Abstract
Background:
α-Synuclein has been related to the pathogenesis of Parkinson’s disease (PD), but it has not thoroughly been investigated in idiopathic rapid eye movement sleep behavior disorder (iRBD).
Objective:
We aimed to explore whether there were different distributions of α-synuclein at a genetic and/or protein level in patients with iRBD.
Methods:
We included 30 patients with iRBD, 30 patients with PD, and 30 age- and sex-matched healthy controls (HCs) in this study. The SNCA methylation and mRNA levels were determined using bisulfite sequencing and quantitative reverse transcription polymerase chain reaction. The plasma levels of exosome α-synuclein were measured using Meso Scale Discovery.
Results:
SNCA methylation showed different distribution among HC, iRBD and PD groups (HC vs RBD: p = 0.011; HC vs PD: p < 0.001; RBD vs PD: p = 0.027). However, plasma exosomal α-synuclein levels were only elevated in patients with PD compared to those in HCs (p = 0.027), and were associated with the SNCA methylation only in the PD group (p = 0.030, r = –0.397).
Conclusion:
SNCA hypomethylation in leukocytes existed both in patients with iRBD and those with PD, indicating that SNCA methylation could be a potential biomarker for early PD diagnosis.
INTRODUCTION
Idiopathic rapid eye movement (REM) sleep behavior disorder (iRBD) is characterized by vocalizations, jerks, and motor behaviors during REM sleep, often associated with REM-related dream content [1]. iRBD is associated with a high risk of progression to a neurodegenerative disease, such as Parkinson’s disease (PD) [1–6], which is characterized by the classical motor symptoms of parkinsonism that are due to α-synuclein aggregation [7]. As a manifestation of early-stage synucleinopathy diseases, iRBD is now regarded as a window for early potential disease-modifying intervention [8, 9]. Hence, it is important to determine the possible biomarkers for predicting the conversion from iRBD to PD.
The presence of α-synuclein in the gastrointestinal tract and skin tissue has been recognized in patients with PD, and similar findings are now emerging in patients with iRBD which suggested that RBD symptoms possibly reflected an underlying synucleinopathy [10, 11]. Phosphorylated α-synuclein was also found in the colonic mucosal, salivary glands, and skin biopsy samples of patients with iRBD and PD [12–15]. Moreover, many studies have shown that the plasma exosome α-synuclein level was substantially higher in patients with PD compared to that in controls [16]. Simultaneously, SNCA methylation levels in the blood were reported to be associated with PD progression [17, 18]. Methylation is a critical epigenetic mechanism to regulate SNCA transcription which transfers a methyl group from S-adenosyl methionine to cytosine residues at the CpG islands of SNCA intron-1 [19]. The above findings indicated that different forms of peripheral α-synuclein could be used as biomarkers of PD [13, 14].
In our study, we measured the SNCA methylation and mRNA expression levels in peripheral leukocytes, as well as the plasma exosome α-synuclein levels both in patients with iRBD and patients with PD. We aimed to build the peripheral α-synuclein profile in iRBD, which could provide potential biomarkers for the prediction of conversion from iRBD to PD in the future.
METHODS
Participants
This case-control study was approved by the Medical Ethics Committee of the Ruijin Hospital affiliated to the Shanghai Jiaotong University School of Medicine. All participants provided written informed consent. Outpatients with iRBD and outpatients with PD from the Department of Neurology, Ruijin Hospital, as well as age- and sex-matched healthy controls (HCs) were enrolled in this study. The diagnosis of PD was performed according to the International Parkinson and Movement Disorder Society (MDS) diagnostic criteria by at least two neurologists skilled in movement disorders [20]. Based on video-polysomnography analyzed by SINBAR method, iRBD patients were recruited according to standard International Classification of Sleep Disorders-Third Edition criteria (ICSD-3) excluding motor signs of Parkinsonism or secondary causes [21]. To avoid alterations of L-DOPA-induced epigenetic effects, we included only PD participants (onset of motor symptoms and diagnosis of PD within the past few years) who were naïve to L-DOPA medication. Subjects were excluded if they had a major organ dysfunction, other neurological disorders, or a family history of movement disorders.
