanes quickly [44, 98]. Supporting and engaging participants throughout a study’s course is thus important. In addition, capturing data from participants in uncontrolled (real-world) environments is not easy. Poor connectivity (e.g., to Wi-Fi) and limited storage capacity of sensors are potential obstacles that need to be overcome.
While these digital devices are appealing, additional work is required to ensure their validity. Validation is the “process of ensuring that the digital measurement tool is meeting its intended use by generating objective data that accurately represents the concept of interest... that it purports to be measuring” [99]. Validation seeks to answer whether researchers built the right tool to assess measures of how someone feels, functions, or survives, such as gait. Validity is divided into two components, analytic validation and clinical validation [99, 100]. The former assesses whether algorithms accurately process the data (i.e., is the calculation of gait speed from accelerometry data true?). The latter evaluates whether the measurement (e.g., gait speed) reflects an important health domain. Large scale public-private partnerships, such as Mobilise-D, are seeking to develop clinically valid digital mobile assessments that can be applied to PD and other chronic medical conditions. These efforts in multiple sclerosis, for example, employed digital measures alongside traditional measures to assess ambulation in the clinic and in a real-world setting [101]. While larger scale studies and rigorous validation of novel digital measures is needed for PD, the field should recognize that comparison against traditional measures may not be the best approach for such validation, particularly because many of the digital measures can be more accurate and sensitive than traditional “gold” standards. After confirming their reliability and correlation with clinical scales, digital measures may be directly validated against pathophysiological biomarkers in PD.
Finally, analysis of data from deep phenotyping studies requires new methodologic and analytic approaches and new collaborations. Deep phenotyping studies generate large datasets that are characterized by high volume, variety, velocity, veracity, and value [102]. Each of these characteristics comes with its own challenges that require multi-disciplinary teams to address. Such data is often highly contaminated with irrelevant factors such as idiosyncratic behaviors, environmental settings and changes, and effects of comorbid conditions. Analysis needs to account for this substantial noise in the data [103]. Hybrid approaches that combine techniques from multiple fields will be essential. In addition to clinicians, research teams often need computer scientists (artificial intelligence and machine learning), data scientists (high dimensional data), electrical engineers (signal processing), and persons with PD to manage and interpret the data.
FUTURE DIRECTIONS
Deep phenotyping offers at least three great potential advances. First, deep clinical phenotyping will enhance our understanding of the nature of the disease, including its expression, pathophysiology, and potential modulators [7, 104]. Detailed, frequent, real-world assessments of PD will provide a more complete picture of the disease’s manifestations, their sequence of development, their impact on individuals’ everyday life, and their relationship to pathology. Current PD phenotypes are based on traditional scales and likely incomplete, if not inaccurate [105]. What we see in our clinic today is an arbitrary snapshot that may not reflect the real-world. New tools can monitor features of the disease inside and outside the clinic. They will allow us to identify new potential modulators of Parkinson’s course, such as weather, altitude, diurnal patterns, sleep, activity, and other factors that have either not been considered or are difficult to measure.
Second, deep phenotyping could help us evaluate new treatments objectively and efficiently. Current PD clinical trials are large, long, expensive, and prone to fail [106, 107]. In this century, the U.S. Food and Drug Administration (FDA) has approved only three new classes of medications for PD (adrenergic agonists, atypical antipsychotics, and adenosine A2A receptor antagonists), each of which benefits a small subset of patients. Moreover, despite extensive discussion and large efforts [108–110], no therapies that target the underlying pathology have emerged. A new approach is needed, and deep phenotyping could generate the objective, sensitive, and high-definition assessments required to evaluate a new generation of therapies [17].
Such approaches are gaining traction. In the U.S., the 21st Century Cures Act introduced the use of real-world data, including sensor data, to support regulatory decision making [111]. The inclusion of real-world data and the resulting real-world evidence reflects the rapid acceleration in the “use of computers, mobile devices, wearables and other biosensors to gather huge amounts of health-related data” [111]. These data can, according to the FDA, “allow us to better design and conduct clinical trials and studies in the health care setting to answer questions previously thought infeasible” [111]. In 2019, the FDA issued a guidance document that indicated that real-world evidence could be used to “provide evidence in support of the effectiveness or safety of a new drug approval” [112] and a similar guidance for devices [113]. Also in 2019, the European Medicines Agency deemed top stride velocity (95th percentile measured at the ankle) as “an appropriate endpoint in studies to support regulatory decision making on medicines for the treatment of Duchenne Muscular Dystrophy” [114]. The FDA has already approved one treatment (dalfampridine) for multiple sclerosis based on gait speed [115, 116]. Such measures could be readily applied to PD.