Measurement of SNCA methylation and mRNA expression levels
Leukocytes were harvested from blood samples after centrifugation for 5 min at 2000 rpm and after adding 0.17 M ammonium chloride solution to lyse the red blood cells (RBCs). Genomic DNA was extracted from leukocytes using the Genomic DNA blood kit (GENERAY, Shanghai, China). Total RNA was extracted using the standard Trizol method (Tiangen, Beijing, China). Bisulfite conversion of genomic DNA was performed using EpiTect Fast Bisulfite Conversion Kits (Qiagen). A 426 bp SNCA promotor fragment containing 22 CpG sites was analyzed by bisulfite sequencing (Supplementary Figure 1). The SNCA promotor fragment was amplified by chain reaction (PCR) and cloned into the pGEM-T easy vector (Promega, Madison, WI, U.S.A.). At least 10 independent clones were sequenced for each subject. Quality control was performed by BiQ analyzer (quality control software for DNA methylation data from bisulfite sequencing). The level of DNA methylation was determined as the percentage of methylated CpG. For each subject, methylation level (methylated CpG/total CpG) at individual CpG sites and the mean methylation level of all the 22 CpG sites were calculated. cDNA was synthesized using PrimeScript RT reagent Kit with gDNA Eraser (Takara Biotech, Co., Dalian, China). Quantitative reverse transcription polymerase chain reaction (RT-qPCR) was performed on ABI 7500 Real-Time PCR System (Applied Biosytems, Inc., Foster City, CA), using TB Green Premix Ex Taq II (Takara Biotech, Co., Dalian, China). The primer sequences for methylation and RT-qPCR are described in Supplementary Tables 1 and 2.
Measurement of α-synuclein in plasma exosomes
Plasma samples were diluted with phosphate-buffered saline (PBS) after mixing it with buffer A and buffer B, followed by incubating the mixture with dynabeads on a rotator for 1 h at 4°C. Next, the mixture with dynabeads was incubated with PBS on a rotator for 15 min at 4°C and the supernatant that contains the plasma exosomes was collected.
Plasma exosomes were characterized based on the size and shape using transmission electron microscopy (TEM). The size of the samples was directly determined by nanoparticle tracking analysis (NTA) using a NanoSight LM10 microscope (NanoSight Ltd, Salisbury, UK). The degree of purity was verified using western blot with positive exosomal marker CD63 (Abcam, Cambridge, MA, USA) and Alix (Abcam, Cambridge, MA, USA), and negative exosomal marker GM130 (Abcam, Cambridge, MA, USA). The level of α-synuclein was determined using U-PLEX Human α-Synuclein Kit on the QuickPlex SQ120 platform (Meso Scale Discovery Co., Rockville, MD, USA).
Statistical analysis
All statistical analysis was performed using Statistical Product and Service Solutions (SPSS, version 21.0; IBM Corp., Armonk, NY). The Chi-square test was used to analyze data as frequencies and percentages. The normality of the distribution of descriptive statistics was tested by the Shapiro–Wilk test. One-way analysis of variance was used to ascertain the difference between variables. For post hoc multiple comparisons between HC, iRBD and PD groups, the LSD test was applied. Pearson’s correlation test was performed for determining the correlation between DNA methylation and the α-synuclein level in exosomes after adjusting for gender and age. For all analyses, p < 0.05 was considered statistically significant.
RESULTS
Clinical characteristics
The demographic and clinical data of all participants were presented in Table 1. There were no significant differences in age or gender among the three groups (PD, iRBD, and HC). 16 PD patients were classified as H-Y1-1.5 stage (early stage) and 14 PD patients as H-Y 2-3 (advanced stage). The scores of UPDRS were significantly different in part II between iRBD and PD patients.
Demographic and clinical data of subjects
HC, healthy controls; RBD, idiopathic rapid eye movement (REM) sleep behavior disorder; PD, Parkinson’s disease.
Comparison of the SNCA methylation level in HC, RBD, PD groups
We performed a methylation analysis of several PD-associated genes, but we only included herein the results that showed the difference in SNCA methylation levels among the three groups (PD, iRBD, and HC) (Supplementary Table 3). As shown in Fig. 1A, there was a significant difference in the mean methylation level of SNCA among the three groups. Moreover, a detailed comparison of the SNCA methylation levels at individual CpG sites among the three groups was further performed (Fig. 1C). Compared to HCs, in patients with PD, hypomethylated DNA was observed at the CpG sites 4, 6, 9, 10, 12, 15, 17, 18, and 21. In addition, the DNA of patients with PD was significantly hypomethylated at the CpG sites 17 and 18 compared to that of patients with iRBD. Furthermore, receiver operating characteristic (ROC) curves were performed to evaluate the diagnostic power of the SNCA methylation levels. The area under the ROC curve was 0.862 for the difference between the HC and PD groups and 0.679 for the difference between the HC and iRBD groups (Fig. 1B and Supplementary Table 4).