Third, a prerequisite for personalized care for PD is individual data [117]. These data can be patient-reported, genetic, biological, or clinical. In a 2013 review, Dr. Walter Maetzler and colleagues predicted such an application. They wrote, “Measuring (Parkinson’s) disease-related outcomes objectively ... , continuously ... , unobtrusively ... , and with high ecological validity ... could boost the efficiency and relevance of (patient) visits, and improve patient care. This clinical wish appears to [be] coming within reach with the advent of new, wearable technology that can quantitatively collect, analyze, and deliver data to both the patient and the doctor” [17].
Personalized medicine for PD is just beginning. Current treatment decisions are largely based on ad hoc assessments [118, 119]. Other diseases are further ahead. Individuals with diabetes are no longer dependent on in-clinic assessments to dose their insulin. Now, blood glucose levels are measured continuously and dosing of medication is tailored to these data in real time, resulting in greater potential for optimal disease control. Future objective assessments of PD (e.g., of gait) could enable tailoring of medical or surgical treatments to the individual [120, 121]. Deep brain stimulation parameters, for example, could be automatically adjusted based on continuous, real-world assessments from implanted, sensing leads, a process that is underway [122–124]. Patients could customize their activities and treatments based on sensor data and foster self-efficacy [125].
Fourth, deep phenotyping will enable the discovery of new disease subtypes [126]. Clustering individuals into groups with similar progression or traits can power new studies that seek to understand the biological differences across these groups. The future (Fig. 2) will see expanded deep phenotyping efforts. These may include generating phenotypes specific to genetic sub-types of PD (e.g., due to LRRK2 or GBA mutations) that could inform gene-directed clinical trials [127]. Previous studies using traditional rating scales have suggested that these genetic sub-populations have different features and progression rates [128, 129]. Deeper phenotyping will likely be able to confirm or refute such findings and identify many other hidden differences. Deep phenotyping of individuals at risk for or with prodromal PD will permit better understanding of the nature of PD and its evolution. This will be especially important for assessing the effectiveness of novel interventions aimed at these groups.