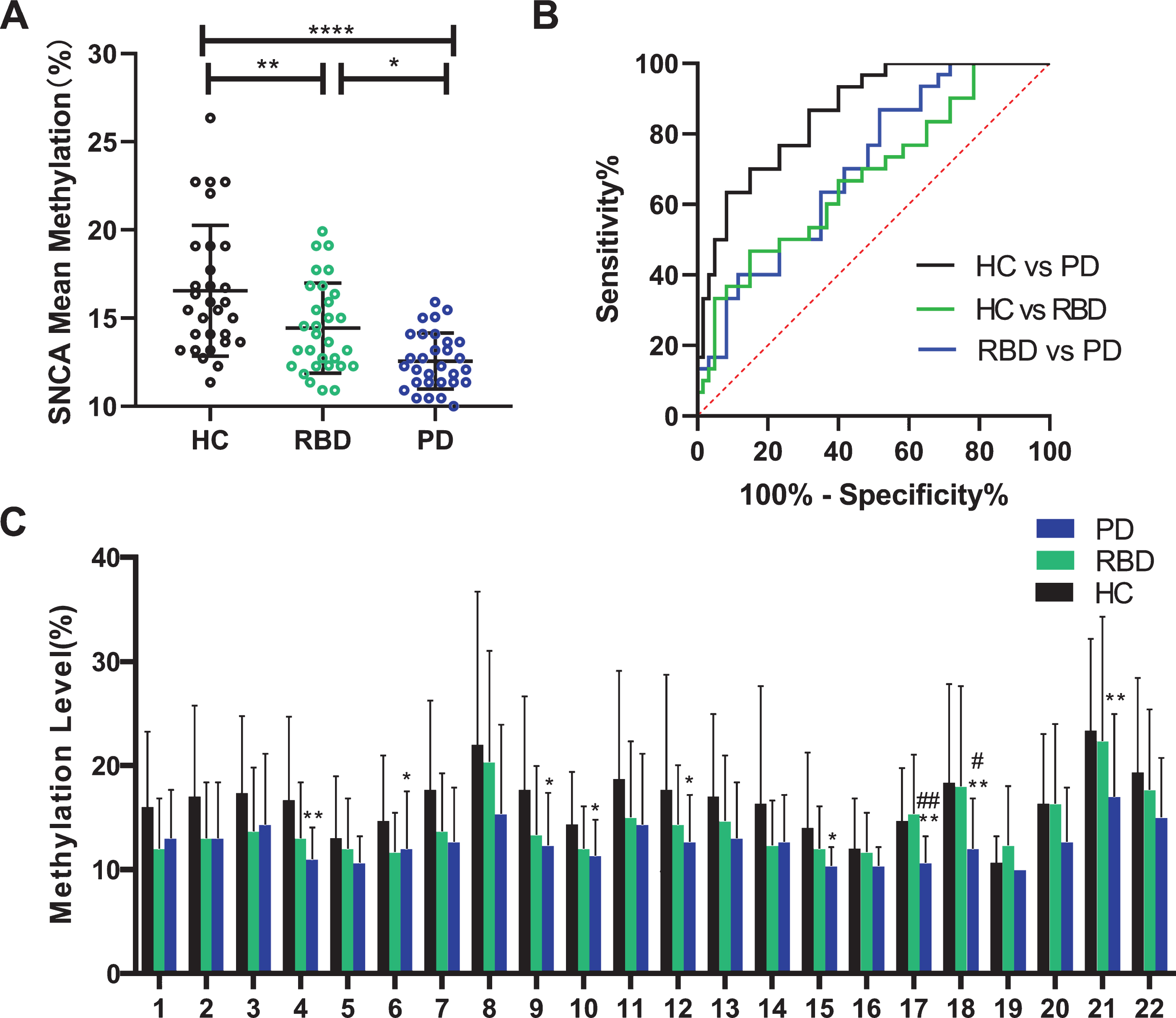
Methylation levels of SNCA in leukocytes from RBD, PD patients and healthy controls. A) Mean methylation of SNCA in all participants. B) Diagnostic performance of mean methylation of SNCA in HC, RBD and PD groups. C) Site-specific methylation Level of CpG island in SNCA gene. #RBD vs PD; *HC vs PD. HC, healthy controls; RBD, idiopathic rapid eye movement (REM) sleep behavior disorder; PD, Parkinson’s disease; *p < 0.05; **p < 0.01; ****p < 0.001.