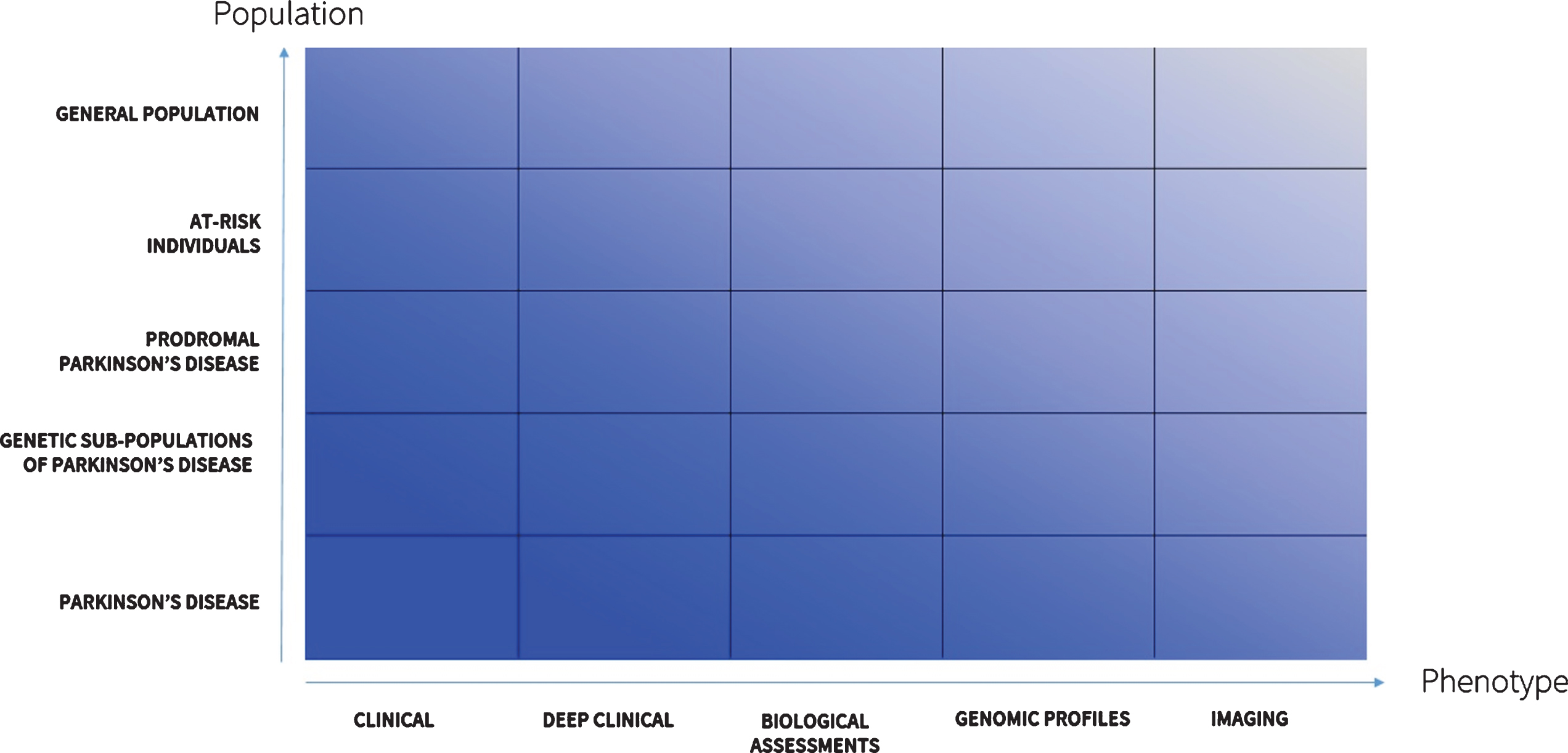
Landscape of deep phenotyping of Parkinson’s disease.
The mathematician Dr. Freeman Dyson said “New directions in science are launched by new tools much more often than by new concepts. The effect of the concept-driven revolution is to explain old things in new ways [6]. The effect of a tool-driven revolution is to discover new things that have to be explained” [130]. We now have these new tools. New insights await us.
CONFLICTS OF INTEREST
Ray Dorsey has served as a consultant to 23andMe, Abbott, Abbvie, American Well, Biogen, Clintrex, DeciBio, Denali Therapeutics, GlaxoSmithKline, Grand Rounds, Karger, Lundbeck, MC10, MedAvante, Medical-legal services, Mednick Associates, National Institute of Neurological Disorders and Stroke, Olson Research Group, Optio, Origent Data Sciences, Inc., Prilenia, Putnam Associates, Roche, Sanofi, Shire, Sunovion Pharmaceuticals, Teva, UCB, and Voyager Therapeutics. He has received honoraria from the American Academy of Neurology, American Neurological Association, and University of Michigan. He has received research support from Abbvie, Acadia Pharmaceuticals, AMC Health, BioSensics, Burroughs Wellcome Fund, Davis Phinney Foundation, Duke University, Food and Drug Administration, GlaxoSmithKline, Greater Rochester Health Foundation, Huntington Study Group, Michael J. Fox Foundation, National Institutes of Health/National Institute of Neurological Disorders and Stroke, National Science Foundation, Nuredis, Inc., Patient-Centered Outcomes Research Institute, Pfizer, Prana Biotechnology, Raptor Pharmaceuticals, Roche, Safra Foundation, Teva Pharmaceuticals, and University of California Irvine. Dr. Dorsey provides editorial services for Karger Publications has ownership interests in Blackfynn, a data integration company, and Grand Rounds, an online second opinion service.
Larsson Omberg has received research support from the National Institutes of Health/National Institute of Neurological Disorders and Stroke.