Furthermore, these different SNCA methylation levels were revalidated in another independent cohort. The significance persisted after the analysis was performed to 50 iRBD patients, 100 matched PD patients and 100 matched controls (Supplementary Figure 3). As expected, the mean methylation levels were significantly decreased in patients with iRBD (p = 0.012) and PD (p < 0.001), compared to healthy controls. The CpG sites 17 and 18 continued to be significant in the comparison between PD and iRBD groups (Supplementary Figure 3C).
Comparison of the expression of SNCA in HC, RBD, PD groups
To evaluate the expression of SNCA in different groups, we measured the level of mRNA and exosomal protein. Plasma exosomes were verified for purity and size distribution in Supplementary Figure 1. First, no significant difference was found in the SNCA mRNA expression among the three groups (data not shown). Second, it was found that the concentrations of α-synuclein in plasma exosomes were significantly higher than those of HCs only in patients with PD (p = 0.027, Fig. 2A), but not in patients with iRBD. The ROC curves of the exosomal α-synuclein levels were shown in Fig. 2B and Supplementary Table 5. The discrimination between the HC and PD groups was more accurate than that between the HC and iRBD groups or that between the iRBD and PD groups, with a cutoff value of 69.65pg/ml. To explore whether the α-synuclein levels were associated with disease severity, we divided the patients with PD into two groups: the early PD group (H-Y 1-1.5) and the advanced PD group (H-Y 2-3). Exosomal α-synuclein levels of advanced PD were higher than early PD (92.2±26.4pg/ml vs 193.7±30.2pg/ml, p = 0.017; Fig. 2C).What’s more, exosomal α-synuclein levels showed a significant correlation with PD duration time (p = 0.025, r = 0.452, Fig. 2D).
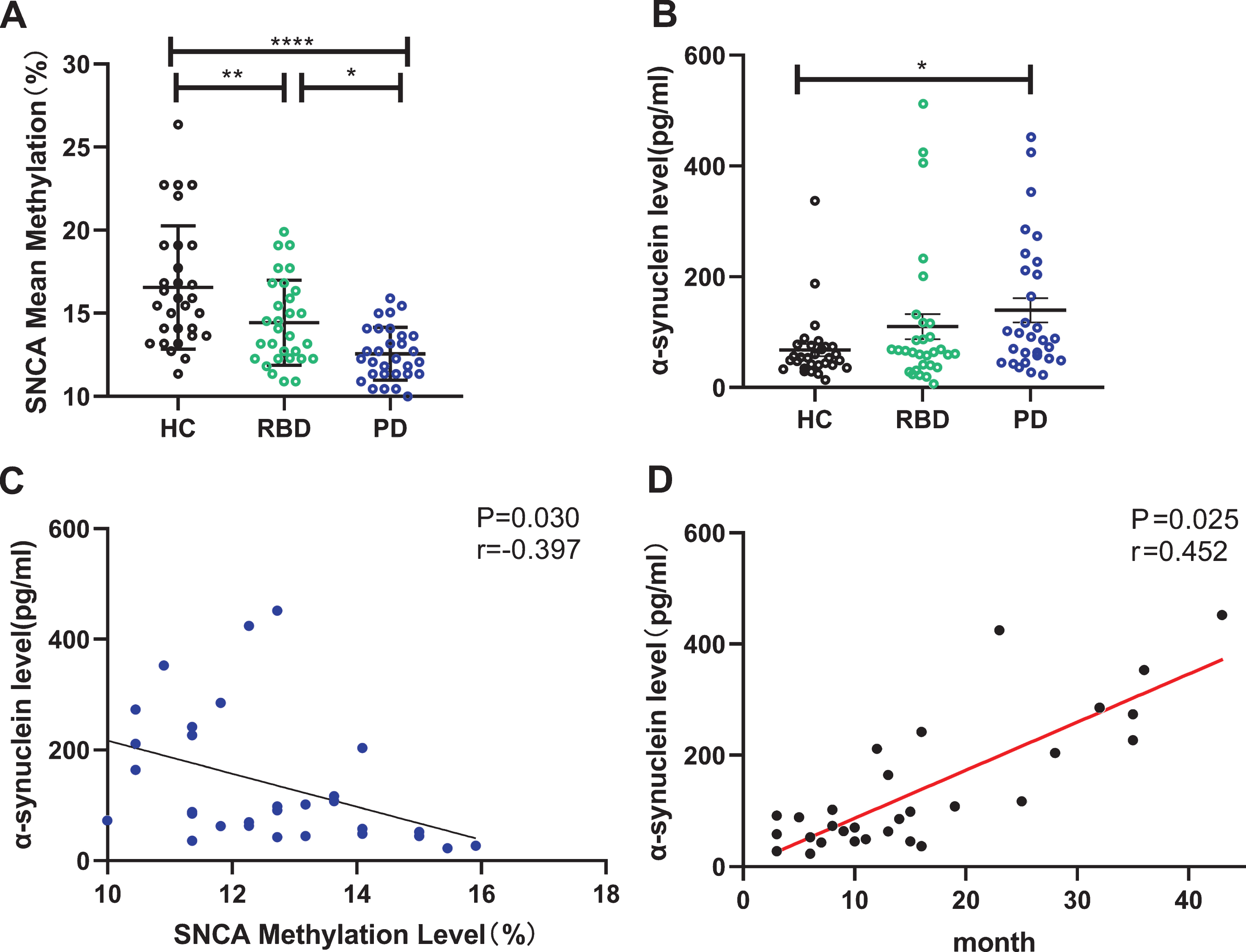
Diagnostic performance of exosomal α-synuclein and correlation to disease severity in PD group. A) α-synuclein level in plasma exosomes from HC, RBD and PD groups. B) Diagnostic performance of exosomal α-synuclein in HC, RBD and PD groups. C) Different exosomal α-synuclein levels in PD patients of early PD and advanced PD. D) Significant correlation between SNCA methylation level and PD duration. HC, healthy controls; RBD, idiopathic rapid eye movement (REM) sleep behavior disorder; PD, Parkinson’s disease.
Correlation between SNCA methylation and plasma exosomal α-synuclein and their risk for PD
We analyzed the relationship between the genetic level and protein expression of α-synuclein. As was shown in Fig. 3A, there was a clearly negative relationship between SNCA methylation level and plasma exosome α-synuclein level in all participants (p = 0.024, r = –0.238, Fig. 3A). Additionally, the SNCA methylation level was associated with the plasma exosomal α-synuclein levels in patients with PD (p = 0.030, r = –0.397, Fig. 3D). However, such a correlation was not found in HCs and patients with iRBD (Fig. 3B, C). Next, we performed binary logistic regression to evaluate whether the SNCA methylation levels and the α-synuclein levels were associated with the increased risk of PD between HC and PD groups (Supplementary Table 6). Lower SNCA methylation levels showed 3.9-fold risk (p = 0.045), while higher exosomal α-synuclein levels exhibited 4.0-fold increased risk for PD (p = 0.012).
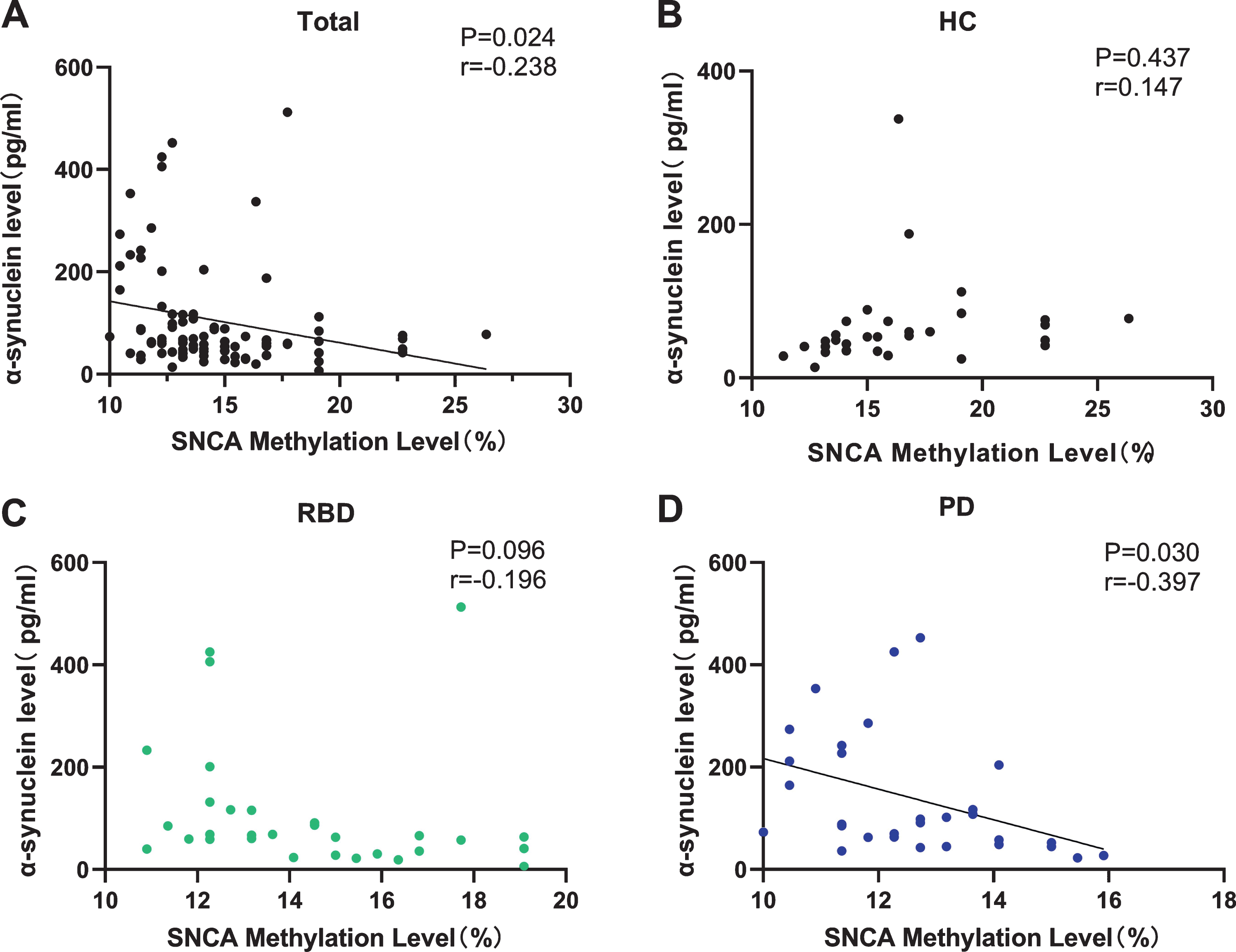
Correlation between SNCA methylation level and exosomal α-synuclein. A) Significant correlation between SNCA methylation level and exosomal α-synuclein in all subjects. B) No correlation between SNCA methylation level and exosomal α-synuclein in HC group. C) No correlation between SNCA methylation level and exosomal α-synuclein in RBD group. D) Significant correlation between SNCA methylation level and exosomal α-synuclein in PD group. HC, healthy controls; RBD, idiopathic rapid eye movement (REM) sleep behavior disorder; PD, Parkinson’s disease.