Jamie Adams has received research support from Biogen, BioSensics, Empire Clinical Research Investigator Program, Michael J. Fox Foundation, National Institutes of Health/National Institute of Neurological Disorders and Stroke, Pfizer, and Safra Foundation.
Katherine Amodeo serves as Co-Investigator for the Lewy Body Dementia Association Research Center of Excellence (LBDA-RCOE) at the University of Rochester Medical Center. She has served as PI and Co-I for PD and DLB clinical trials funded by Acadia Pharmaceuticals, EIP Pharma, Michael J. Fox Foundation, National Institutes of Health/National Institute of Neurological Disorders and Stroke, and Roche. She serves as medical monitor for a PD trial funded by Biogen. Dr. Amodeo’s fellowship in Movement Disorders was funded by the Safra Foundation and Michael J. Fox Foundation.
Erika Augustine has served as a consultant to Biomarin Pharma, Neurogene, Regenxbio, and Beyond Batten Disease Foundation. She has received research support from Abeona Therapeutics, Neurogene, and National Institutes of Health/National Institute of Neurological Disorders and Stroke. Dr. Augustine has served as central rater for a clinical trial funded by Signant Health.
Zachary Kabelac is co-founder of Emerald Innovations, Inc., which produces similar radio wave devices to those used in the reported research. Dr. Kabelac has been issued two patents: US9753131B2 and US20170042432A1.
Dina Katabi is co-founder of Emerald Innovations, Inc., which produces similar radio wave devices to those used in the reported research. Dr. Katabi has five patents, either pending or issued: WO2015102713A3, US20190188533A1, WO2018183106A1, WO2018013192A2, and US20170042432A1.
Karl Kieburtz has served as a consultant to Blackfynn, Clintrex, Genentech/Roche, and Novartis. He has received research support from National Institutes of Health (National Institute of Neurological Disorders and Stroke and National Center for Advancing Translational Sciences) and Michael J. Fox Foundation. Dr. Kieburtz has ownership interests in Clintrex, Hoover Brown LLC, and Safe Therapuetics.
Max Little has received an academic grant from the Michael J. Fox Foundation.
Suchi Saria has received research support from American Heart Association, Child Health Imprints, Defense Advanced Research Projects Agency, National Institutes of Health/National Institute of Neurological Disorders and Stroke, and National Science Foundation.
Giovanni Schifitto has received research support from the National Institutes of Health/National Institute of Neurological Disorders and Stroke.
Ruth Schneider has received research support from Acadia Pharmaceuticals, Biohaven Pharmaceuticals, Canadian Institute of Health, Massachusetts General Hospital, Michael J. Fox Foundation, National Institutes of Health/National Institute of Neurological Disorders and Stroke, Nuredis, Inc., and Pfizer.
Gaurav Sharma has received research support from MC10.
Ira Shoulson is founder and principal manager of Grey Matter Technologies LLC and is compensated by the company for work related to the reported research.
Christopher Tarolli has received research support from American Academy of Neurology Institute, BioSensics, Greater Rochester Health Foundation, Michael J. Fox Foundation, and National Institutes of Health/National Institute of Neurological Disorders and Stroke. He has received honoraria from the American Academy of Neurology and Davis Phinney Foundation. Dr. Tarolli has received clinical training support from Medtronic.
Michael McDermott has received research support from National Institutes of Health/National Institute of Neurological Disorders and Stroke and Parkinson Study Group. Dr. McDermott has been compensated by Cynapsus Therapetuics, Prilenia, and Voyager Therapeutics for service on a Data Safety and Monitoring Board.
Emma Waddell, Roy Adams, Rafayet Ali, Abigail Arky, Karthik Dinesh, Ehsan Hoque, Alistair Glidden, Estella Jensen-Roberts, Daniel Kinel, Karlo Lizarraga, Taylor Myers, Sara Riggare, Spencer Rosero, Anna Stevenson, and Jiebo Luo have no disclosures.
Footnotes
ACKNOWLEDGMENTS
Research reported in this publication was supported by the National Institute of Neurological Disorders and Stroke of the National Institutes of Health under Award Number P50NS108676. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