DISCUSSION
In this study, the different distributions of α-synuclein at a genetic and protein level were investigated to establish the potential biomarkers for RBD and PD. Our study found that (1) SNCA methylation was significantly decreased in both the iRBD and PD groups compared with HCs; (2) plasma exosomal α-synuclein levels were elevated in patients with PD compared to those in HCs associated with disease severity; (3) SNCA methylation level was associated with the plasma exosomal α-synuclein levels in patients with PD; (4) the SNCA methylation levels and the α-synuclein levels were associated with the increased risk of PD.
PD is characterized by the degeneration of dopaminergic neurons in the substantia nigra and is diagnosed based on the motor symptoms of bradykinesia, rigidity, and tremor [22]. However, neurodegeneration spreads over the central nervous system (CNS) long before the onset of typical motor signs [23]. This long prodromal period provides an important opportunity to diagnose PD much earlier than it is currently being diagnosed. RBD is considered to be a prodromal marker that is predictive of PD according to the MDS criteria [24], and the strong correlation between RBD and synucleinopathy may propose that some pathological changes typical for PD even appear at the RBD stage [25]. TMEM175 p.M393T may lead to impaired autophagy lysosomal pathway to induce α-synuclein aggregation which was recently reported to be associated with RBD [26]. Meanwhile a recent study discovered that several 5’ SNCA variants were associated with iRBD and PD patients with probable RBD [27]. However, investigation of iRBD biomarkers in the peripheral blood has not been thoroughly performed. Hence, we conducted a case-control study to identify possible biomarkers in patients with iRBD, which would provide evidence for the possibility of early prediction of synucleinopathy.
In the present work, we found that SNCA was hypomethylated in the leukocytes of patients with iRBD and PD compared with controls. This result is consistent with that of previous studies, which have found hypomethylation of SNCA in the brains of patients with PD and increased synucleinopathy in the brains of patients with iRBD [28, 29]. There have also been some discrepancies regarding the methylation level of SNCA in the peripheral blood. Some studies reported no evident changes in the DNA methylation level within the SNCA promoter region [30]. However, it should be noted that the medication L-DOPA increased the SNCA methylation in recent research [31]. Hence, in our study, the patients with PD were drug naïve to avoid such possible effects of this medication.
The CpG sites 17 and 18 of the SNCA gene were significantly different in the iRBD and PD groups. The genotype SNP rs3756063 lying between the CpG site 17 and 18 showed a significant correlation with the DNA methylation state of SNCA both in brain and blood PD samples [32]. Moreover, the genetic variant rs3756063 was associated with a possible RBD status in patients with PD, supporting the variety in SNCA methylation related to RBD symptoms [33]. Thus, this study further reinforced the notion that the changes in the methylation level of the SNCA genes in peripheral leukocytes may reflect the alterations in the methylation level in the brains of patients with PD or iRBD to a certain degree.
The association between iRBD and synucleinopathies is mainly due to the impairment of specific brainstem areas, particularly the mesencephalon and pontine tegmentum, which are involved in the pathogenesis of both conditions [34]. Nevertheless, only a very limited number of studies have been performed in patients diagnosed with iRBD at the prodromal stage, without evidence of motor signs. It was found that the levels of α-synuclein in the cerebrospinal fluid and serum in the PD with RBD group were elevated compared with those in the PD without RBD group[35]. Our results indicated that the alteration in SNCA methylation may occur even in the very early stage of PD.
It has been reported that RBCs are the major source of α-synuclein in the blood [36]. Considering the abundance and fragility of the RBCs, α-synuclein levels in the plasma may be artificially elevated by contamination with intact or lysed RBCs. Therefore, we selected plasma exosomes to evaluate the α-synuclein expression. The α-synuclein levels in plasma exosomes were significantly different between the HC and PD groups, but not in the RBD group. Recent studies showed that plasma exosomal α-synuclein likely originated from the CNS and correlated with the disease severity [16]. Furthermore, the SNCA methylation levels in our study were associated with the exosomal α-synuclein expression, which was consistent with the results of a study on brains with PD [29]. Thus, our findings indicated that SNCA methylation may play a role in the activation of the SNCA expression.
However, the SNCA mRNA levels in this study showed no obvious change in patients with iRBD and PD. Previous results on SNCA mRNA levels in patients with PD were conflicting [34, 35]. A small sample size, contamination with RBCs [37, 38], and the unavoidable degradation of mRNA [39] may partially explain the different results among studies. Similarly, small sample size may also lead to that α-synuclein in plasma exosomes failed to reach statistical significance in iRBD vs HC. Meanwhile, considering the increasing accumulation of α-synuclein during disease progression, most of our enrolled PD patient were at the earlier stages which may partially contribute to the decreased difference. Also, other forms of α-synuclein (phosphorylated and oligomeric α-synuclein) may be needed to form an ‘ideal’ panel to distinguish RBD and PD in plasma exosomes.
Taken together, our results revealed SNCA hypomethylation in the leukocytes of patients with iRBD and PD that correlated with the plasma exosomal α-synuclein level in patients with PD. These findings indicated that SNCA hypomethylation may occur earlier than plasma exosomal α-synuclein expression in iRBD. Therefore, SNCA methylation could be a potential biomarker for early PD diagnosis.
CONFLICT OF INTEREST
The authors have no conflict of interest to report.
Footnotes
ACKNOWLEDGMENTS
We thank all the patients and healthy controls who participated in this study.
This study was supported by grants from the National Key Research and Development Program (2016YFC1306505); the National Natural Science Foundation of China (81873778); the National Natural Science Foundation of China (81501097); the Shanghai Municipal Science and Technology Commission-Basic Key Project (18JC1420300); and the Shanghai Clinical Collaboration Construction Project of Chinese and Western Medicine [ZY(2018-2020)-FWTX-1104].
